Week 6 Lab: Gibson Assembly
Day 1: Preparation of DNA Fragments
Background
In this two-day lab, we modified the color-generating chromophore of the purple Acropora millepora chromoprotein (amilCP) to create a variety of orange, pink, and blue mutants.
We performed two sets of Polymerase Chain Reactions (PCR) to prepare for a Gibson Assembly. The insert PCR region spans the 24 base pairs before the chromophore to just beyond the gene transcription terminator. The forward primer includes an intentional mismatch for site-directed mutagenesis of the mUAV DNA plasmid. These mutants were then expressed in chemically competent E. coli cells.
PCR Reaction Setup
We prepared a backbone reaction alongside four color-specific reactions: Blue, Light Pink, Magenta, and Orange.
1. Backbone DNA Fragment
Primers: Backbone Fwd and Backbone Rev
| Reagent | Stock Conc. | Desired Conc. | Volume (µL) |
|---|---|---|---|
| Template mUAV Plasmid | 38.5 ng/µL | 20 ng | 0.8 |
| Backbone Forward Primer | 5 µM | 0.5 µM | 2.5 |
| Backbone Reverse Primer | 5 µM | 0.5 µM | 2.5 |
| Phusion HF PCR Mix | 2X | 1X | 12.5 |
| Nuclease-free water | — | — | 6.8 |
| Total Volume | — | — | 25.0 |
2. Color DNA Fragments
Primers: Color Fwd and Color Rev
| Reagent | Stock Conc. | Desired Conc. | Volume (µL) |
|---|---|---|---|
| Template mUAV Plasmid | 38.5 ng/µL | 20 ng | 0.8 |
| Color Forward Primer | 5 µM | 0.5 µM | 2.5 |
| Color Reverse Primer | 5 µM | 0.5 µM | 2.5 |
| Phusion HF PCR Mix | 2X | 1X | 12.5 |
| Nuclease-free water | — | — | 6.8 |
| Total Volume | — | — | 25.0 |
After mixing, the tubes were placed in the thermocyclers. The backbone was run on one specialized program while the color mutations were run on another.

Note: DpnI digest was skipped as our reactions did not contain methylated DNA.
Purification & Analysis
We purified the PCR products using the Zymo DNA Clean & Concentrator kit.
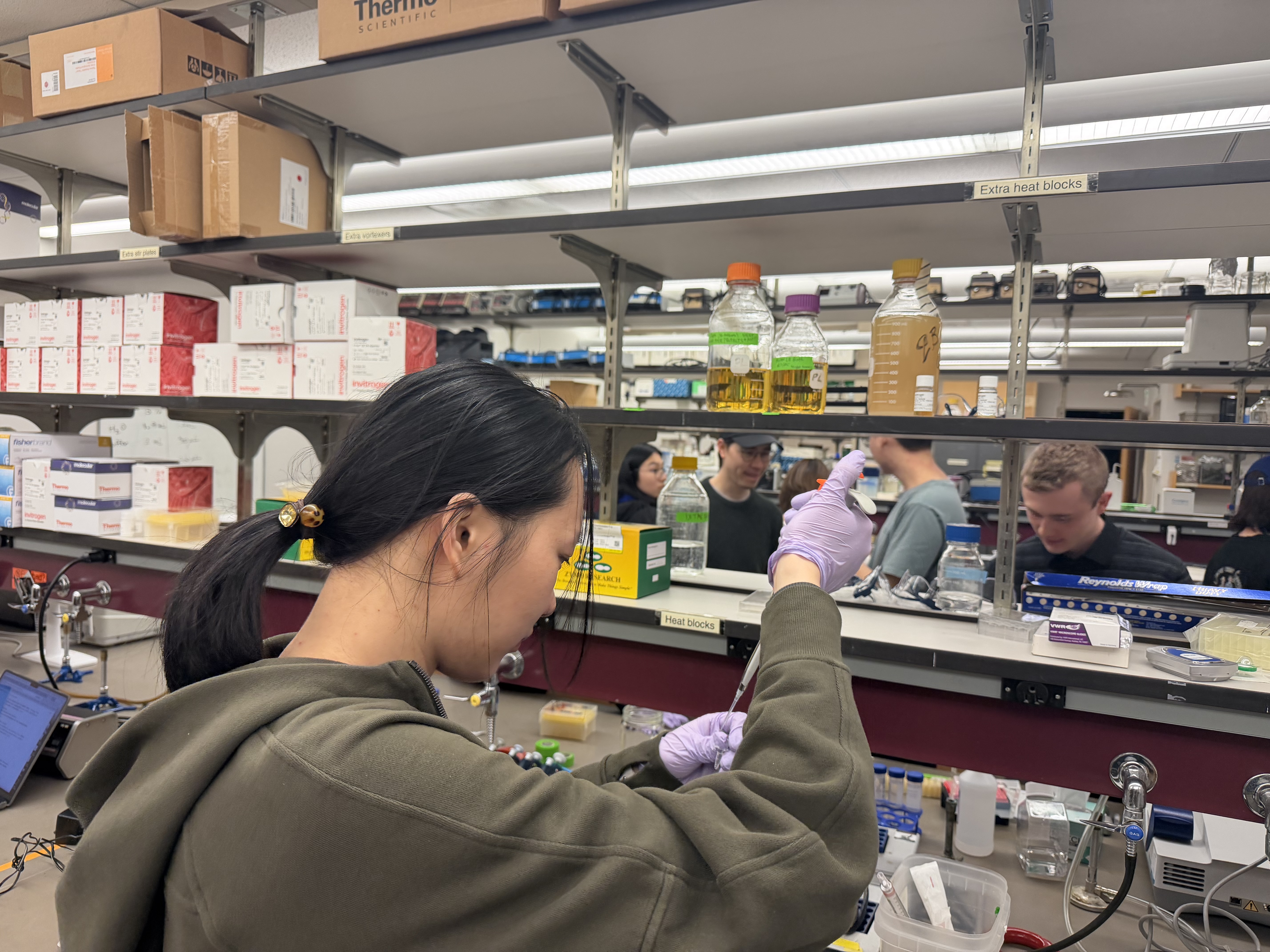


We utilized Zymo-Spin columns, performed two washes, and eluted the DNA for storage. The listed protocol was for 50µL of PCR product. We made 25µL total in our PCR reactions, so we used 20µL of PCR product (saved 5µL) in the purification process and scaled the other volumes to equal proportion. Gel electrophoresis was performed to verify the amplification.


Gel Results: We had a 1kb ladder furthest left, and Lane 1 contains the native plasmid. Lanes 2–5 show the expected amplified fragments for the Gibson Assembly, all roughly around 650bp. Our gel results were very convincing, we didn’t see formation of primer dimers or miscellaneous bands indiciating reduced PCR efficacy and polymerase fideltiy. Each sample contained a strong band at the expected bp length. Samples were then stored in fridge until the next lab day.
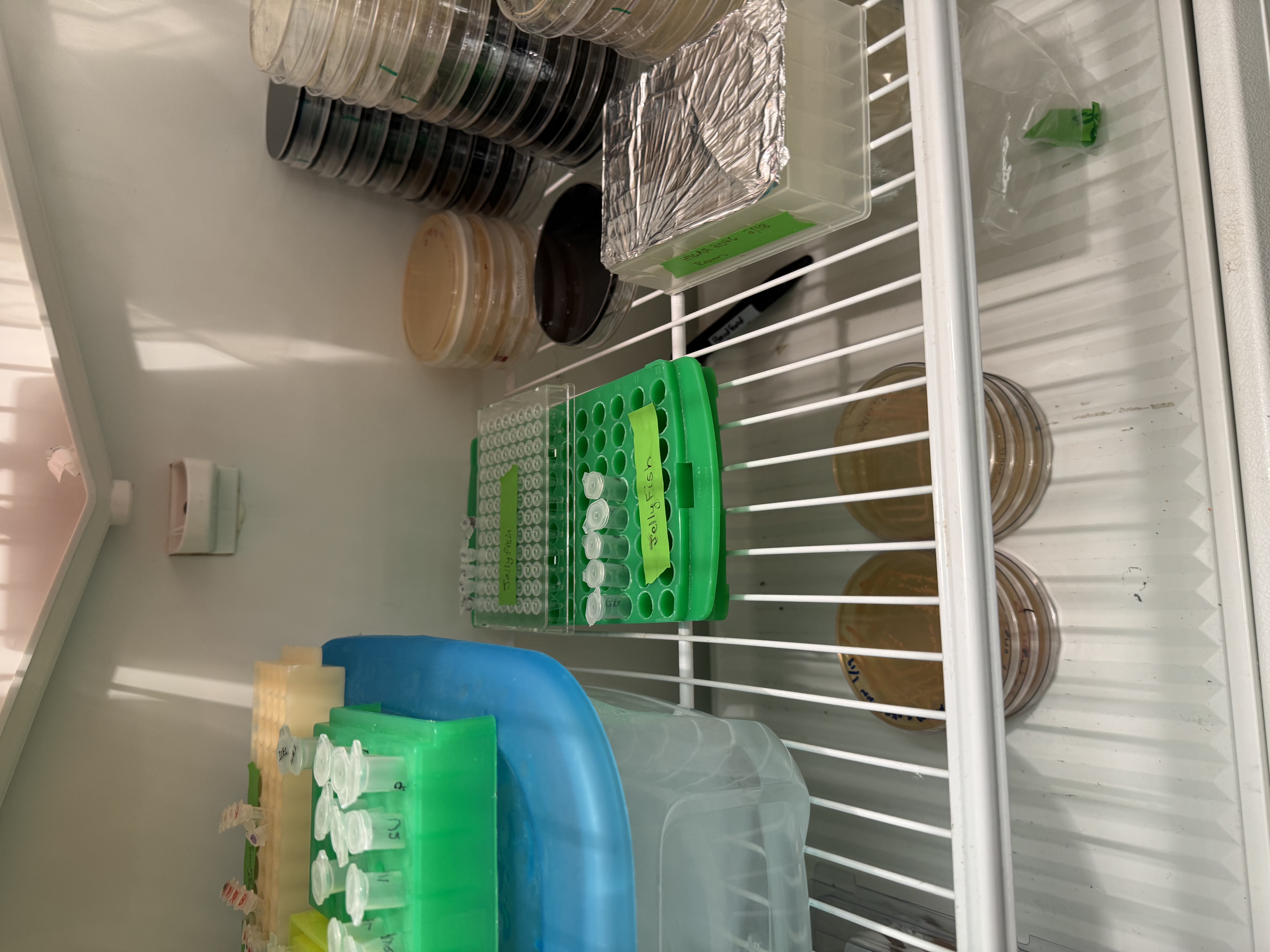


Day 2: Assembly & Transformation
Gibson Assembly
We diverged slightly from the standard protocol by utilizing unpurified PCR products for the assembly. This decision was made for a couple of reasons. We realized we had very small samples of each color in both the purified and unpurified results, effectively forcing us to chose between assaying the concentration via nano-drop or assembling the reaction with our theoretical maxium concentration. Additionally, other groups were reporting very low (almost negligible) concentrations when they tested purified PCR products. So, we decided to use our unpurified products and followed the rest of the protocol as stated (given our gel result, we deemed it reasonable to assume that the majority of DNA content was the desired amplicon.
