<Sameen Nasar> — HTGAA Spring 2026

About me
🧬 London | LSHTM 📈
INTERESTS: Biology, Genomics, Epidemiology, Public Health, Genetic Engineering, Political Economy, Science Fiction, Comic Books, Nutrition
Contact info
Homework
Class Assignment — DUE BY START OF FEB 10 LECTURE
(1) First, describe a biological engineering application or tool you want to develop and why. This could be inspired by an idea for your HTGAA class project and/or something for which you are already doing in your research, or something you are just curious about.
By leveraging biological engineering tools, such as CRISPR systems, I would like to develop highly specific nucleic acid biosensors and synthetic circuits to detect M. tuberculosis and resistance mutations with high precision and speed. The inspiration for this comes from working on my MSc project, where I studied the genomic epidemiology of multi-drug-resistant tuberculosis (MDR-TB) using WGS data. My work focused on downstream analyses (phylogenetics, transmission clustering, regression, and machine learning), with particular attention to population structure and epidemiological interpretation. However, when working on my project, I found that genomic data of MDR-TB is geographically imbalanced, limiting the representativeness of global MDR-TB patterns and, ultimately, timely detection and treatment. This is especially true in high burden countries. As a result, I would like to explore the application of biosensors and genetic circuitry to add an additional layer of surveillance alongside traditional methods; biosensor or genetic circuit engineered to detect specific MDR-TB resistance markers or lineage-specific sequences, potentially using luminescence as a real-time readout to provide rapid, high-throughput signals.
Brief on the biology and possible mechanism for the tool: 🛠️ 🧬
Unlike many other bacteria that can share drug‑resistance genes with each other through horizontal gene transfer, Mycobacterium tuberculosis mainly becomes drug resistant through mutations in its own DNA (Single Nucleotide Polymorphisms (SNPs), insertions/deletions (indels)) [1]. Simultaneously, the ability of M. tuberculosis to persist within human hosts exposes it to prolonged immune pressure, driving adaptive changes in virulence‑associated loci such as phoR, mymA and the mce1 operon that can influence how different lineages transmit or interact with particular human populations [2¬–4]. As a result, the proposed bio-engineering tool could take the form of a bio-sensor, where CRISPR-based device could be programmed to recognise TB resistance mutations or an engineered genetic circuit that only produces a light or electrical signal when multiple resistance signatures are present. Such a device would convert the presence of specific mutations into a measurable output that can be rapidly read and fed into surveillance models.
(2) Next, describe one or more governance/policy goals related to ensuring that this application or tool contributes to an “ethical” future, like ensuring non-malfeasance (preventing harm). Break big goals down into two or more specific sub-goals. Below is one example framework (developed in the context of synthetic genomics) you can choose to use or adapt, or you can develop your own. The example was developed to consider policy goals of ensuring safety and security, alongside other goals, like promoting constructive uses, but you could propose other goals for example, those relating to equity or autonomy.
Governance Goal 1: Prevent harm or misuse
As genomic data can be geo-located and time-stamped, there are risks for community stigmatization and political duress. Therefore, to mitigate against these risks, the governance goal should implement frameworks that: (i) Require ethical review and oversight of bio-sensor data and its secondary uses (ii) Establish strict guidelines on the limits of how precise location data can be shared or publicized (iii) Establish clear accountability mechanisms for state and private actors
Governance Goal 2: Promote equity in data collection, analysis and development
To prevent further exacerbation of inequities biological data collection and usage, the framework will implement mechanisms that ensure: (i) Control of locally generated data by implementing country (ii) Inclusion of implementing country as equal partners in analysis and interpretation (iii) Prioritization of under-sampled regions to improve representativeness and combining outputs with timely access to treatment and care.
(3) Next, describe at least three different potential governance “actions” by considering the four aspects below (Purpose, Design, Assumptions, Risks of Failure & “Success”). Try to outline a mix of actions (e.g. a new requirement/rule, incentive, or technical strategy) pursued by different “actors” (e.g. academic researchers, companies, federal regulators, law enforcement, etc). Draw upon your existing knowledge and a little additional digging, and feel free to use analogies to other domains (e.g. 3D printing, drones, financial systems, etc.).
- Purpose: What is done now and what changes are you proposing?
- Design: What is needed to make it “work”? (including the actor(s) involved - who must opt-in, fund, approve, or implement, etc)
- Assumptions: What could you have wrong (incorrect assumptions, uncertainties)?
- Risks of Failure & “Success”: How might this fail, including any unintended consequences of the “success” of your proposed actions?
Governance Action 1: Regulation and creation of standards for early-stage bio-sensor development
Purpose: Early-stage bio-sensor development research is guided by bio/genetic engineering but requires safety and bio-security risks. I am proposing specific standards and regulatory requirements for early-stage biosensor design, ensuring safety, transparency, and responsible innovation before deployment. This could be in the form of new regulatory support or reference diagnostics.
Design: Actors may include public health agencies, national regulators in science, and diagnostic developers. Establish validation criteria, accuracy thresholds, metadata standards, and geolocation safeguards. In addition, embed standards into existing public health TB surveillance programmes.
Assumptions: This initiative assumes that regulators will be quick to evaluate bio-sensor technologies. Also assumes public health surveillance will be quick to agree and implement technology across the existing surveillance system.
Risk of failure: Bureaucracy may hinder technological innovation and deployment. Unintended consequences include a premature reliance on bio-sensor technology which could lead to false positive cases and mis-directed public health strategies.
Governance Action 2: Pre-sequencing rapid signal regulatory pathways
Purpose: Currently, bio-sensor outputs such as CRISPR signals and genomic data are not integrated in low to middle-income countries Therefore, I would like to propose the creation of formal pathways that enable rapid biosensor signals to feed into surveillance systems before whole genome sequencing (WGS), with defined quality, privacy, and data use standards.
Design: Actors include public health agencies, national regulators in science, and diagnostic developers. Actors may also include international bodies such as the WHO. There may be potential to expand the WHO’s ‘attributes and principles on genomic data-sharing platforms supporting surveillance of pathogens’ [5–7].
Assumptions: This assumes developers implement required standards and metadata. Also assumes public health agencies can incorporate new signal streams effectively.
