Week 2 HW :DNA Read, Write and Edit
Part 1: Benchling & In-silico Gel Art
- Make a free account at benchling.com
- Import the Lambda DNA.
- Simulate Restriction Enzyme Digestion
- Design using Ronan’s website: Cactus Digest. I found this tool very useful. For the longest time I was just trying to get a good pattern randomly. Then i realised I can edit individual columns. With some more digging it became clear that I need to find restriction enzymes that slice the DNA in the bands that appear like something on the digest. So for the lane 5 I used EcoRV and BanHI to get the nice trunk of the Cactus design. For lane 4 and 6 i used SacI since it only makes two bands. And for lane 3 and & I used EcoRI and BamHI to make the 2 stems of the cactus. A little throwback to where I am from (Rajasthan).
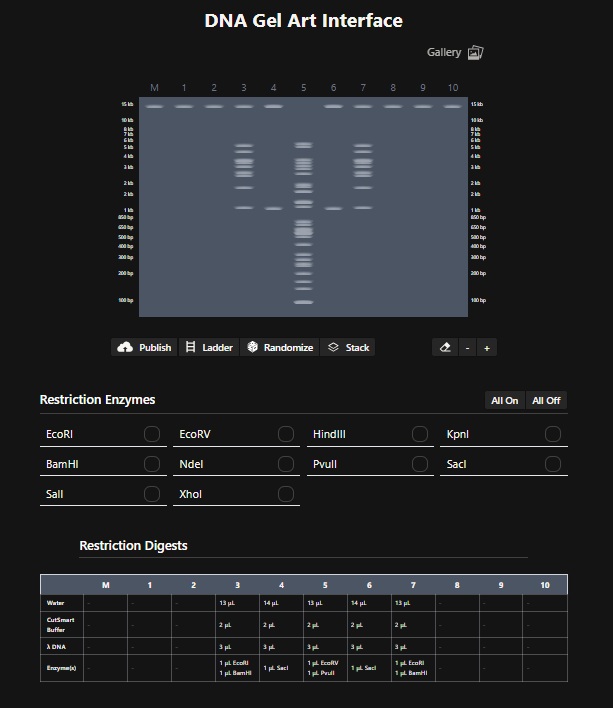

- Create a pattern/image in Benchling- I then recreated ronans website design by performing the digest on the lamda phage genome downloaded from GenBank in Benchling using the digest enzymes that ronans website described. The following is a screenshot of the Virtual Digest in Benchling
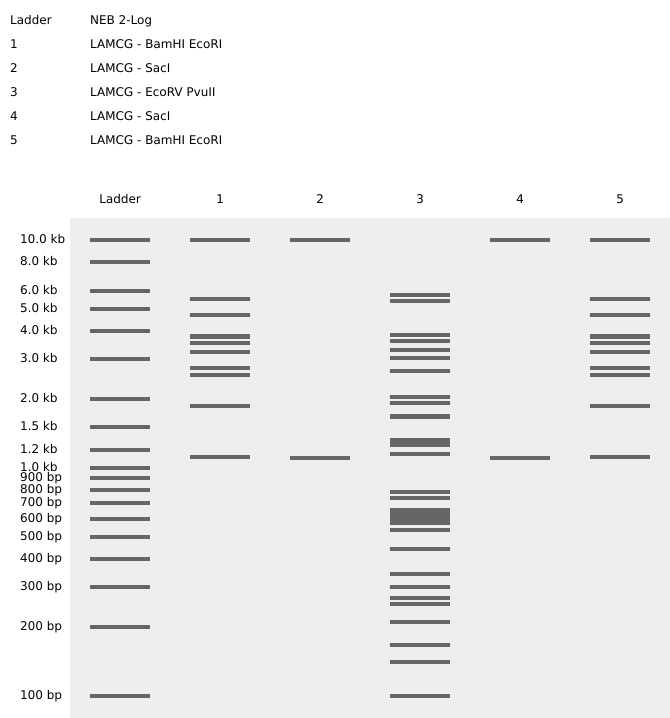