| Reagent | Stock Conc. (ng/µL) | Desired Conc (ng/µL) | Volume (µL) |
|---|---|---|---|
| Backbone Fragment | 50 | 25 | 0.5 |
| Color Fragment (Single) | 50 | 50 | 1.0 |
| Gibson Assembly Mix | 2X | 1X | 5.0 |
| Nuclease-free water | — | — | 3.5 |
| Total Volume | — | — | 10.0 |
The reaction was incubated at 50°C in the thermocycler for 30 minutes.
Transformation
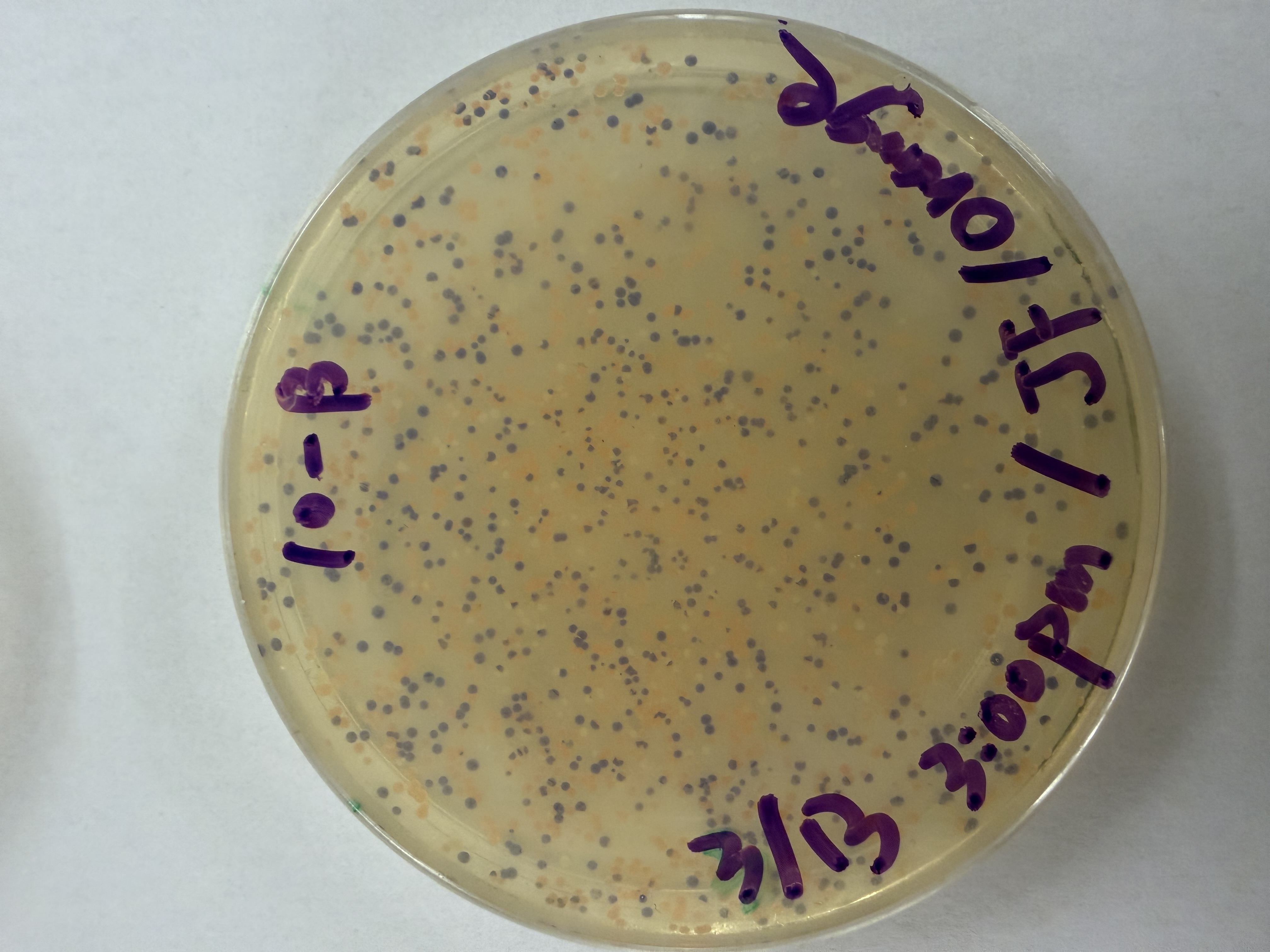
We compared two competent E. coli strains: DH5α and 10-beta.
We thawed our competent cells, and mixed 20µL of cells with 4µL from each of our Gibson Assembly reactions and incubated for 30 minutes on ice. We also did an additional reaction with just the native plasmid (DH5α with mUAV) to serve as a positive control for successful transformation. To create the control, I only utilized 1µL of the plasmid to try and somewhat match the total concentration of the samples.
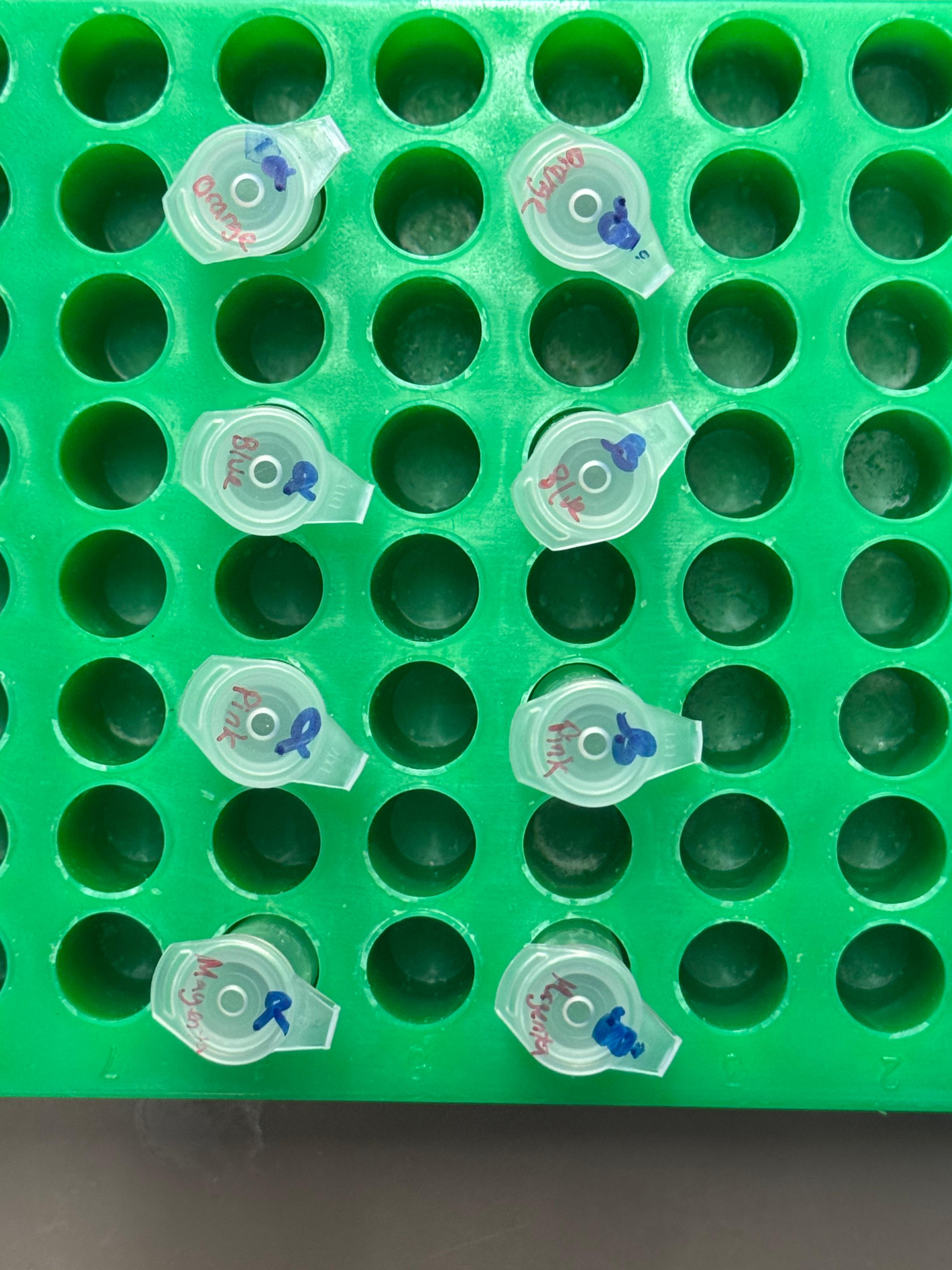


At this next step, we again diverged slightly from the protocol. We transfered the transformation reactions into PCR tubes containing 100µL of SOC medium and performed the heat shock step in that solution in the thermocycler. The heat shock importantly was only 45 seconds, and the cells were put on ice immediately after.