Risk of failure: Disagreements about implementation into existing surveillance pathways. State agencies may lack technical expertise to train workers to evaluate, interpret and act on rapid biosensor signals. This could lead to misinterpretation and/or delayed action
Governance Action 3: Ethical data access and sharing standards (with local and community engagement requirements)
Purpose: Many genomic and bio-engineering projects lack consistent standards for privacy, consent, equity, and local engagement. A proposed change could be the mandatory implementation of ethical standards for data access combined with mandatory local/community engagement, ensuring transparency, and equitable benefit-sharing.
Design: Develop standardised model data agreements which specify permissible uses, benefit-sharing obligations, and consent mechanisms. Furthermore, advisory boards and steering committees can be established to ensure engagement, feedback, and regular assessment of processes.
Assumptions: This assumes that communities where the technology is planned to be implemented will agree to engage meaningfully. It also assumes that cross-country coordination on ethical standards will be possible.
Risk of failure: Strict data provisions may slow down implementation, collection and action. There may be failure to engage communities as they may view the initiative to engage them as superficial.
(4) Next, score (from 1-3 with, 1 as the best, or n/a) each of your governance actions against your rubric of policy goals. The following is one framework but feel free to make your own:
| Does the option: | Option 1 | Option 2 | Option 3 |
|---|---|---|---|
| Regulation and creation of standards for early-stage bio-sensor development | Pre-sequencing rapid signal regulatory pathways | Ethical data access and sharing standards | |
| 🦠🛡️Enhance Biosecurity | |||
| • By preventing incidents | 1 | 2 | 2 |
| • By helping respond | 2 | 3 | 2 |
| 🧪Foster Lab Safety | |||
| • By preventing incidents | 2 | 2 | 2 |
| • By helping respond | 3 | 3 | 2 |
| 🌱Protect the environment | |||
| • By preventing incidents | 1 | 2 | 2 |
| • By helping respond | 2 | 1 | 1 |
| ⚖️Other considerations | |||
| • Minimizing costs and burdens to stakeholders | 1 | 1 | 1 |
| • Feasibility | 2 | 2 | 2 |
| • Not impede research | 2 | 2 | 2 |
| • Promote constructive applications | 1 | 1 | 1 |
5. Last, drawing upon this scoring, describe which governance option, or combination of options, you would prioritize, and why. Outline any trade-offs you considered as well as assumptions and uncertainties. For this, you can choose one or more relevant audiences for your recommendation, which could range from the very local (e.g. to MIT leadership or Cambridge Mayoral Office) to the national (e.g. to President Biden or the head of a Federal Agency) to the international (e.g. to the United Nations Office of the Secretary-General, or the leadership of a multinational firm or industry consortia). These could also be one of the “actor” groups in your matrix.
Based on the inputs and ranking in the matrix above, I would prioritize the following:
(i) Regulation and creation of standards for early-stage bio-sensor development (ii) Ethical data access and sharing standards with local community engagement
Together both these actions would address both the technical and social foundations required for responsible deployment of biosensors. Standards would ensure that biosensors are developed safely, setting incentive structures to develop lab safety protocols and enforce biosecurity. Local community engagement, training, and capacity building will help build trust, protect rights, and enable effective use of surveillance data across settings.
References
Richard M. Jones, Kristin N. Adams, Hassan E. Eldesouky, and David R. Sherman “The evolving biology of mycobacterium tuberculosis drug resistance.” Frontiers in Cellular and Infection Microbiology 2022.
Sebastien Gagneux “Ecology and evolution of mycobacterium tuberculosis.” Nature Reviews Microbiology 2018.
Qingyun Liu, Jianhao Wei, Yawei Li, Mei Wang, Jun Su, et al. “Mycobacterium tuberculosis clinical isolates carry mutational signatures of host immune environments.” Science Advances 2020.
Á. Chiner-Oms, L. Sánchez-Busó, J. Corander, S. Gagneux, S. R. Harris, et al. “Genomic determinants of speciation and spread of the mycobacterium tuberculosis complex.” Science Advances 2019.
World Health Organization. Attributes and principles of genomic data-sharing platforms supporting surveillance of pathogens with epidemic and pandemic potential. World Health Organization; 2025.
Carter L, Yu MA, Sacks J, Barnadas C, Pereyaslov D, Cognat S, et al. Global genomic surveillance strategy for pathogens with pandemic and epidemic potential 2022–2032. Bulletin of the World Health Organization. 2022 Apr 1;100(04):239–9A.
Trump BD, Florin MV, Perkins E, et al. Biosecurity for Synthetic Biology and Emerging Biotechnologies: Critical Challenges for Governance. 2021 Sep 8. In: Trump BD, Florin MV, Perkins E, et al., editors. Emerging Threats of Synthetic Biology and Biotechnology: Addressing Security and Resilience Issues [Internet]. Dordrecht (DE): Springer; 2021. Chapter 1. Available from: https://www.ncbi.nlm.nih.gov/books/NBK584259/ doi: 10.1007/978-94-024-2086-9_
Assignment (Week 2 Lecture Prep)
Homework Questions from Professor Jacobson
Nature’s machinery for copying DNA is called polymerase. What is the error rate of polymerase? How does this compare to the length of the human genome. How does biology deal with that discrepancy?
The error rate of polymerase is 1 error per 10⁶ nucelotides, where this can range from expected error frequency from 1 error per 104 to approximately 106 [1]. The human genome has 3 x 109 base pairs, this is around 3 billion nucleotides. This is much larger (approx. 3000 times) than 10⁶-nucleotide error rate of polymerase. Biology deals with this through a process of proofreading; cells use polymerase proofreading and mismatch repair to reduce errors to just a few per genome per replication [2].
How many different ways are there to code (DNA nucleotide code) for an average human protein? In practice what are some of the reasons that all of these different codes don’t work to code for the protein of interest?
Average Human Protein: 1036 bp As 1 codon = 3 nucleotides
∴ Total amino acids = 1036/3 ~ 345
Given 3 nuclotide-codons and 1 codon codes for 1 amino acid, there are 3345 different ways to code for an average human protein.
Given 3345 DNA sequences code for the same protein, only some of it works due to codon preferences and bias, repetitive or unstable sequences, and mRNA folding [3].