Part 2: No remote lab access yet! 🤞
Part 3: DNA Design Challenge
3.1 Protein Choice:
Since I am working on building a smart hydrogel I did some literature review and found that Elastin Like Polypeptides (ELPs) are good candidates. ELPs are recombinant protein polymers inspired by the natural protein Elastin, found in the extra cellular matrix, which is responsible for the flexibility of our bodies. ELPs are thermally responsive biopolymers and capable of having customizable properties. ELPs undergo something called an inverse temperature transition. This means that when heated, normal polymers dissolve but ELPs do the opposite. So ELPs stay soluble below a certain temperature and above a transition temperature (Tt) they undergo hydrophobic collapse and aggregate. ELPs interestingly are just repeat pentapeptides with amino acidv(AA) sequence Val-Pro-Gly-X-Gly (VPGXG)n where X can be any canonical amino acid except proline. This interchangebaility of X gives ELPs their customizability and such engineered ELPs exhibit excellent potential as customizable platforms for drug delivery and tissue repair.
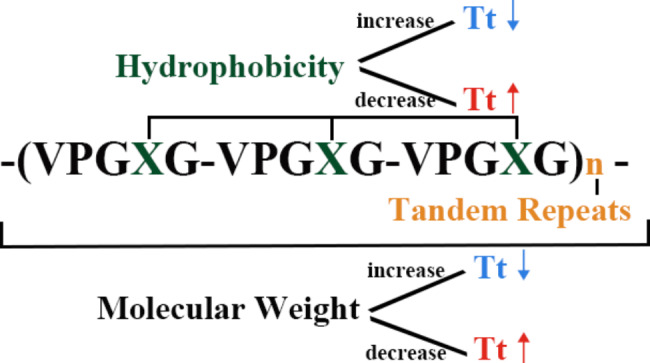

structure of ELPs and its impact on its own Tt (Guo Y, Liu S, Jing D, Liu N, Luo X. The construction of elastin-like polypeptides and their applications in drug delivery system and tissue repair. J Nanobiotechnology. 2023 Nov 11;21(1):418. doi: 10.1186/s12951-023-02184-8. PMID: 37951928; PMCID: PMC10638729.)
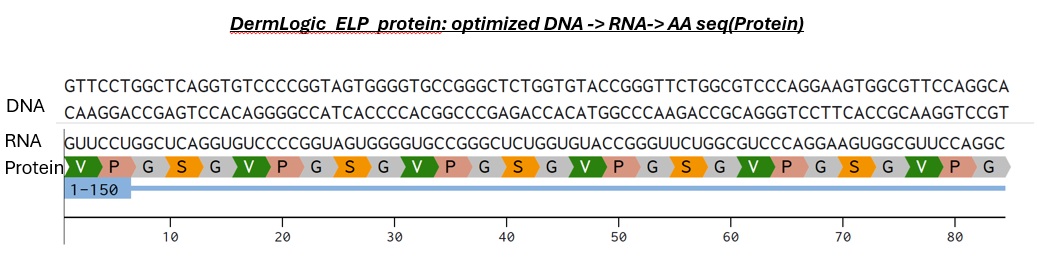
For this weeks assignment I designed a short ELP consisting of only 10 repeats with X= Serine to creat a moderately hydrophilic scaffold:. So the AA sequence will be:
VPGSGVPGSGVPGSGVPGSGVPGSGVPGSGVPGSGVPGSGVPGSGVPGSG
3.2 Reverse Translate: Protein (AA) sequence to DNA (NA) sequence:
For this part I used reverse translation tool online for this as suggested in the homework instructions and got this as a result
“>reverse translation of Untitled to a 150 base sequence of most likely codons.
gtgccgggcagcggcgtgccgggcagcggcgtgccgggcagcggcgtgccgggcagcggc gtgccgggcagcggcgtgccgggcagcggcgtgccgggcagcggcgtgccgggcagcggc gtgccgggcagcggcgtgccgggcagcggc
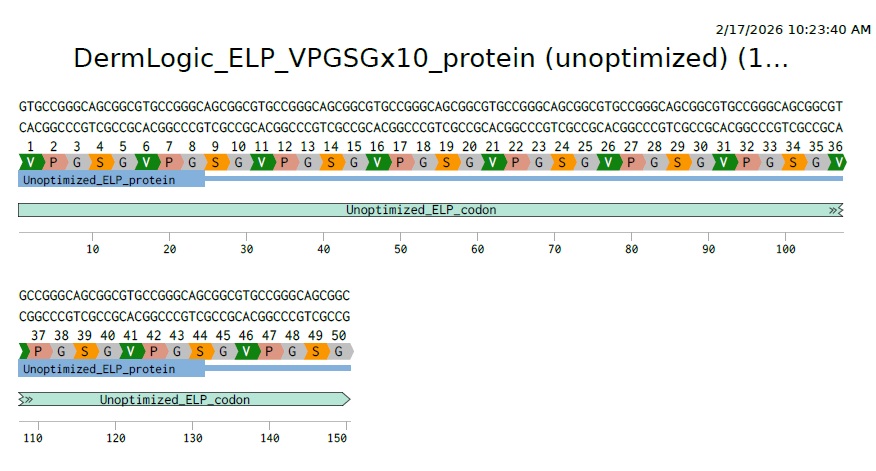