- Incubation: Mixed competent cells with Gibson products and incubated on ice for 30 minutes.
- Heat Shock: Performed in the thermocycler for 45 seconds in SOC medium, then immediately returned to ice.
- Outgrowth: Incubated for 60 minutes (it was really ~45 minutes) using a makeshift shaker made from a pipette tip box.
We plated the entire incubation (~124µL) of each transformation onto LB-Agar plates with Chloramphenicol using glass beads for even spreading.


Results
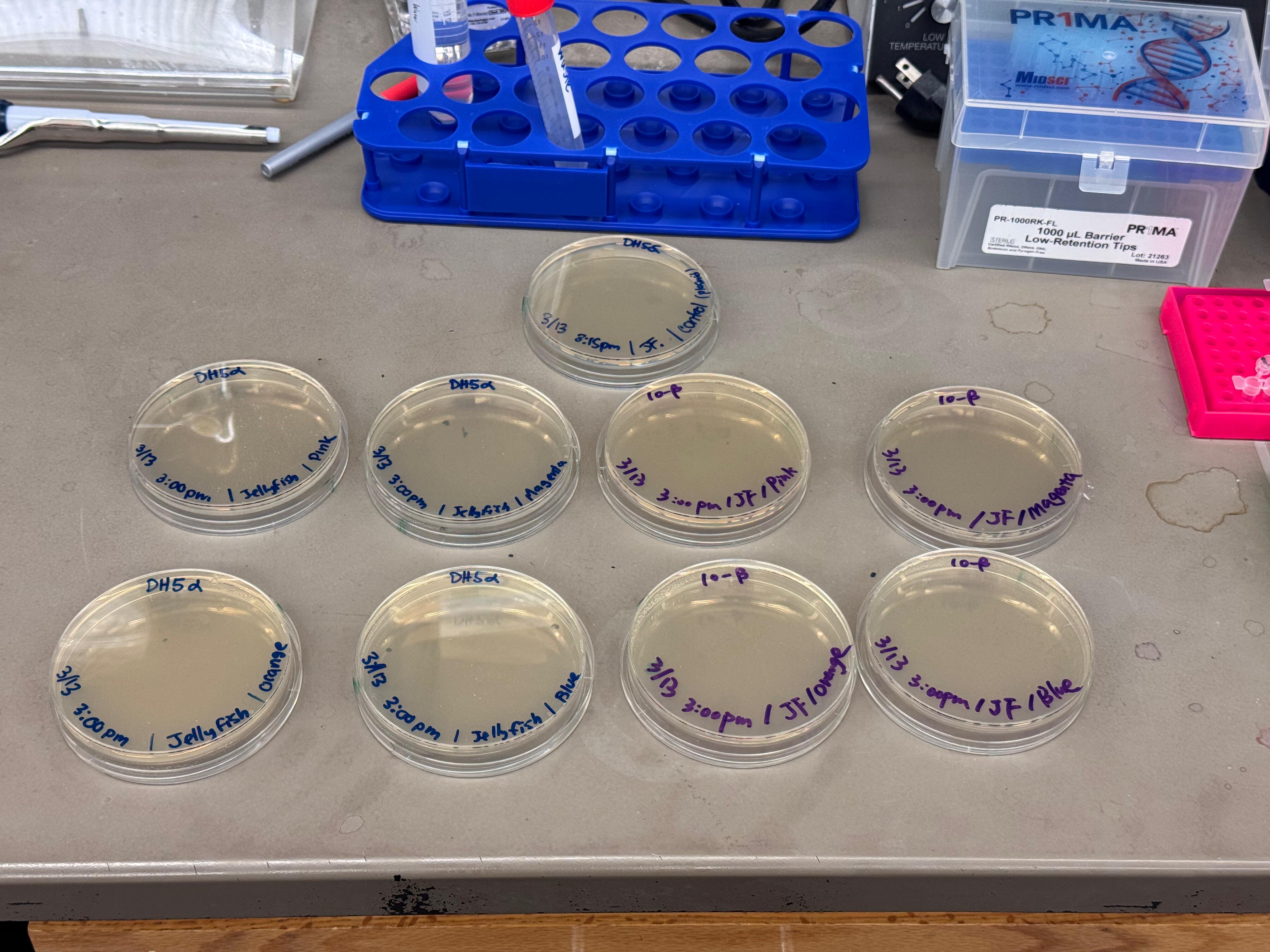