References
Kunkel TA, Bebenek K. DNA replication fidelity. In: Brenner S, Miller JH, editors. DNA Replication and Human Disease. Bethesda (MD): National Center for Biotechnology Information (US); 2002. Available from: [https://www.ncbi.nlm.nih.gov/books/NBK9940/](https://www.ncbi.nlm.nih.gov/books/NBK9940/]
Alberts B, Johnson A, Lewis J, Raff M, Roberts K, Walter P. Molecular Biology of the Cell. 4th ed. New York: Garland Science; 2002. ISBN: 0-8153-3218-1, 0-8153-4072-9.
Lin J, Chen Y, Zhang Y, Lin H, Ouyang Z, et al. Deciphering the role of RNA structure in translation efficiency. BMC Bioinformatics. 2022;23:559
Homework Questions from Dr. LeProust
What’s the most commonly used method for oligo synthesis currently?
Oligonucleotide synthesis is the chemical process of making short fragments of DNA or RNA with a defined sequence, typically using step‑by‑step addition of nucleotide building blocks on a solid support [1]. For enzyme-free synthesis, the process involves sequentially adding nucleotide units to a growing chain, typically using solid- or liquid-phase synthesis [2]. The most common method is solid phase oligo phosphoramidite synthesis. As it is now automated and uses high quality short sequences, it is widely used in biotech companies around the world [3–4].
Why is it difficult to make oligos longer than 200nt via direct synthesis?
As length is increased, chemical synthesis becomes less efficient. As a result, there is a loss in product yield, greater rate of error accumulation (higher substitution or deletion rates), and an increased difficulty in purifying the final product due to the introduction of truncated and mis-incorporated oligos [5].
Why can’t you make a 2000bp gene via direct oligo synthesis?
As oligosynthesis adds one nucleotide at a time, increasing length will lead to a greater accumulation of errors (substitutions/deletions). The truncated or defective sequences become increasingly difficult to purify [6]. Therefore, direct synthesis of a 2000bp gene is not practical despite surface-based methods and capture-based purification [7].
References
Beaucage SL, Caruthers MH. Deoxynucleoside phosphoramidites—A new class of key intermediates for deoxypolynucleotide synthesis. Tetrahedron Letters. 1981;22(20):1859–62. doi:10.1016/S0040-4039(01)90461-7.
Bachem. What is oligonucleotide synthesis & how does it work? [Internet]. Bubendorf: Bachem; 2024 Aug 26 [cited 2026 Feb 10]. Available from: https://www.bachem.com/articles/oligonucleotides/how-does-oligonucleotide-synthesis-work/
ScienceDirect. Oligonucleotide synthesis [Internet]. Amsterdam: Elsevier; 2024 [cited 2026 Feb 10]. Available from: https://www.sciencedirect.com/topics/biochemistry-genetics-and-molecular-biology/oligonucleotidesynthesis
ATDBio. Solid-phase oligonucleotide synthesis: The Phosphoramidite method [Internet]. Southampton: ATDBio; 2024 [cited 2026 Feb 10]. Available from: https://atdbio.com/nucleic-acids-book/Solid-phase-oligonucleotide-synthesis#The-Phosphoramidite-method
Kosuri S, Church GM. Large-scale de novo DNA synthesis: technologies and applications. Nat Methods. 2014;11:499–507. doi:10.1038/nmeth.2918.
Pichon M, Hollenstein M. Controlled enzymatic synthesis of oligonucleotides. Commun Chem. 2024;7:138. doi:10.1038/s42004-024-01216-0.
Yin Y, Arneson R, Yuan Y, Fang S. Long oligos: direct chemical synthesis of genes with up to 1728 nucleotides. Chem Sci. 2025;16:1966–73. doi:10.1039/D4SC06958G.
Homework Question from George Church
Choose ONE of the following three questions to answer; and please cite AI prompts or paper citations used, if any.
1. [Using Google & Prof. Church’s slide #4] What are the 10 essential amino acids in all animals and how does this affect your view of the “Lysine Contingency”?
Essential amino acids are defined as the amino acids that the animal body cannot synthesize, and therefore must obtain from diet. The essential amino acids in animals are are: isoleucine, leucine, lysine, threonine, tryptophan, methionine, histidine, valine, and phenylalanine. In addition, cysteine and tyrosine are often described as conditionally essential because they cannot be synthesized de novo in animals and are instead produced from methionine and phenylalanine, respectively [1].
Given lysine is one of essential amino acids that is universal for all animals, the “Lysine Contingency” is not an exclusive real control mechanism. Even if it hypothetically existed and could be removed, animals could easily source it from food, either meats, beans, or grains.
References
- Hou Y, Wu G. Nutritionally essential amino acids. Adv Nutr. 2018;9(6):849–851. doi:10.1093/advances/nmy054
Homework Week 2
Part 1: Benchling & In-silico Gel Art
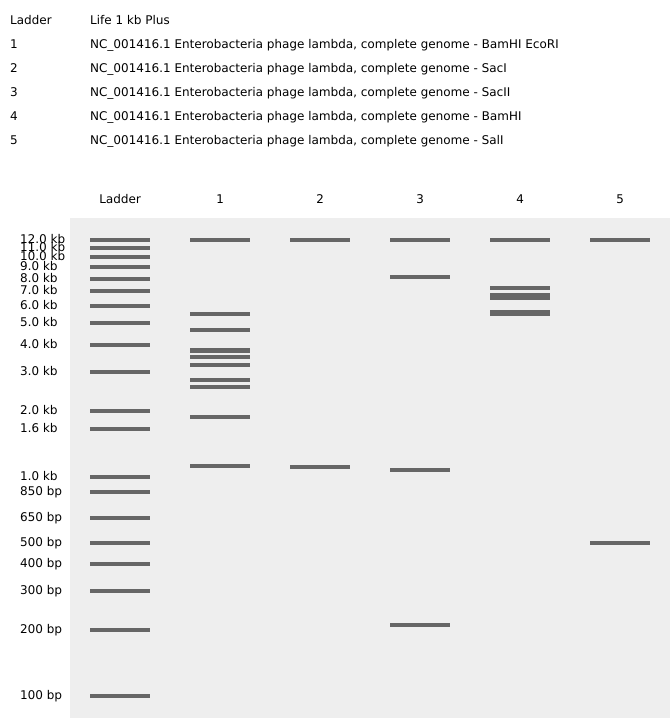
Import the Lambda DNA. Simulate Restriction Enzyme Digestion with the following Enzymes:
EcoRI
HindIII
BamHI
KpnI
EcoRV
SacI
SalI
Create a pattern/image in the style of Paul Vanouse’s Latent Figure Protocol artworks.
