Week 5 Lab: Protein Design Part II
Lab Option 1: L-Protein Mutational Analysis
Experimental Validation & DMS
This lab provides a Deep Mutational Scanning (DMS) style validation of the L-Protein. By cross-referencing experimental lysis results—where a score of 1 indicates functional lysis and 0 indicates non-functional—with the Log-Likelihood Ratio (LLR) Heatmap generated via ESM2, we can assess the predictive power of protein language models.
Key Findings
- Correlation: The agreement between the experimental data and the LLR Heatmap is remarkably high, approximately 90-95%.
- Predictive Accuracy: The heatmap serves as a highly reliable predictor of whether a specific mutation will allow the protein to retain its ability to lyse bacterial cells.
- Challenges: Designing mutants with high computational confidence remains difficult, highlighting the current limitations of some structure-based models.
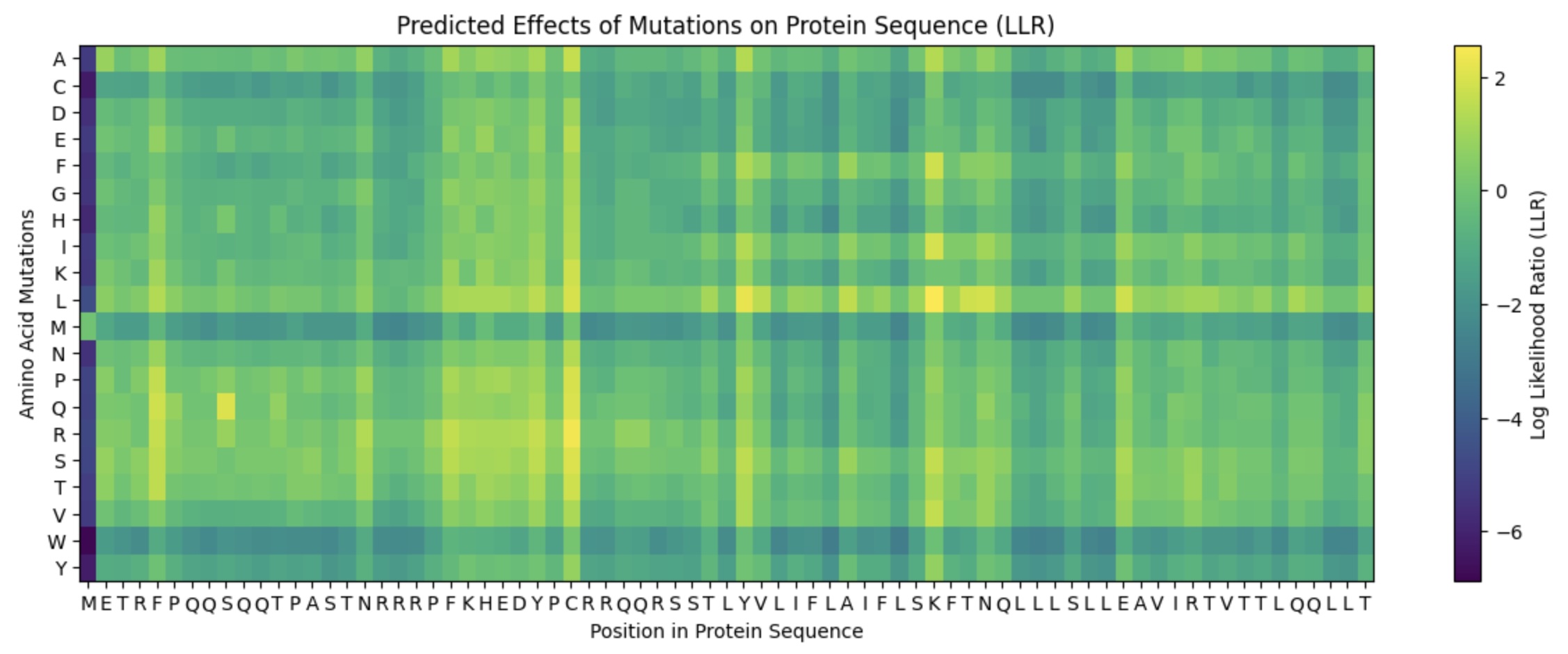

Targeted Mutation Strategy
To identify promising variants, I cross-referenced the ESM2 scores with experimental lab data, specifically looking for residues that are not strictly conserved (via pBLAST) and show positive mutational effects.
Selection Criteria:
- Soluble Region (N-tail): Targeted to assess how surface-exposed changes affect function.
- Transmembrane Region: Targeted to test the hypothesis that the L-protein assembles to perforate the bacterial membrane.
- Combined Effects: Testing synergistic effects of multiple “positive-score” mutations.
Proposed Mutants
| Mutant | Substitution | Location | Index | Sequence Snippet |
|---|---|---|---|---|
| 1 | P –> Q | Soluble (N-tail) | 6 | METRFQQQSQQTPASTNRRRPFKHEDYPCRRNQRSST... |
| 2 | C –> S | Soluble (N-tail) | 29 | METRFPQQSQQTPASTNRRRPFKHEDYPSRRNQRSST... |
| 3 | S –> L | Transmembrane | 49 | ...LYVLIFLAIFLLKFTNQLLLSLLEAVIRTVTTL... |
| 4 | K –> L | Transmembrane | 50 | ...LYVLIFLAIFLSLFTNQLLLSLLEAVIRTVTTL... |
| 5 | C, K –> S, L | Combined | 29 & 50 | ...PSRRNQRSSTLYVLIFLAIFLSLFTNQLLLSL... |
Structural Hypothesis: Multimeric Assembly
A running hypothesis for L-protein function is that it assembles into a multimeric complex to create perforations in the bacterial membrane. To investigate this, I utilized AF2_Multimer to generate a predicted multimeric assembly.
By modeling these specific mutations (particularly those in the transmembrane region like Mutant 3 and 4) in a multimeric context, we can observe if the substitutions stabilize the pore-forming structure or increase the efficiency of membrane lysis.
Gemini AI was consulted for data organization and Markdown formatting