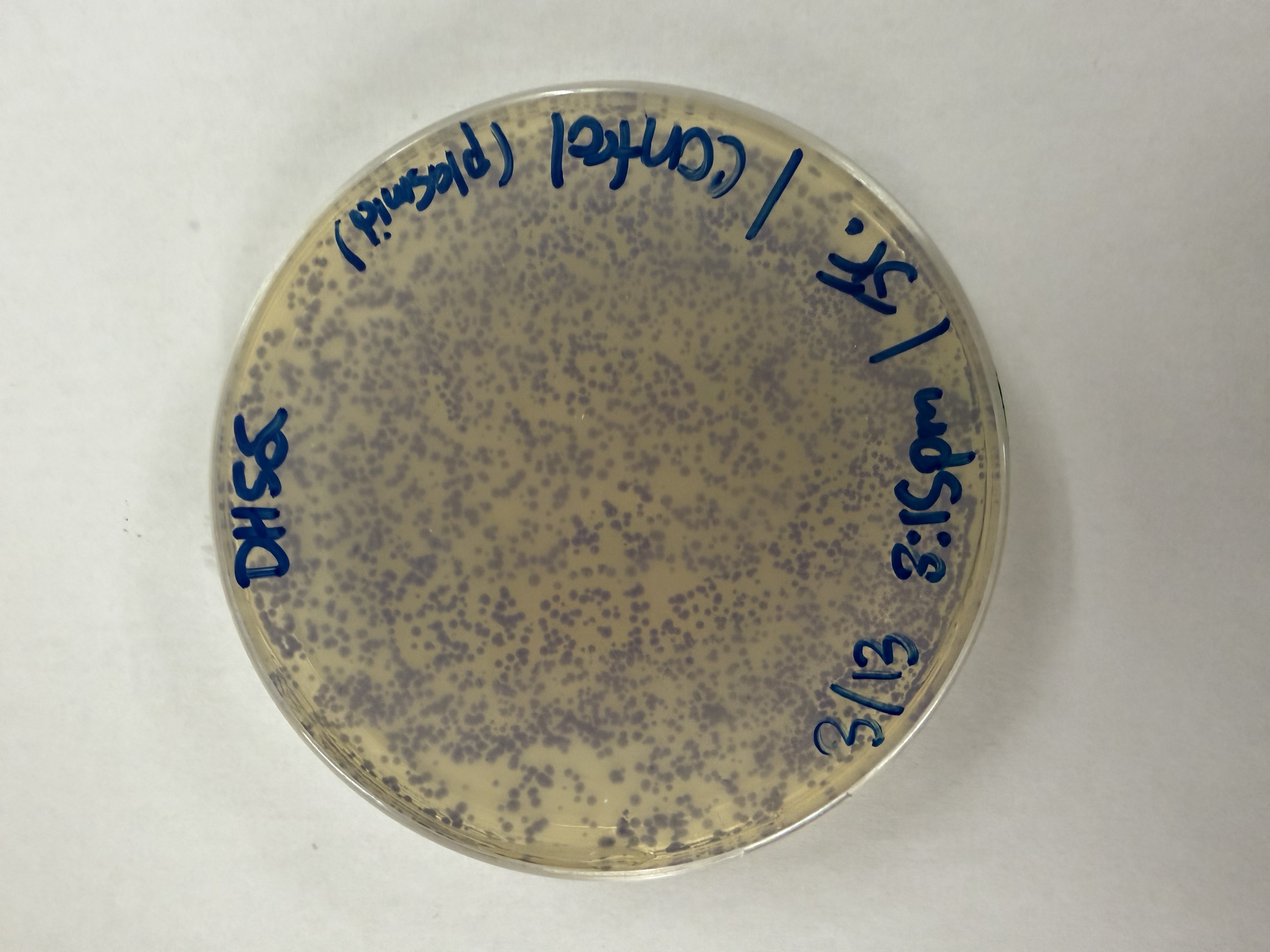
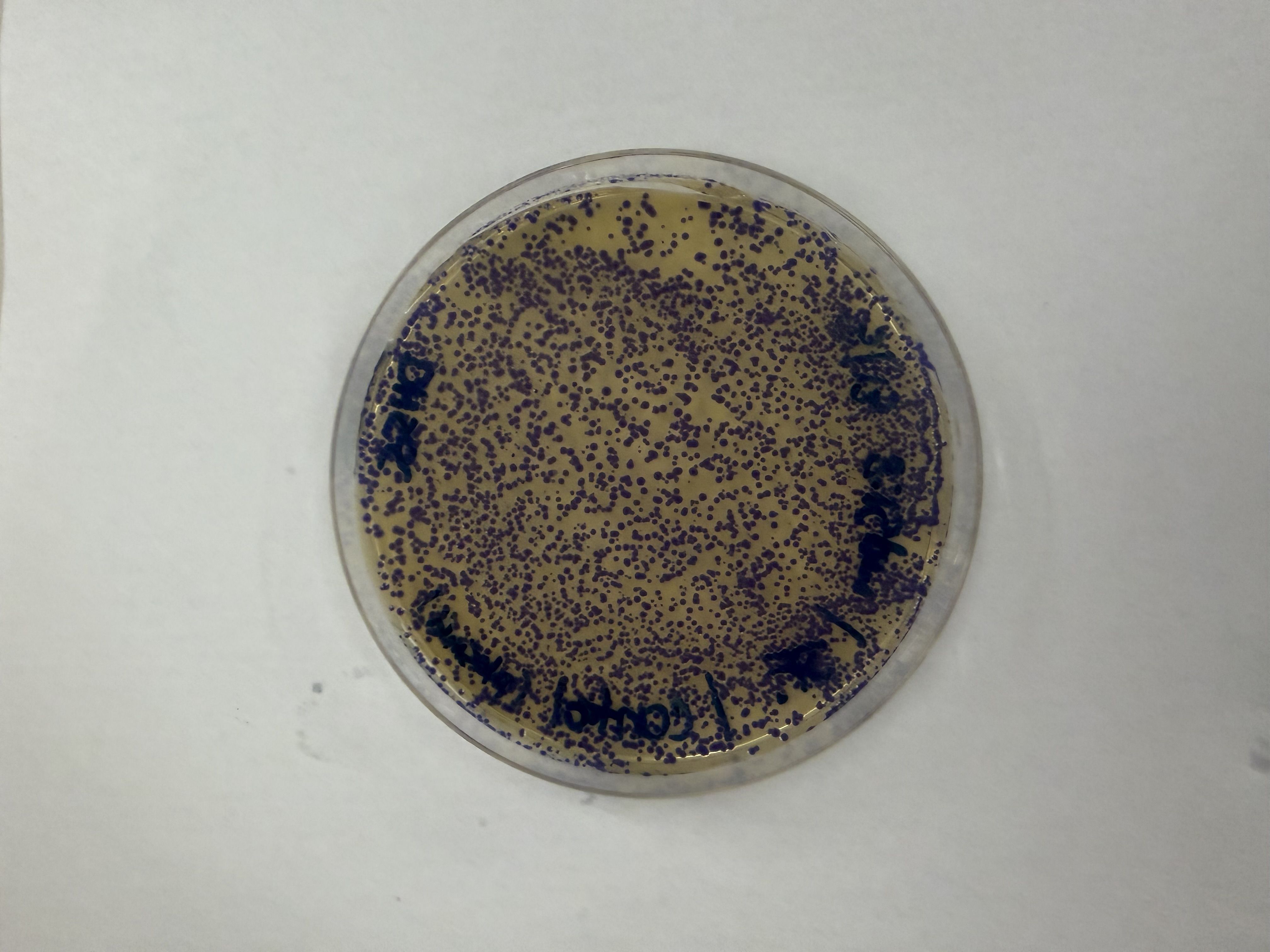
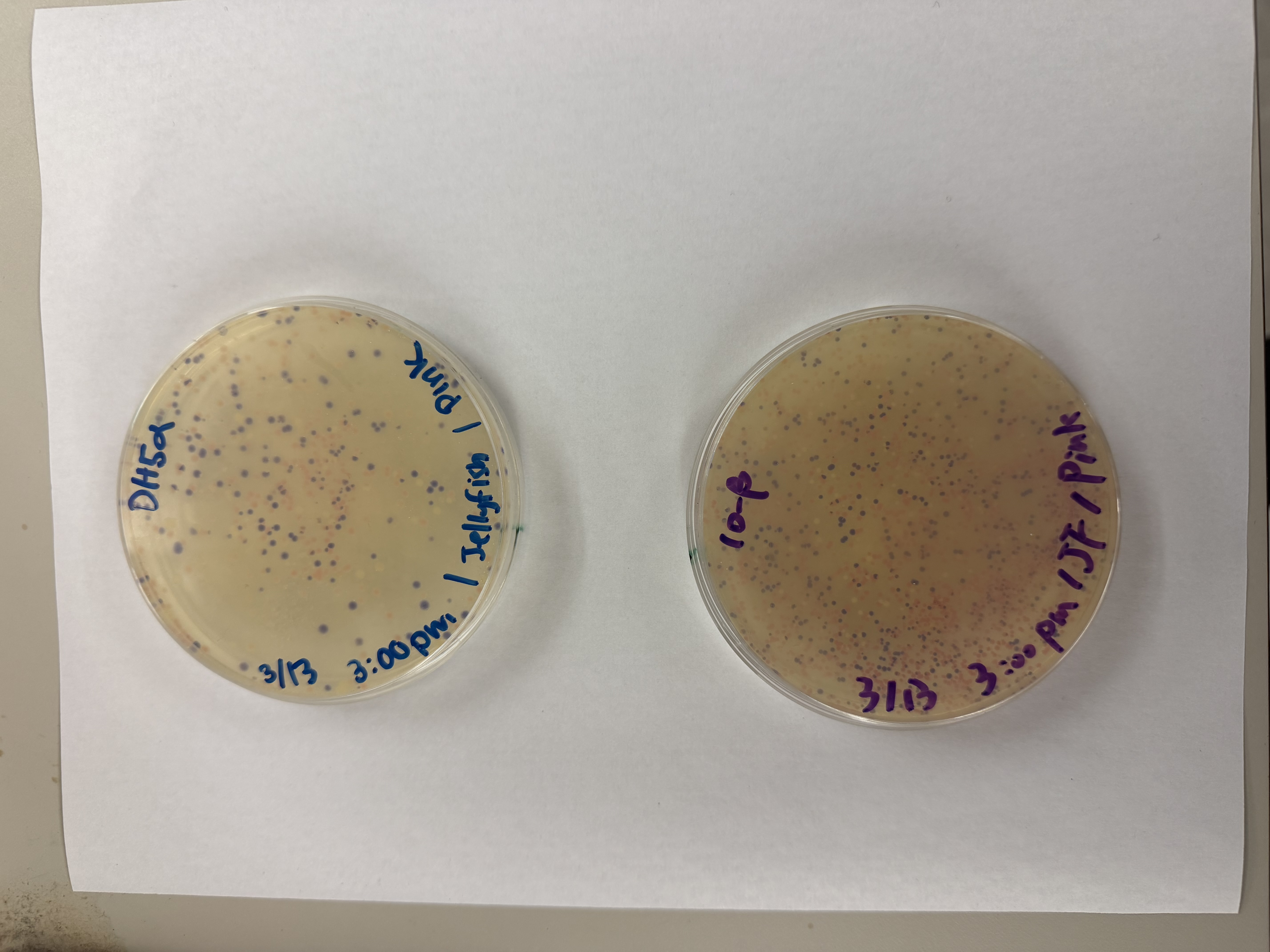
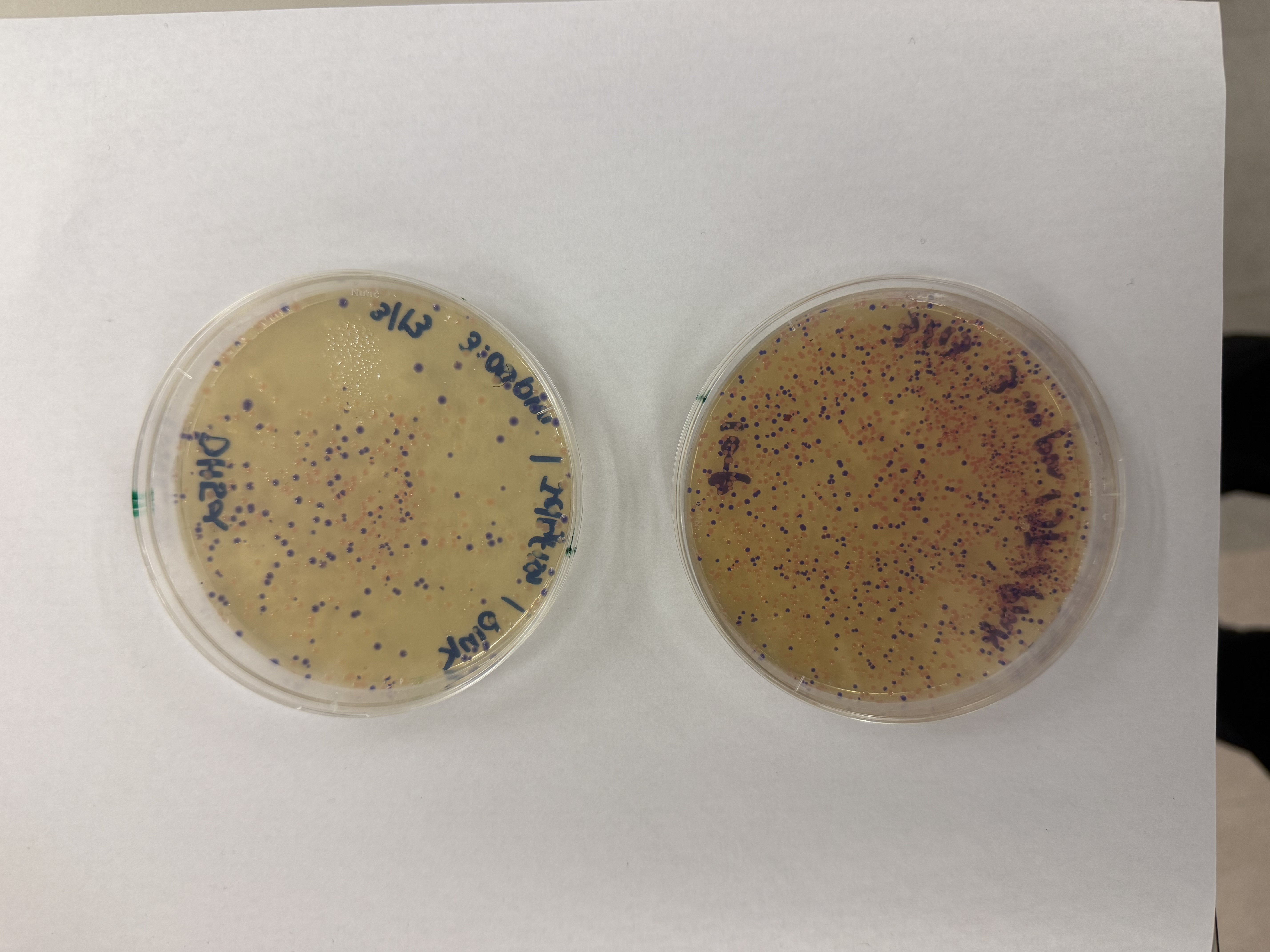
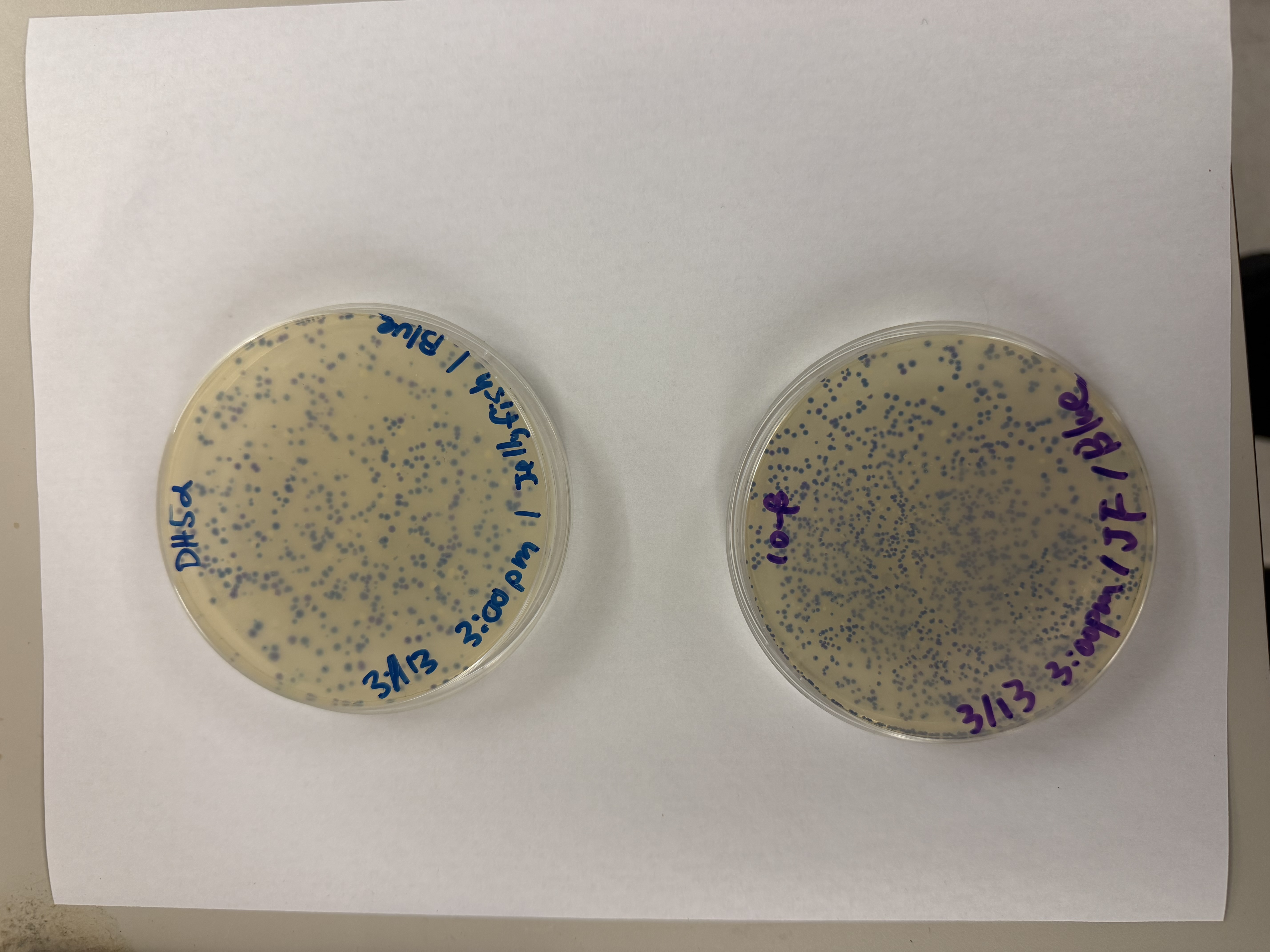
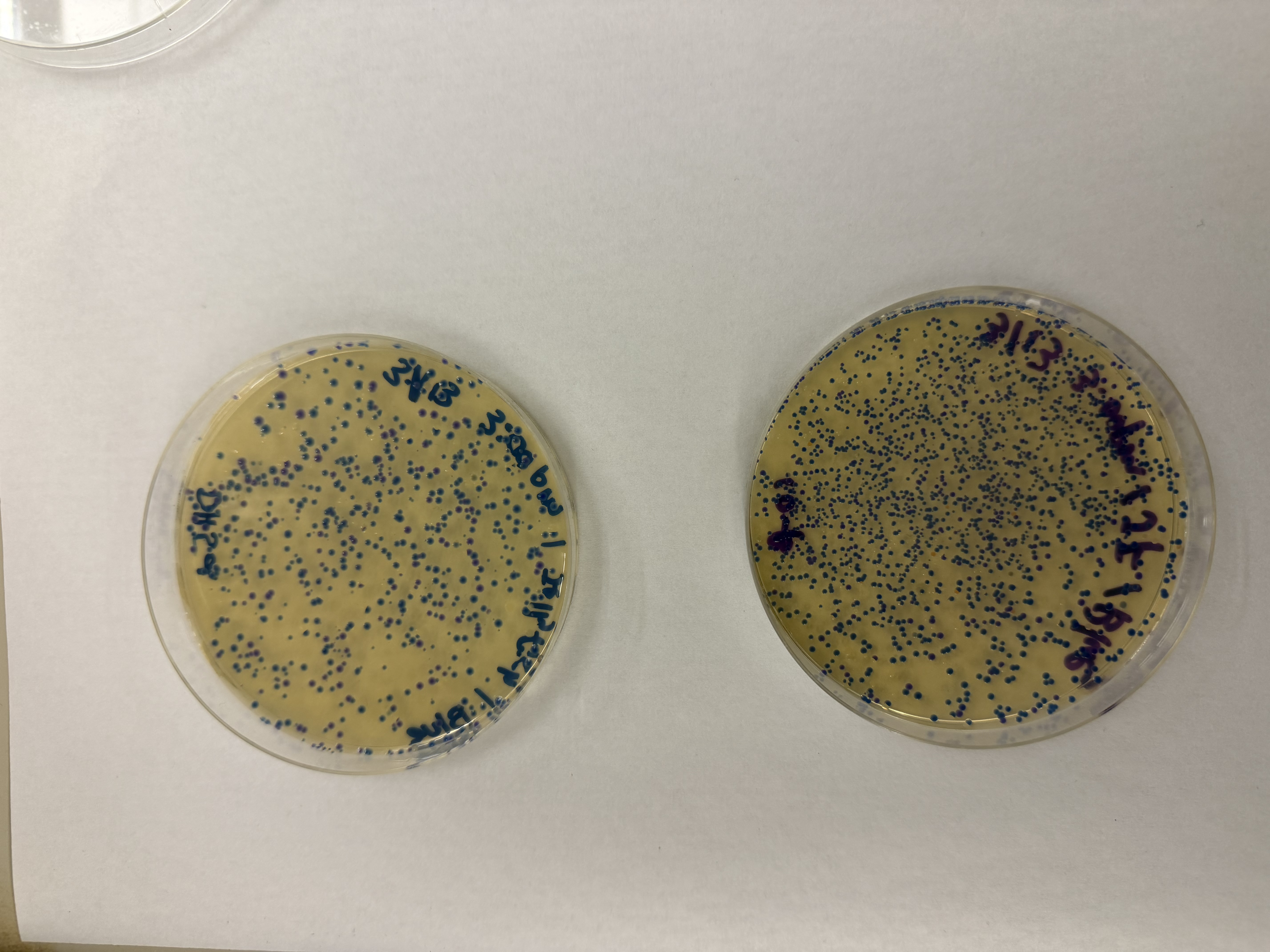
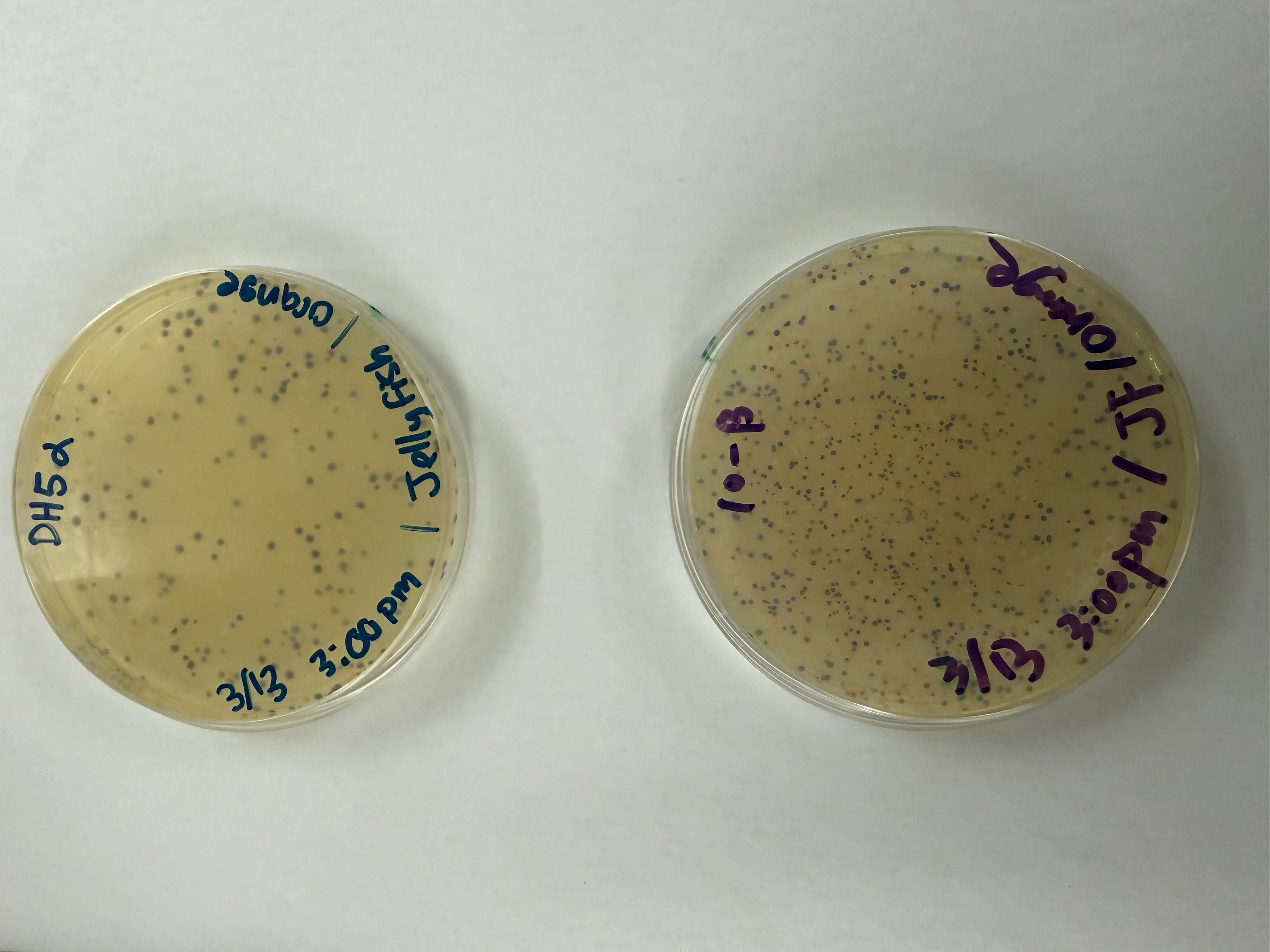