You might find Ronan’s website a helpful tool for quickly iterating on designs!
Playing around with the digest enzymes


.png)
.png)
Getting an “S”, well…sort of:


Part 3: DNA Design Challenge
3.1. Choose your protein.: In recitation, we discussed that you will pick a protein for your homework that you find interesting. Which protein have you chosen and why? Using one of the tools described in recitation (NCBI, UniProt, google), obtain the protein sequence for the protein you chose. [Example from our group homework, you may notice the particular format — The example below came from UniProt]
I have chosen Tumor Necrosis Factor- Alpha (TNF-α).
Why:
Reasons for choosing this protein include my interest in dermatology and chronic diseases. It is a key inflammatory cytokine in many skin and insulin resistant conditions. I am interested in psoriasis, particularly plaque psoriasis and its relation to insulin resistance and diabetes [1]. This is because this is something my Mum has suffered from the last couple of years, recently developing some pre-diabetes.
Protein Sequence:
NP_000585.2 tumor necrosis factor [Homo sapiens] MSTESMIRDVELAEEALPKKTGGPQGSRRCLFLSLFSFLIVAGATTLFCLLHFGVIGPQREEFPRDLSLI SPLAQAVRSSSRTPSDKPVAHVVANPQAEGQLQWLNRRANALLANGVELRDNQLVVPSEGLYLIYSQVLF KGQGCPSTHVLLTHTISRIAVSYQTKVNLLSAIKSPCQRETPEGAEAKPWYEPIYLGGVFQLEKGDRLSA EINRPDYLDFAESGQVYFGIIAL
References
- Moller DE. Potential role of TNF-α in the pathogenesis of insulin resistance and type 2 diabetes. Trends Endocrinol Metab. 2000 Aug;11(6):212-217. doi:10.1016/S1043-2760(00)00272-1.
3.2. Reverse Translate: Protein (amino acid) sequence to DNA (nucleotide) sequence.
To obtain the nucleotide sequence encoding TNF-α, I retrieved the validated human mRNA record (NCBI RefSeq: NM_000594.4) from NCBI. From this record, I extracted the coding sequence (CDS), which corresponds to the protein sequence NP_000585.2. Only the CDS was used for downstream codon optimization. See below:
ATGAGCACTGAAAGCATGATCCGGGACGTGGAGCTGGCCGAGGAGGCGCTCCCCAAGAAGACAGGGGGGCCCCAGGGCTCCAGGCGGTGCTTGTTCCTCAGCCTCTTCTCCTTCCTGATCGTGGCAGGCGCCACCACGCTCTTCTGCCTGCTGCACTTTGGAGTGATCGGCCCCAGAGGGAAGAGTTCCCCAGGGACCTCTCTCTAATCAGCCCTCTGGCCCAGGCAGTCAGATCATCTTCTCGAACCCCGAGTGACAAGCCTGTAGCCCATGTTGTAGCAAACCCTCAAGCTGAGGGGCAGCTCCAGTGGCTGAACCGCCGGGCCAATGCCCTCCTGGCCAATGGCGTGGAGCTGAGAGATAACCAGCTGGTGGTGCCATCAGAGGGCCTGTACCTCATCTACTCCCAGGTCCTCTTCAAGGGCCAAGGCTGCCCCTCCACCCATGTGCTCCTCACCCACACCATCAGCCGCATCGCCGTCTCCTACCAGACCAAGGTCAACCTCCTCTCTGCCATCAAGAGCCCCTGCCAGAGGGAGACCCCAGAGGGGGCTGAGGCCAAGCCCTGGTATGAGCCCATCTATCTGGGAGGGGTCTTCCAGCTGGAGAAGGGTGACCGACTCAGCGCTGAGATCAATCGGCCCGACTATCTCGACTTTGCCGAGTCTGGGCAGGTCTACTTTGGGATCATTGCCCTGTGA
3.3. Codon optimization. Once a nucleotide sequence of your protein is determined, you need to codon optimize your sequence. You may, once again, utilize google for a “codon optimization tool”. In your own words, describe why you need to optimize codon usage. Which organism have you chosen to optimize the codon sequence for and why?
For codon optimization, I chose the online codon optimizing tool:
https://en.vectorbuilder.com/tool/codon-optimization.html
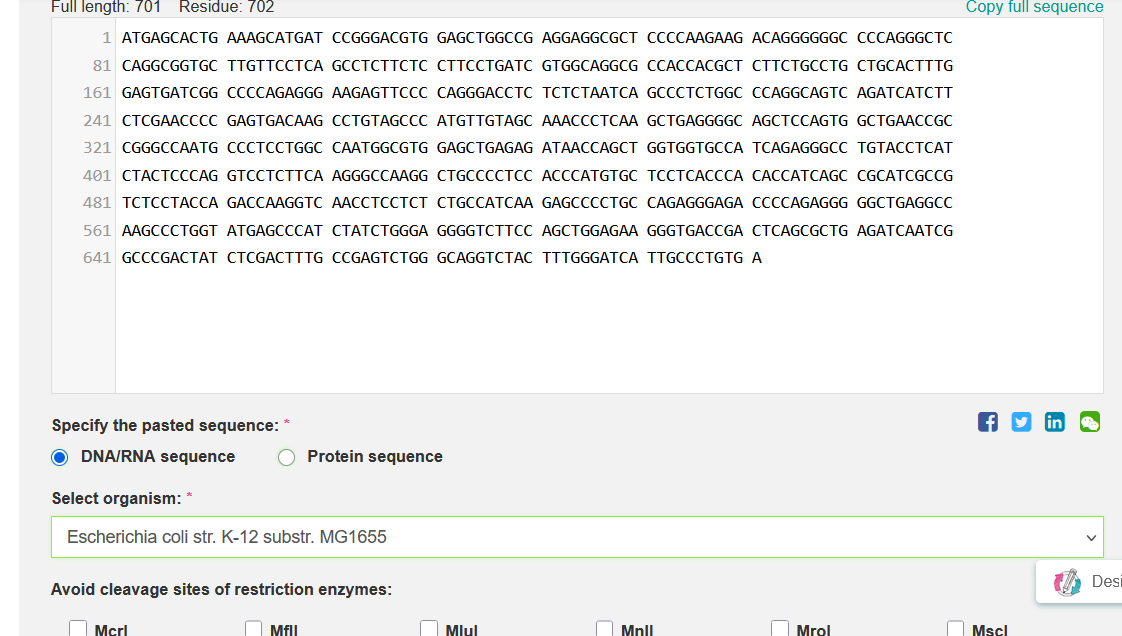

From my input:
I got: Pasted Sequence: GC=59.84%, CAI=0.49
From my output:
Improved DNA[1]: GC=59.97%, CAI=0.92
For CAI (Codon Adaptation Index), this indicates strong expected expression.