3.3 Codon optimization:
- Why optimize?
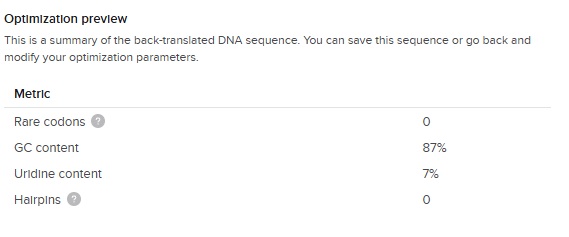

Now the problem here is that its a highly repetitive DNA with repeats of GTG CCG GGC AGC GGC repeated 10 times. Its correct biologically but as mentioned in Dr Prousts lecture this is what you want to avoid for synthesis stability. So we need to figure out a way to make the DNA sequence less repetative eventhough the AA are repetative. We can do this because each amino acid has more than one codon that can code for it. So lets find the options for each of our AAs. Here they are:
- Valine: GTT, GTC, GTA, GTG
- Proline: CCT, CCC, CCA, CCG
- Glycine: GGT, GGC, GGA, GGG
- Serine: AGT, AGC, TCT, TCC, TCA, TCG
Now we need to find a way to use these options to create a new DNA sequence thats less repetative and more synthesis friendly.
Codon optimization on benchling
On Benchling I uploaded the 10 repeats of my AA sequence (
VPGSGVPGSGVPGSGVPGSGVPGSGVPGSGVPGSGVPGSGVPGSGVPGSG). Then selecting the whole sequence choose back translate. Choosing E.Coli (O157:H7 EDL933). Then Match codon usage to target organism method.
The output of the codon optimization gave the following optimization preview results:
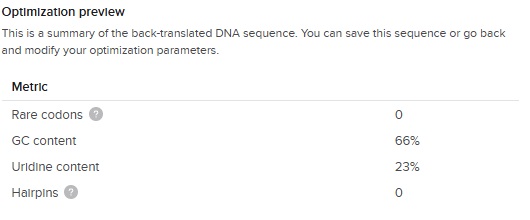