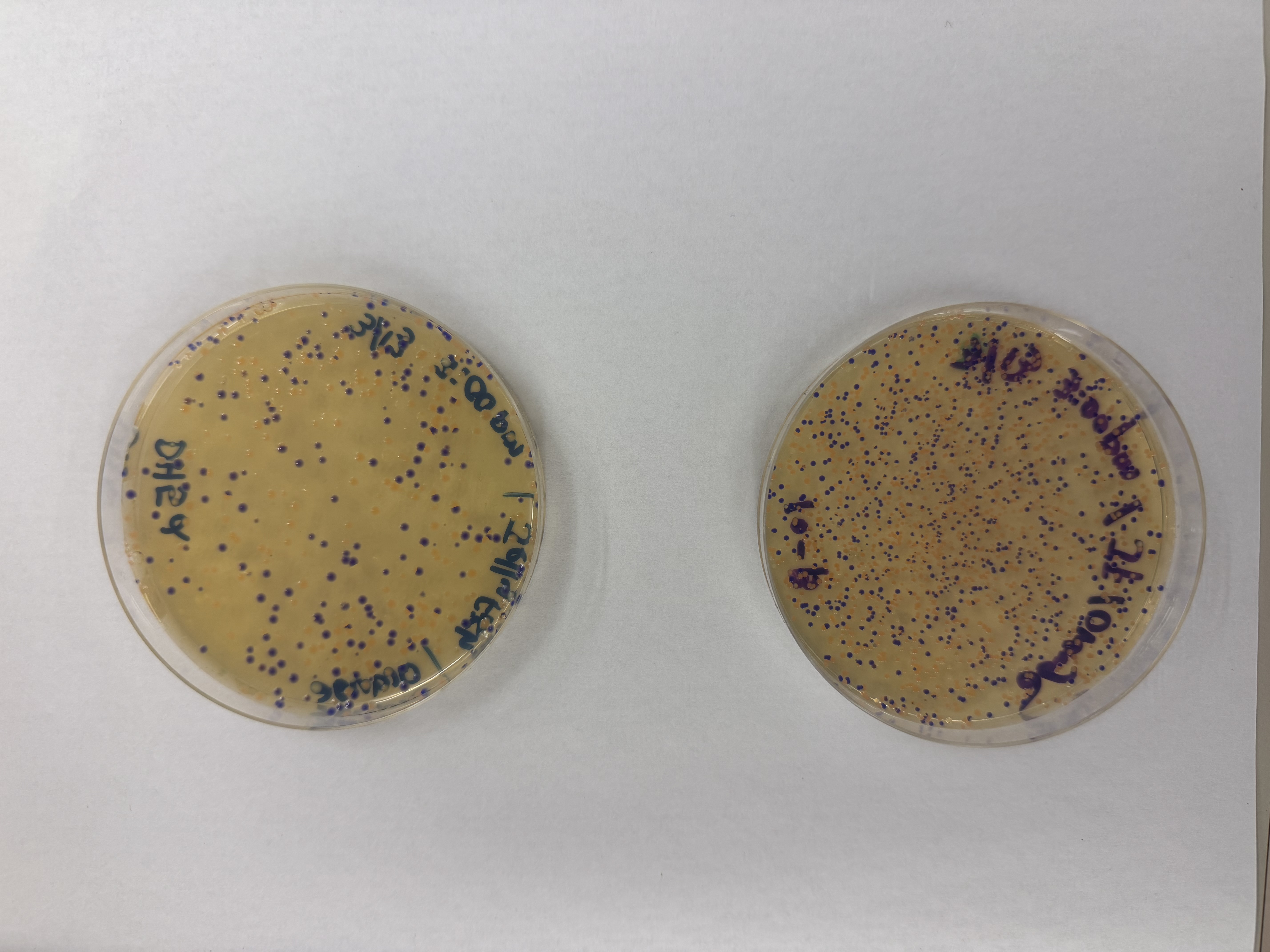
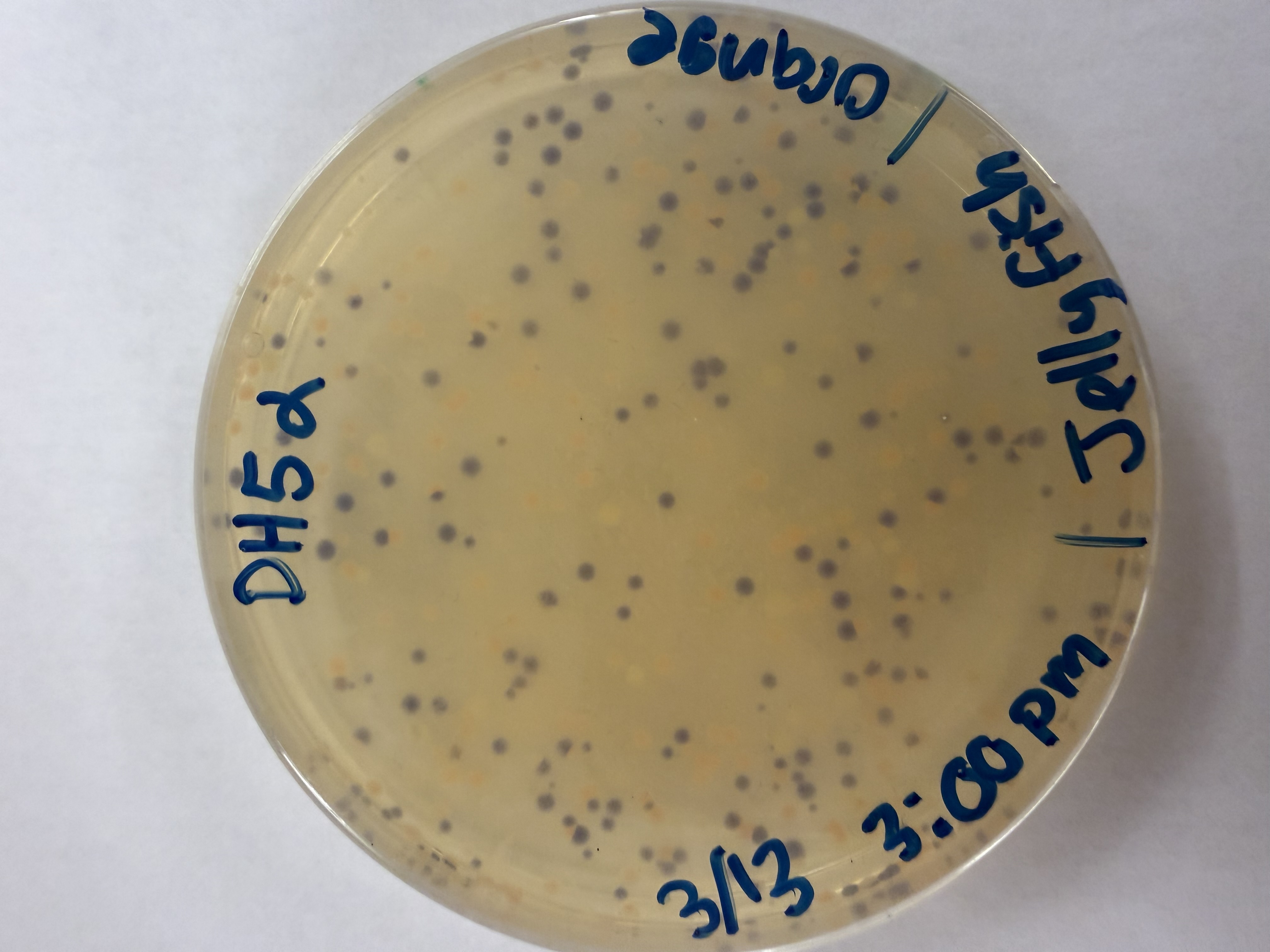
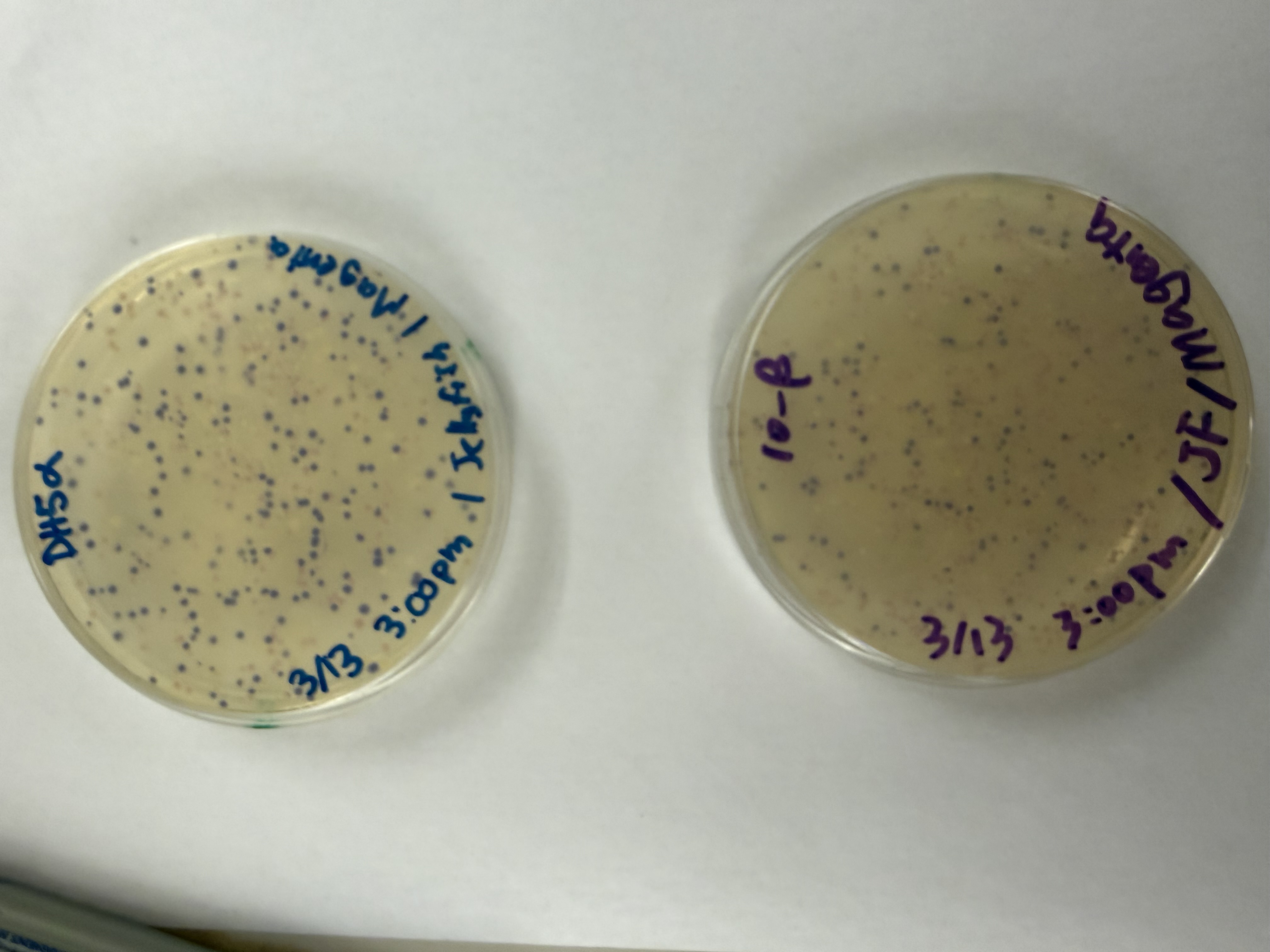
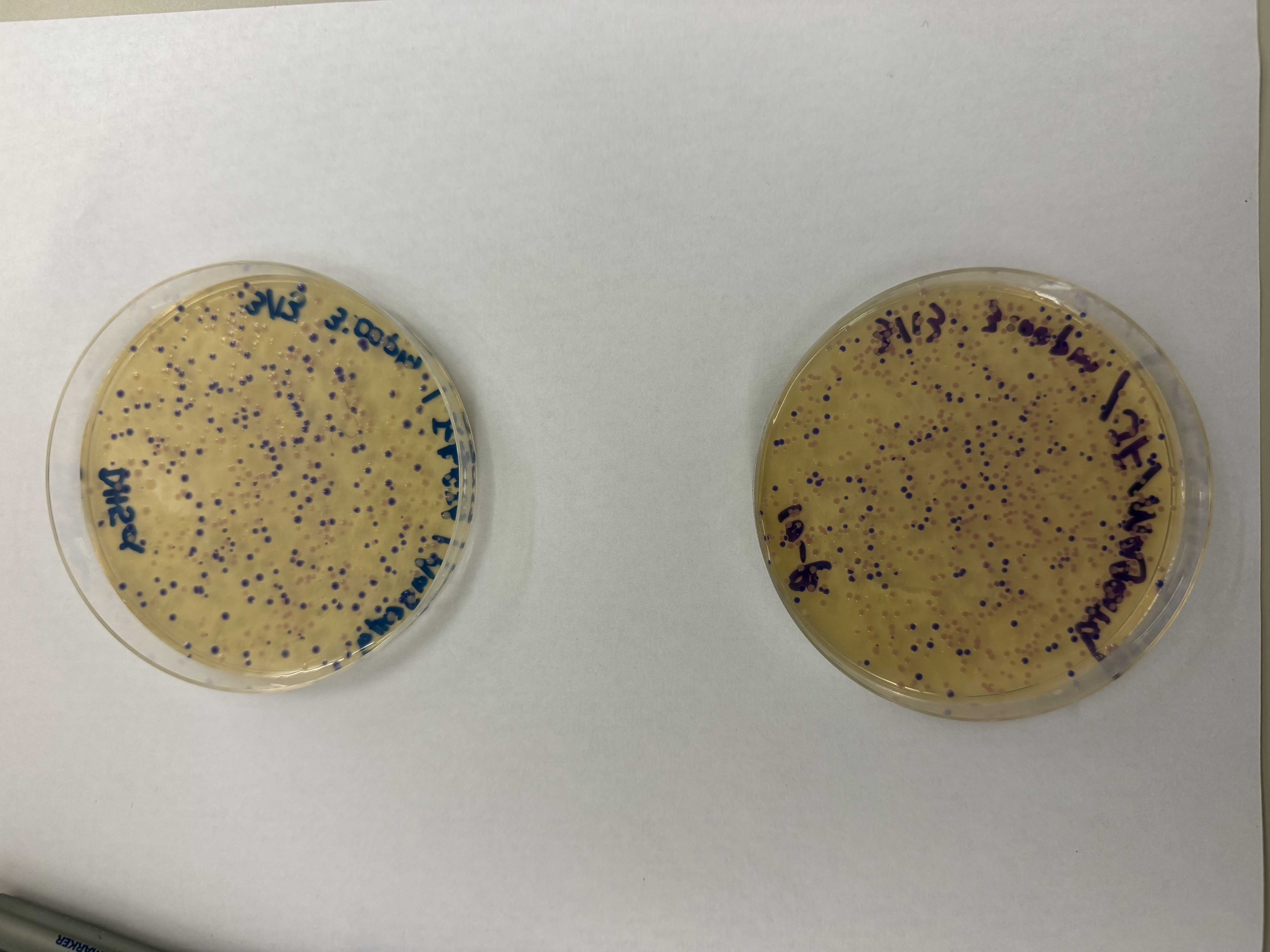
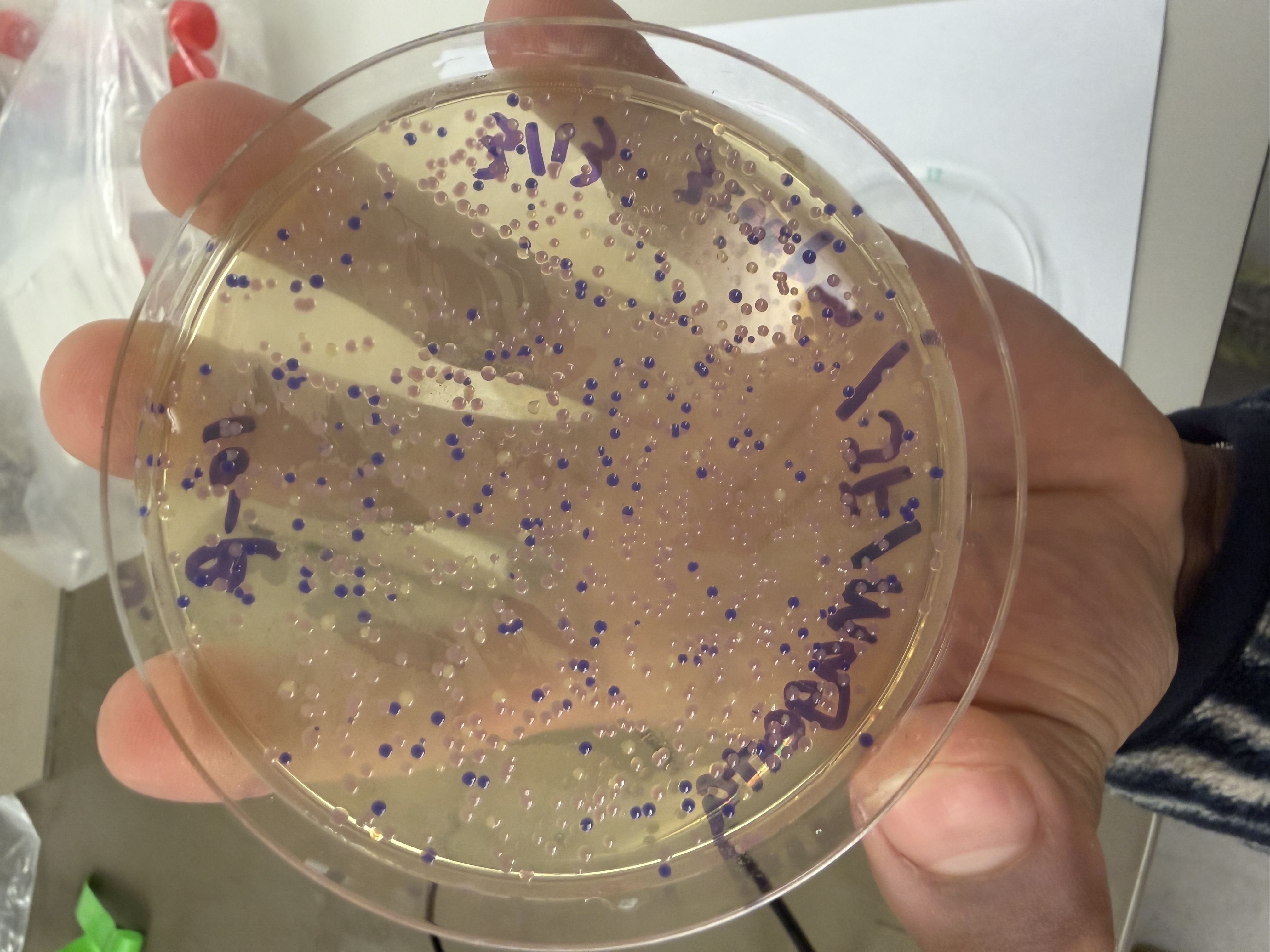
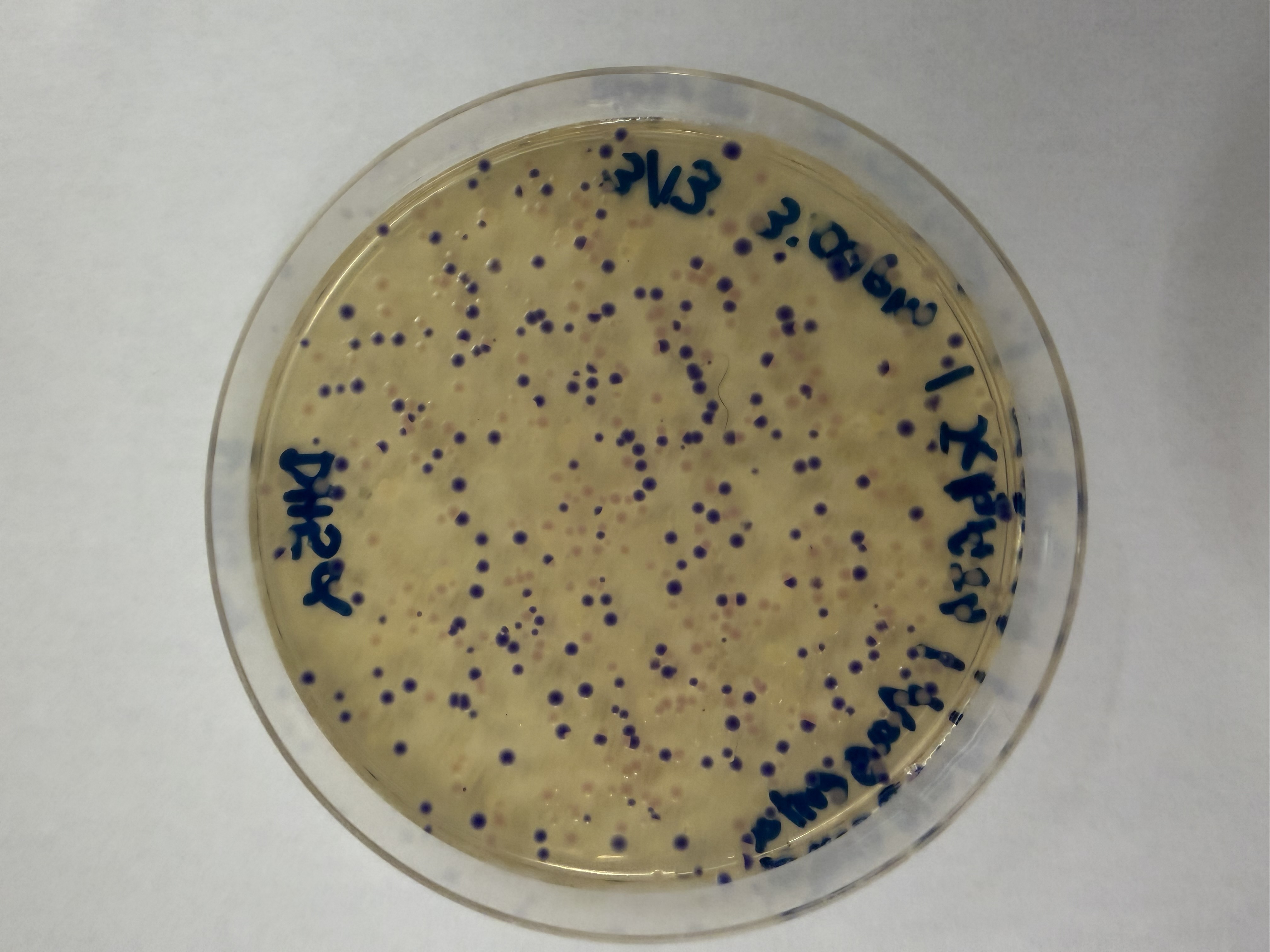