For GC content, after optimization it remained near 60%, within a suitable range for Escherichia coli, supporting stable and efficient gene synthesis.
I selected Escherichia coli strain K-12 MG1655 as the target organism for codon optimization because it is a well-studied laboratory strain with a completely sequenced and annotated genome [1–2].
Codon Optimized TNF-Alpha for improved expression of Escherichia coli
CTGAGCCCGTTCAACAACCCGCTGCTGCGCCCGTTTCTGATTCTGTATGAACATTAAAAACATGATCCGGGCCGTGGCGCAGGTCGCGGCGGCGCGCCGCAGGAAGATCGTGGCGCACCGGGCTTACAGGCCGTGCTGGTTCCGCAGCCGCTGCTGCTGCCGGATCGCGGCCGTCGTCACCATGCCCTGCTGCCGGCGGCCCTGTGGTCGGATCGTCCGCAGCGTGAAGAATTTCCGCGCGATCTGAGCCTGATTAGCCCGCTGGCGCAGGCCGTGCGTAGCAGCAGCCGCACCCCGTCAGATAAACCGGTGGCGCACGTGGTGGCAAATCCGCAGGCCGAAGGTCAGCTGCAGTGGCTGAATCGTCGCGCGAATGCCCTGTTAGCCAATGGTGTGGAACTGCGCGATAATCAGCTGGTGGTGCCGTCAGAAGGTCTGTACCTGATCTATTCGCAGGTGCTGTTTAAAGGCCAGGGCTGTCCGAGCACCCATGTGCTGCTGACCCACACCATTAGCCGCATTGCGGTGAGCTACCAGACCAAAGTGAACCTGCTTTCTGCGATTAAAAGCCCGTGCCAGCGTGAAACCCCGGAAGGCGCGGAAGCGAAACCGTGGTACGAACCGATTTATCTGGGCGGCGTGTTCCAGCTGGAAAAAGGCGATCGTCTGAGCGCGGAAATTAATCGCCCGGATTATCTGGATTTTGCGGAAAGCGGTCAGGTGTATTTCGGCATTATTGCCTTGTAA
References
Lukjancenko O, Wassenaar TM, Ussery DW. Comparison of 61 sequenced Escherichia coli genomes. Microb Ecol. 2010 Nov;60(4):708-20. doi:10.1007/s00248-010-9717-3. PMID:20623278; PMCID:PMC2974192.
Yannai A, Katz S, Hershberg R. The codon usage of lowly expressed genes is subject to natural selection. Genome Biol Evol. 2018 May;10(5):1237–46. doi:10.1093/gbe/evy084.
3.4. You have a sequence! Now what?
What technologies could be used to produce this protein from your DNA? Describe in your words the DNA sequence can be transcribed and translated into your protein. You may describe either cell-dependent or cell-free methods, or both.
After codon optimizing the TNF- α DNA sequence, it can be used to produce protein either through cell-dependent or cell-free systems.
For cell-dependent systems, the DNA will first need to be cloned using and inserted into an expression vector, this is then introduced into live host cells such as E. coli or eukaryotic cells, where cellular machinery transcribes the DNA into mRNA and then translates the mRNA into TNF‑α protein during growth and metabolism; this is seen in standard biotechnology production processes [1–2].
For cell-free systems, crude cell extracts provide all the machinery for transcription, translation, protein folding, and energy metabolism [3]. Therefore, when the codon optimized DNA is added, the TNF‑α protein will be produced in-vitro and under controlled conditions.
Both these methods rely on the flow of information from DNA to mRNA to protein; the Central Dogma of Molecular Biology.
References
Lukjancenko O, Wassenaar TM, Ussery DW. Comparison of 61 sequenced Escherichia coli genomes. Microb Ecol. 2010 Nov;60(4):708-20. doi: 10.1007/s00248-010-9717-3. Epub 2010 Jul 11. PMID: 20623278; PMCID: PMC2974192.
Swartz JR. Advances in Escherichia coli production of therapeutic proteins. Curr Opin Biotechnol. 2001 Oct;12(5):195–201. doi:10.1016/s0958-1669(00)00198-5. PMID:11513436.
Carlson ED, Gan R, Hodgman CE, Jewett MC. Cell-free protein synthesis: Applications come of age. Biotechnol Adv. 2012 Sep-Oct;30(5):1185-94. doi:10.1016/j.biotechadv.2011.09.016. PMID:22001003; PMCID:PMC3359644.
3.5. [Optional] How does it work in nature/biological systems?
1. Describe how a single gene codes for multiple proteins at the transcriptional level. 2. Try aligning the DNA sequence, the transcribed RNA, and also the resulting translated Protein!!! See example below. [Example shows the biomolecular flow in central dogma from DNA to RNA to Protein] Special note that all “T” were transcribed into “U” and that the 3-nt codon represents 1-AA.