The GC content (66 %) is slightly higher than the E.Coli average (around 50%) but great considering our first DNA sequence was 87 % !
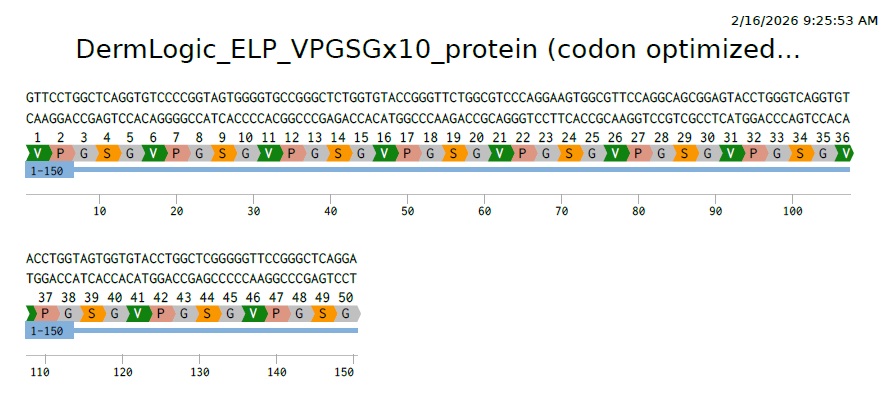

So this is our final optimized 150 bp sequence. It translates to VPGSG 10 times. It has clearly become less repetative.
3.4. You have a sequence! Now what?
What technologies could be used to produce this protein from your DNA? Describe in your words the DNA sequence can be transcribed and translated into your protein. You may describe either cell-dependent or cell-free methods, or both.
Production Technology: Ideally, I would like to produce this ELP using a cell-free protein synthesis (CFPS) system. Cell-free system will allow me to rapidly prototype different ELP lengths without the time-consuming steps of cloning and cell-culture. However, since I am learning the foundational workflows of synthetic biology and tools, I have chosen to codon optimize and design the sequence in E.Coli.
Why this works: Since the common cell free systems (such as TX-TL) are actually made from e.coli extracts, optimizing the codon for e.coli is a robust strategy that will allow this DNA to be used in both living cells and cell-free synthesis.
Mechanism (Central Dogma): To produce the protein the DNA is transcribed by RNA Polymerase (recognizing the J23106 constitutive promoter in my design in Part5 below) into messenger RNA. This single stranded mRNA will then be recognized by the ribozomes present in e.coli at the ribozome binding site and thus translating the codon-optimized sequence we made earlier. The ribozome will read the codons three neucleotides at a time and then recruit tRNA with matching amino acids like Valine-Proline etc (that we want and we optimized in benchling for e.coli so they have these tRNAs present and there is no rare codons) and links them together until reaching a stop codon in the mRNA (the T1 terminator) and releasing the ELP polymer. Quite magical how this is identical in all living cells.
3.5 How does it work in nature/biological systems?
- Describe how a single gene codes for multiple proteins at the transcriptional level.
- Despite the universality there are important variations between various organism to control this process of DNA code becoming protein. In eukaryotes A single gene can code for multiple proteins primarily through Alternative Splicing. During this process, the pre-mRNA (containing both coding and non-coding regions) is produced from transcription. Then something called a spliceosome splices the introns and stitches together the exons. The alteration crucially comes from the spliceosome which doesn’t always stitch together in the same order. It can skip certain exons or include others depending on the cells need. This results in distinct mRNA transcripts from the same DNA template which are then translated to unique protein isoforms with different structures and thus functions.
This is what it would look like to have a gene that codes for our ELP polymer with introns and exons in Eukaryotes.
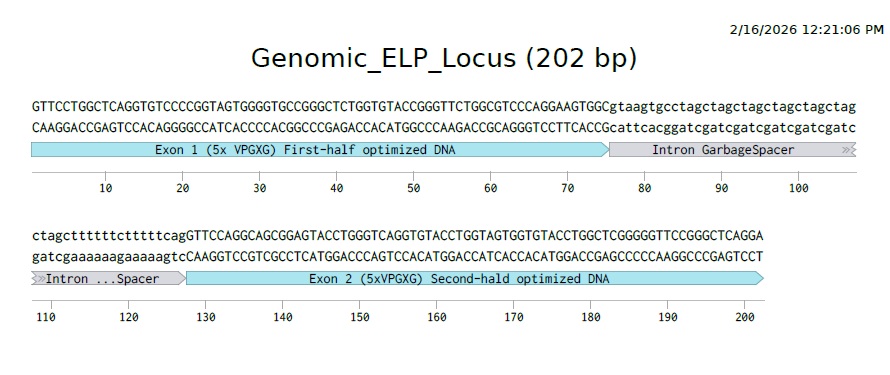