After 72 hours of incubation, the results were highly successful! We achieved the targeted chromophore mutations across both cell lines.
Analysis
The positive control confirmed the effectiveness of the assembly. While some purple colonies (native plasmid) were present on all plates, each plate showed distinct colored colonies (Orange, Pink, Blue, Magenta), indicating successful Gibson Assembly and transformation.
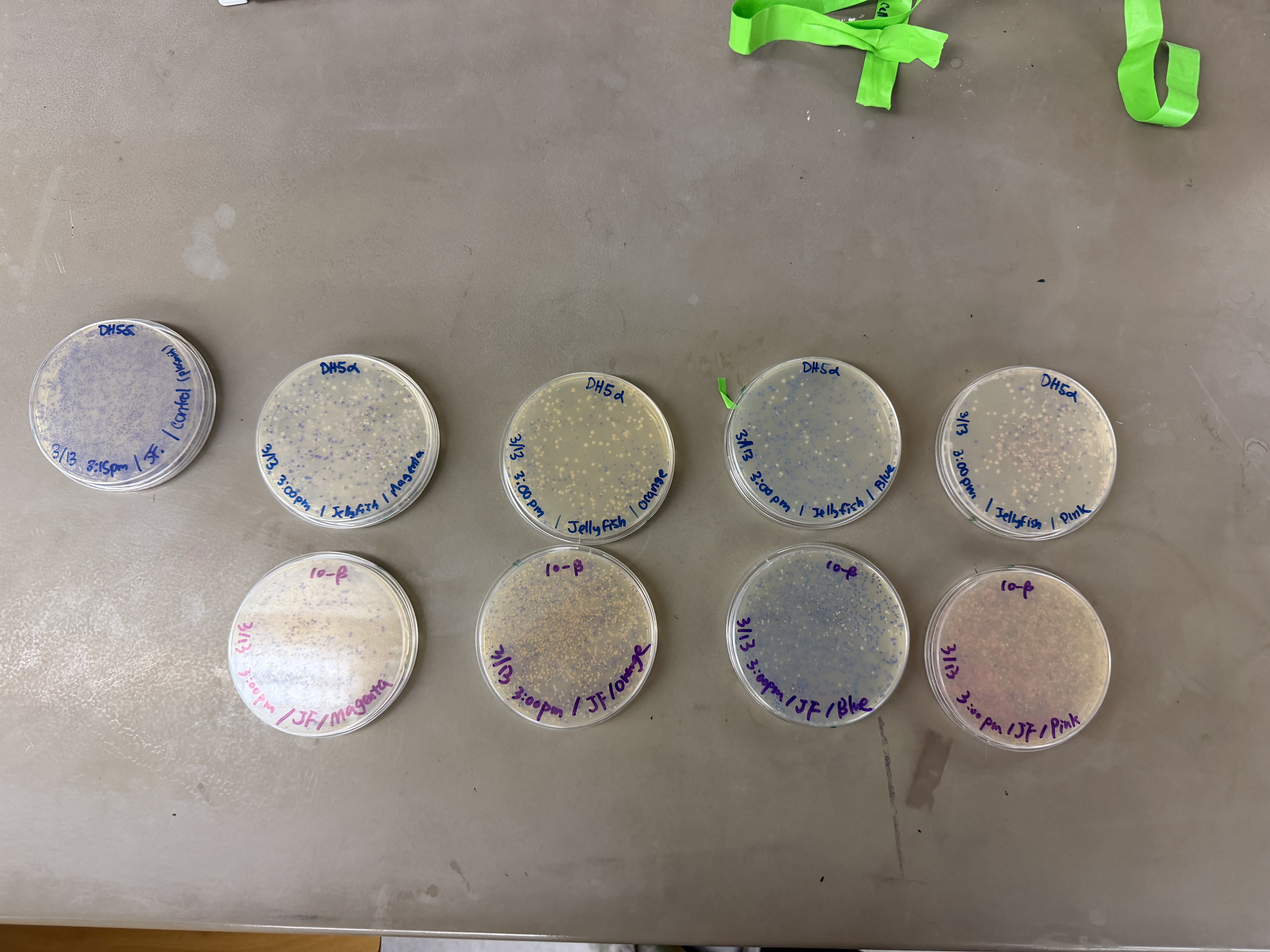


Control
Pink
Blue
Orange
Magenta
Gemini AI was consulted for formatting and content organization