Part 4: Prepare a Twist DNA Synthesis Order
4.2. Build Your DNA Insert Sequence
Link to the sequence (first attempt):
https://benchling.com/s/seq-92QKTmxOZ4NOBZloFYXH?m=slm-ih8RIVqVkxJpGYbdm50f
Link to corrected sequence:
https://benchling.com/s/seq-AKpYnuHnRmdf5XnJxSv8?m=slm-sqc6y4bFyGTTvcXYx3Q9
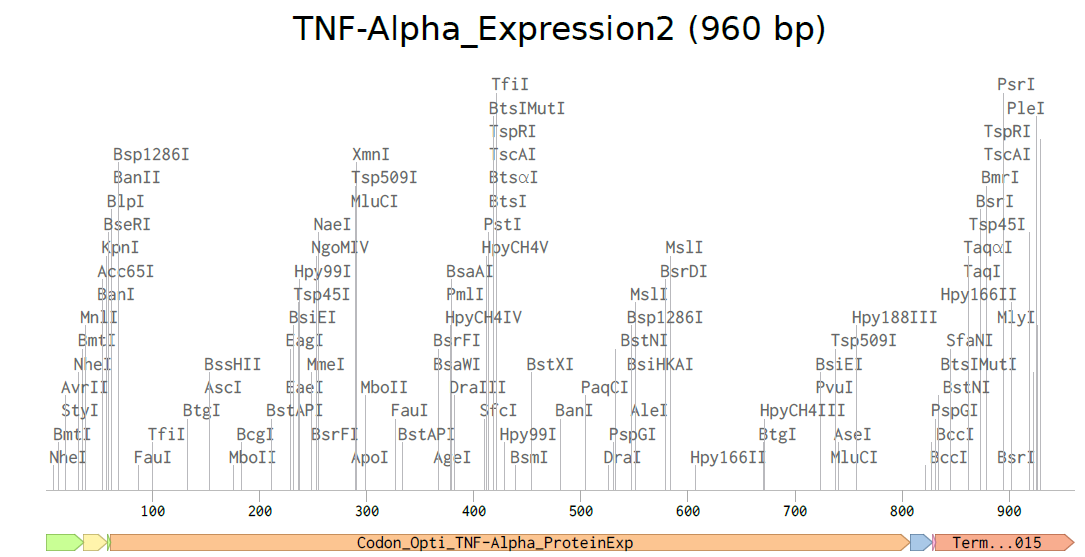

4.3-4.5. Building Expression Cassette and Plasmid
Plasmid with Expression Cassette
https://benchling.com/s/seq-dx10o3kwSJPyNLgmJDGo?m=slm-V5wHDO0G8ZxGTwWVp2A7
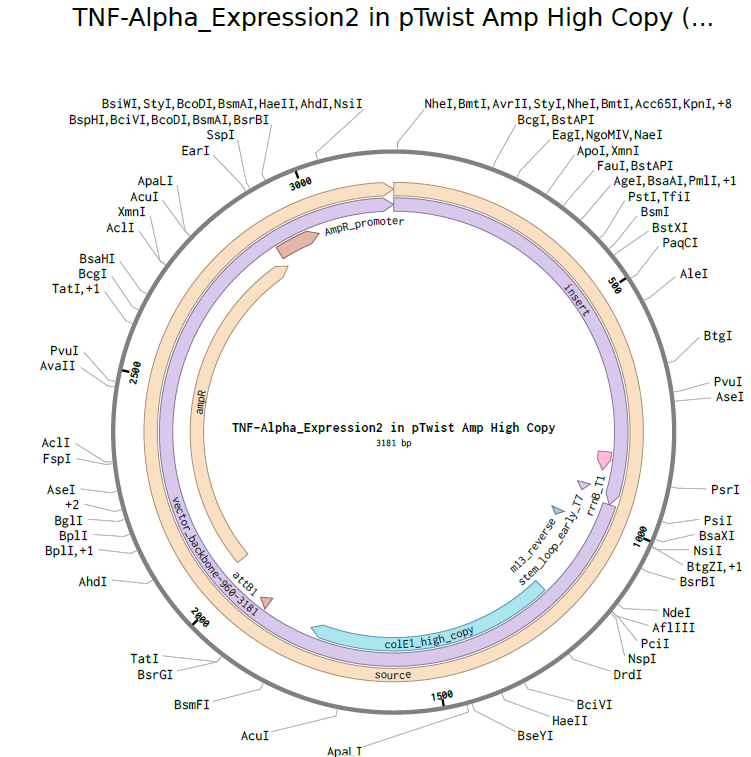

Part 5: DNA Read/Write/Edit
5.1 DNA Read
(i) What DNA would you want to sequence (e.g., read) and why? This could be DNA related to human health (e.g. genes related to disease research), environmental monitoring (e.g., sewage waste water, biodiversity analysis), and beyond (e.g. DNA data storage, biobank).
I would want to sequence Mycobacterium Tuberculosis DNA. I would like to focus on virulence‑associated loci such as phoR, mymA and the mce1, and lineage defining SNPs, such as rpoB, katG, inhA promoter, gyrA, embB.
To integrate with surveillance, I would potentially try to store drug resistance and mutation outputs from my detection bio-tool into a DNA-based archive. This could help build a long-term genomic repository.
(ii) In lecture, a variety of sequencing technologies were mentioned. What technology or technologies would you use to perform sequencing on your DNA and why? Also answer the following questions: 1. Is your method first-, second- or third-generation or other? How so? 2. What is your input? How do you prepare your input (e.g. fragmentation, adapter ligation, PCR)? List the essential steps. 3. What are the essential steps of your chosen sequencing technology, how does it decode the bases of your DNA sample (base calling)? 4. What is the output of your chosen sequencing technology?
To perform sequencing for the drug-resistant DNA, short-read sequencing is ideal for identifying the key resistance driving genes for profiling and analysis. In contrast, long-read sequencing (e.g. Oxford Nanopore) would make rapid detection, which is useful in high-burden regions, but has slightly lower accuracy. Therefore, short-read sequencing is ideal for identifying key resistance-driving genes for profiling and analysis (for e.g. using Illumina) [1]. It involves DNA extraction, fragmentation, adapter ligation, cluster amplification, and sequencing by synthesis, with base-calling software decoding the sequence from fluorescent signals. The output includes high-quality short reads, aligned sequences, and variant calls for resistance and lineage analysis. In contrast, long-read sequencing enables rapid detection in high-burden regions but has slightly lower accuracy and may require deeper coverage.
References
- The CRyPTIC Consortium and the 100,000 Genomes Project. Prediction of Susceptibility to First-Line Tuberculosis Drugs by DNA Sequencing. N Engl J Med. 2018;379:1403–1415. doi:10.1056/NEJMoa1800474.