Prokaryotes like E.coli do not use splicing, they often use Polycistronic mRNA (Operons). In this system, a single promoter drives the transcription of multiple distinct genes arranged in a row. The resulting long mRNA strand contains multiple Ribosome Binding Sites (RBS), allowing ribosomes to translate each gene into a seperate protein independently. This allows bacteria to turn on entire metabolic pathways with a single switch, rather than cutting and pasting RNA like eukaryotes do.
I personally am fascinated with the eukaryotic spliceosome and wonder what it would be like to engineer something as complicated and sofisticated.
Part 4: Prepare a Twist DNA Synthesis Order
4.1: Create Twist and Benchling account
4.2: Build your DNA Insert Sequence
Using the varius parts of the expression casette and the optimized codon we get this (https://benchling.com/s/seq-hjqR0qesr21GIZrfs2Ua?m=slm-glVH1Ltmva9AxlUc2JFg)
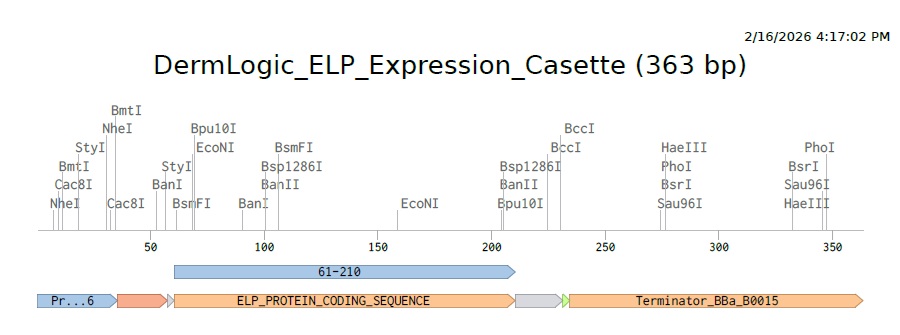


4.3 - 4.6: Final Plasmid from Twist with our ELP insert.
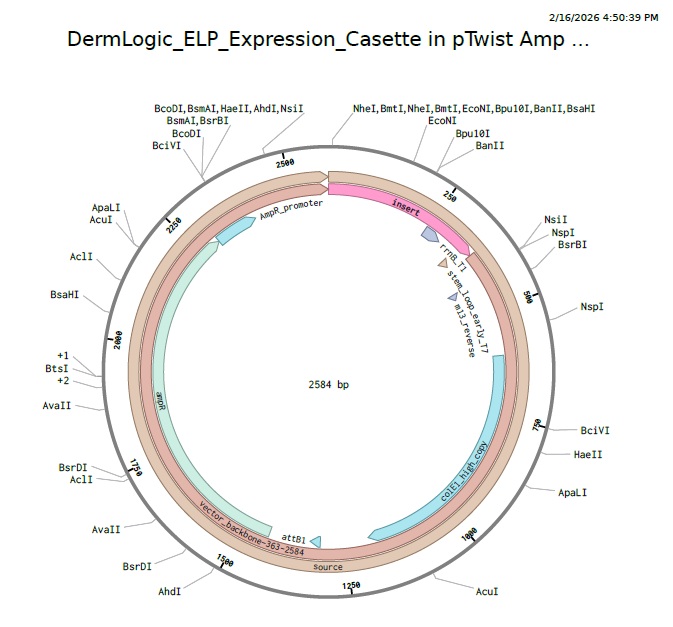

Also our complexity score in twist came out to be standard that means our codon optimization worked and the plasmid is constructable even with our initial repeat codon sequence with high GC content.