5.2 DNA Read
(i) What DNA would you want to synthesize (e.g., write) and why? These could be individual genes, clusters of genes or genetic circuits, whole genomes, and beyond. As described in class thus far, applications could range from therapeutics and drug discovery (e.g., mRNA vaccines and therapies) to novel biomaterials (e.g. structural proteins), to sensors (e.g., genetic circuits for sensing and responding to inflammation, environmental stimuli, etc.), to art (DNA origamis). If possible, include the specific genetic sequence(s) of what you would like to synthesize! You will have the opportunity to actually have Twist synthesize these DNA constructs! :)
I would like to design a genetic circuit that could be integrated into a microbial chassis or a cell-free system, which would enable it to detect molecular signatures for key multi-drug /extra-drug-resistant tuberculosis and activates a fluorescent reporter when present in a sample. Examples of this have been seen in research that looks at how biosensors are used to detect heavy metal in water through recombinase-based logic gates [1]. Such CRISPR‑based detection systems can be programmed with guides targeting lineage‑specific SNPs (e.g., Beijing/East Asian, Indo-American) [2] alongside resistance mutations so that the circuit only activates a fluorescent reporter when both types of signatures are present. Potentially, CRISPR‑Cas12/13 coupled with allele‑specific amplification can discriminate single‑base changes for lineage and resistance detection with high specificity. There is also a possibility of integrating all of this into a microfluidic biosensor, enabling automated, low-volume, rapid, and multiplexed detection suitable for environmental and point-of-care surveillance [3].
(ii) In lecture, a variety of sequencing technologies were mentioned. What technology or technologies would you use to perform sequencing on your DNA and why? Also answer the following questions:
- Is your method first-, second- or third-generation or other? How so?
- What is your input? How do you prepare your input (e.g. fragmentation, adapter ligation, PCR)? List the essential steps.
- What are the essential steps of your chosen sequencing technology, how does it decode the bases of your DNA sample (base calling)?
- What is the output of your chosen sequencing technology?
For this synthesis, I would use synthetic DNA platforms and include CRISPR guide sequences, promoters, and fluorescent reporter proteins. These technologies would allow for quick prototyping, flexibility with design and would allow for automated printers to synthesize sequences up to multiple kilobases accurately.
Essential steps would include: full sequence of nucleotides and CRISPR guides, promoters and reporter proteins; setting the oligonucleotide assembly, this includes making assemblies of short oligos through PCR or ligation. These would need to be further tested and validated to ensure proper functioning of the circuit.
Limitations include, time, fixing errors, and scaling the device. These large constructs and may take time due to the complexity associated with multiple variants.
References
Mathur S, Singh D, Ranjan R. Genetic circuits in microbial biosensors for heavy metal detection in soil and water. Biochem Biophys Res Commun. 2023 Apr 16;652:131–137. doi:10.1016/j.bbrc.2023.02.031.
Napier, G., Campino, S., Merid, Y. et al. Robust barcoding and identification of Mycobacterium tuberculosis lineages for epidemiological and clinical studies. BMC Genome Med 12, 114 (2020). https://doi.org/10.1186/s13073-020-00817-3
Didarian R, Azar MT. Microfluidic biosensors: revolutionizing detection in DNA analysis, cellular analysis, and pathogen detection. Biomed Microdevices. 2025;27:10. doi:10.1007/s10544-025-00741-6.
5.3 DNA Edit
(i) What DNA would you want to edit and why? In class, George shared a variety of ways to edit the genes and genomes of humans and other organisms. Such DNA editing technologies have profound implications for human health, development, and even human longevity and human augmentation. DNA editing is also already commonly leveraged for flora and fauna, for example in nature conservation efforts, (animal/plant restoration, de-extinction), or in agriculture (e.g. plant breeding, nitrogen fixation). What kinds of edits might you want to make to DNA (e.g., human genomes and beyond) and why?
For editing, I would use CRISPR-Cas systems to introduce lineage specific SNPs and resistant mutations into safe mycobacterial strains or cell-free systems [1]. This allow me to test the genetic circuit, validate the CRISPR guides, and generate controls for MDR-TB detection.
(ii) What technology or technologies would you use to perform these DNA edits and why? Also answer the following questions:
- How does your technology of choice edit DNA? What are the essential steps?
- What preparation do you need to do (e.g. design steps) and what is the input (e.g. DNA template, enzymes, plasmids, primers, guides, cells) for the editing?
- What are the limitations of your editing methods (if any) in terms of efficiency or precision?
Essential steps would include: designing guide RNAs to target SNPs/loci related to drug resistance; integrating of editing components into cells or a cell-free platform; functional testing to ensure sequences properly activate the fluorescent reporter protein within the circuit.
Preparation would require designing the guide RNAs and providing either a cell-free system or microbial framework as the host.
Limitations include: possible off-target edits; increased complexity when introducing multiple edits or larger constructs, which can affect throughput and precision.
References
- Molla KA, Yang Y. CRISPR/Cas mediated base editing: technical considerations and practical applications. Trends Biotechnol. 2019 Oct;37(10):1121–1142. doi:10.1016/j.tibtech.2019.03.008. Review of CRISPR base editing systems and how they introduce precise nucleotide changes without double strand breaks.
title: ‘Week 3 HW: Principles and Practices’ weight: 30
Homework Week 3
Post-Lab Questions
One of the great parts about having an automated robot is being able to precisely mix, deposit, and run reactions without much intervention, and design and deploy experiments remotely. For this week, we’d like for you to do the following:
1. Find and describe a published paper that utilizes the Opentrons or an automation tool to achieve novel biological applications.
2. Write a description about what you intend to do with automation tools for your final project. You may include example pseudocode, Python scripts, 3D printed holders, a plan for how to use Ginkgo Nebula, and more. You may reference this week’s recitation slide deck for lab automation details.
While your description/project idea doesn’t need to be set in stone, we would like to see core details of what you would automate. This is due at the start of lecture and does not need to be tested on the Opentrons yet.
(Answer 1)
Paper: An Automated Versatile Diagnostic Workflow for Infectious Disease Detection in Low-Resource Settings
Miren Urrutia Iturritza, Phuthumani Mlotshwa, Jesper Gantelius, Tobias Alfvén, Edmund Loh, Jens Karlsson, Chris Hadjineophytou, Krzysztof Langer, Konstantinos Mitsakakis, Aman Russom, Håkan N. Jönsson, Giulia Gaudenzi
https://doi.org/10.3390/mi15060708
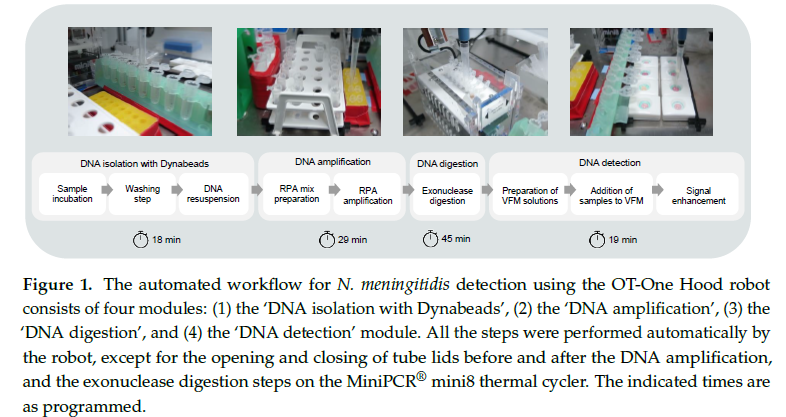
This paper describes how researchers built an automated diagnostic workflow for detection of infectious diseases in low-resource settings [1]. Specifically, they tested for Neisseria meningitidis; a gram-negative bacterium that cause serious meningitis and blood infections in humans.
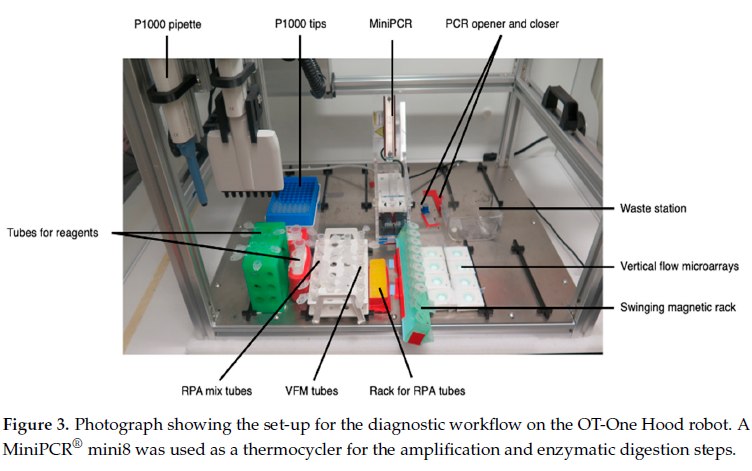
For their workflow, they used Opentrons OT-One-S Hood. This is an open-source liquid handling robot, which can be bought at a relatively low cost. The researchers wrote custom software developed at SciLifeLab Nanobiotechnology division [2] to create scripts for their workflow.
Materials and reagents were organized onto the OT-One-S Hood robot, with racks and tubes with primers, buffers, and enzymes, the MiniPCR® mini8 thermal cycler, magnetic bead racks, waste containers, and microarray holders, to analyze Neisseria meningitidis DNA in both clinical and spiked samples. “Clinical” samples refere to specimens collected from individuals, where “spiked” samples were lab prepared samples where a known amount of Neisseria meningitidis DNA.
The robot then performs all the necessary pipetting steps, RNA amplication of ctrA gene (as its conserved, species-specific gene essential for capsule formation, making it a reliable marker [3]), enzymatic digestion, and deposition onto paper-based microarrays. The only manual steps were the opening and closing of tube lids before and after the DNA amplification, and the exonuclease digestion steps on the MiniPCR® mini8 thermal cycler [1].
The study showed that, automated liquid handling can detect Neisseria meningitidis in low-resource settings, though accuracy and reproducibility were not fully validated.




References
Urrutia Iturritza M, Mlotshwa P, Gantelius J, Alfvén T, Loh E, Karlsson J, Hadjineophytou C, Langer K, Mitsakakis K, Russom A, et al. An automated versatile diagnostic workflow for infectious disease detection in low-resource settings. Micromachines. 2024;15(6):708. doi:10.3390/mi15060708.
Langer K, Joensson HN. Rapid production and recovery of cell spheroids by automated droplet microfluidics. SLAS Technol. 2020;25:111–122.
Rivas L, Reuterswärd P, Rasti R, Herrmann B, Mårtensson A, Alfvén T, Gantelius J, Andersson-Svahn H. A vertical flow paper-microarray assay with isothermal DNA amplification for detection of Neisseria meningitidis. Talanta. 2018;183:192–200.
(Answer 2)
For the automation of my project, I plan to use automation tools to develop and test a CRISPR-based biosensor that would be capable of detecting multi-drug-resistant tuberculosis (MDR-TB) signatures. This workflow would involve high-throughput liquid handling and cell-free protein synthesis. Possible steps would include:
(i) Module setup: This would include arranging reagents, tip racks, thermal cyclers, magnetic bead racks, and microarray holders on an Opentrons OT-2 deck [1]. This would be supplemented by temperature modules for incubation and heater-shaker modules for mixing and precise reaction control
(ii) Automated reaction setup: The robot will then perform pipetting of cell-free lysate, DNA templates, CRISPR guides, and cofactors into 96- or 384-well plate. Then multiple combinations of lineage-specific SNP guides and resistance mutation guides will be tested to evaluate ‘AND-gate logic’.
(iii) Incubation: External devices like a plate reader or miniPCR thermal cycler amplification will be loaded. Then Python scripts will be used to control timing, mixtures, and incubation periods.
(iv) Signal detection and analysis: Fluorescent outputs will be measure using devices such as Spark or PHERAstar FSX for high-throughput plate analysis [1]. This will be a measure of change in fluorescence colour which would indicate successful target detection and amplification.
(v) Microfluidic integration (if possible): If possible, will look to integrate 3D printed holders for small microfluidic chips. These can serve as small test cartridges for running multiple tests at once while minimizing manual handling and contamination risk in low-resource settings.
References
[1] Course Recitation Slides. Lab Automation Overview. Course presentation, [institution or course name if known]; Year [cited 2026 Feb 23]. Available from: https://docs.google.com/presentation/d/e/2PACX-1vQc3zo7Z0b6HK7YeC56p_n2RbHNjUHh1HI66DH0cHbFk0db1HlbF7gILE__NCvhUiYMjIGSOHwHPv2_/pub?start=false&loop=false&delayms=3000#slide=id.g2b9b763dcde_1_131