Part 5: DNA Read/Write/Edit:
5.1 DNA Read:
- (i) What to sequence:
I would like to read the DNA of a Beaver’s gut microbiome and see how it manages to survive eating just trees. I love going camping in the beautiful parks of Ontario such as Algonquin park and always see tons of beavers on my trips. And I am fascinated at the uniqueness of beavers and how they are really architects of their environment much like us humans. And a big part of that is their unique microbiome that lets them survive on something so abundant (trees) and their amazing skills to live under water in the winters. ALso their abilities to hibernate and how thats beneficial to them. Sequencing them could reveal novel enzymes for degrading tough plant materials, or even traits that support their survival under ice during canadian winters.
- (ii) DNA Sequencing approach:
Since the beaver gut probably contains thousands of unknown bacteria and fungi working together to break down wood, I would need to sequence the entire soup of DNA. So Metagenomic sequencing would be a good approach using the Oxford Nanopore Tech (ONT). Nanopore sequencers like MiniON are portable which would be ideal to take with me to algonquin park and sample fresh sample from the field!
This is very third generation. To be more precise it is Single Molecule Real-TIme (SMRT) sequencing. Since Nanopore unlike 2nd gen methods (that required PCR amplification) can read native, single molecule DNA directly in real time without amplification. This allows for extremely long reads critical for assembling complex metagenomes.
Wet Steps I will need a field ready extraction kit like once from Zymo Research to lyse the hardy fungal/bacterial cells. I will also need to ligate motor proteins and sequencing adapters to the ends of DNA fragments which will lead the DNA into the nanopore.
How it Decodes The Nanopore is a protein pore through which current is passed and each combination of neucleotides in the DNA inside the pore creates a unique squiggle that Neural Netorks translate into a nucelotide seq.
Output Primary output is usually a FASTQ file which is the sequence data with quality scores. For the gut soup project we will have to take the collection of millions of reads and compare it with the Metagenome-Assembeled Genomes to identify our wood-degrading microbes of interest.
5.2 DNA Write:
(i) What to synthesize: Thinking about the DermLogic concept all week I think beyond simple ELPs, I would like to synthesize a “Living Bandage” gene circuit. This could be a genetic circuit designed using skin bacteria like S.epidermis that can sense inflammation markers and respond by synthesizing and secreting healing hydrogel matrix directly into the wound.
(ii) I like the concept of De Novo DNA synthesis using polymerase nucleotide conjugates shared by Prof Joe Jacobson. Its a fast and non-toxic compared to the traditional Phosphoramidite Chemistry. Also its capable of long fragment synthesis which is the future of potentially synthesizing several kilobases without need for extensive assembly. But for the gene fragments ad vectors needed for this type of live bandage I would use the Oligo synthesis using Phosphoramidite chemisry used by companies like Twist Biosciences.
5.3 DNA Edit:
(i) What to edit: I would like to edit the connective tissue cells (fibroblasts) in older adults to restore their ability to produce functional elastin. Since reading about the role of elastin in the human body I was wondering if we could somehow mimic the natural elastin in the extra cellular matrix of patients with lack of elastin causing various dieseases and locally edit the ECM to reverse some of the elastin related aging in older adults. Such ELPs could someday help an old person heal their weak broken bones that are still “fixed” by doctors using screws and foreign metal parts and instead hope for a future where we could perhaps edit the local stem cells to secrete high performance ELPs that regenerate the tissue naturally, restoring the elasticity and strength.
(ii) Technologies
Design: Since we are dealing with humans we wouldnt want to use standard CRISPR-Cas9 to cut ds DNA but rather use something advanced in CRISPR technologies like Prime editing. It uses a nickase (a broken Cas9 that can only cut one strand) fused to a reverse transcriptase to find a specific site using pegRNA (guide RNA for the system) in the fibroblast genome and directly write a corrected sequence for elastin without causing any dangerous ds breaks.
Inputs: Firstly we will need the fusion protein delivered as a plasmid or mRNA encoding the nCas-9RT fusion. Thenn we will make the custom guide pegRNA containing the target spacer (DNA site in the fibroblast genome for elastin). Lipid nanoparticle delivery system can be useful for delivery into the fibroblast cell.
Limitations: Its less efficient (20-50% successful cells edited) than standard CRISPR-Cas9 (80%) Designing pegRNA is harder than standard gRNA
*Thanks for reading!*