Week 2 HW: DNA Read, Write, & Edit
Part 1: Benchling & In-Silico Gel Art
Using Benchling, I imported the Lambda DNA sequence and simulated restriction enzyme digestions with EcoRI, HindIII, BamHI, KpnI, EcoRV, SacI, and SalI. The goal was to design a gel art pattern inspired by Paul Vanouse’s Latent Figure Protocol.
The design depicts a figure in a rocky victory pose with a crying face, using 5 lanes of selected enzyme combinations to form the image.
Full virtual digest — all 7 enzymes (reference):
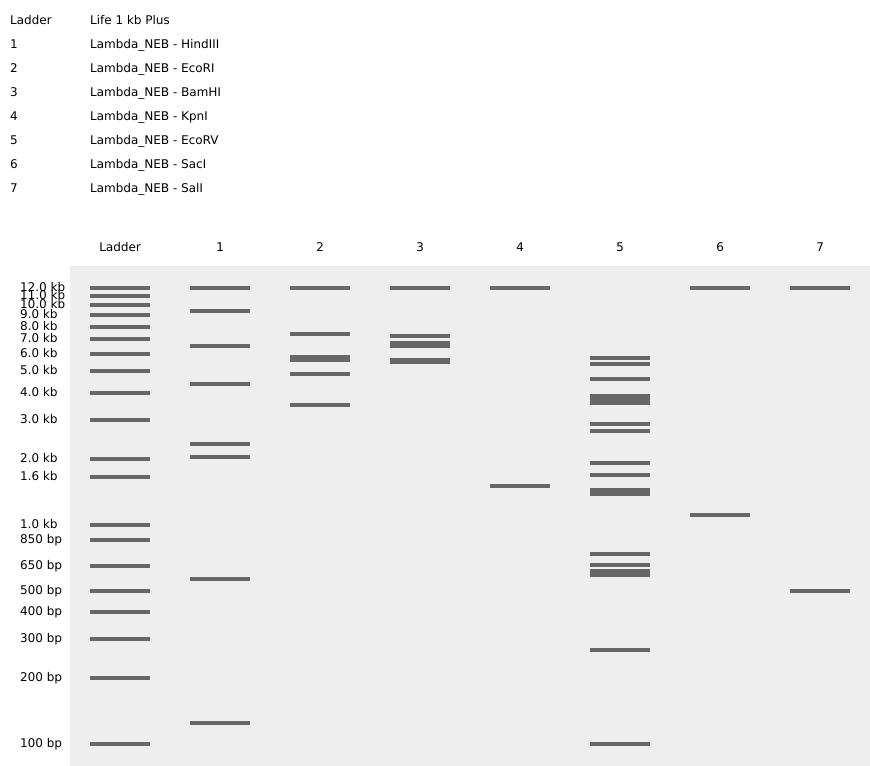


Gel art design — victory pose with crying face:


The final design uses the following lanes:
- Lane 1: Lambda_NEB - BamHI
- Lane 2: Lambda_NEB - EcoRV
- Lane 3: Lambda_NEB - KpnI
- Lane 4: Lambda_NEB - EcoRV
- Lane 5: Lambda_NEB - BamHI
Part 2: Gel Art - Restriction Digests and Gel Electrophoresis
We performed the wet-lab restriction digest and gel electrophoresis experiment designed in Part 1, following the Gel Art protocol. Photos of the gel were taken and will be uploaded shortly.
In progress — wet lab gel photos to be uploaded from Discord.
Part 3: DNA Design Challenge
3.1 Protein of Interest
I selected beta-casein (Bos taurus), a milk protein. Accession: UniProt P02666
3.2 Reverse Translation
The protein sequence was reverse translated to DNA using SMS2 Reverse Translate:
3.3 Codon Optimization
The reverse translated sequence was codon optimized for E. coli expression using VectorBuilder Codon Optimization:
Codon optimization translates a DNA sequence to match the codon preferences of a target organism. Beta-casein is naturally expressed in bovines (Bos taurus), but here I optimized for E. coli since it is a different organism with different preferred codon usage. By remapping the codons, we maximize translational efficiency and protein yield in the new host.
3.4 Protein Production Technologies
With the codon-optimized sequence, the protein can be produced via two approaches:
- Cell-dependent: The codon-optimized DNA is introduced into E. coli cells (e.g. via a plasmid). The cell’s own transcription and translation machinery reads the sequence and produces the protein.
- Cell-free: The necessary cellular components (ribosomes, tRNA, polymerases, etc.) are extracted and used in vitro to transcribe and translate the DNA into protein, without needing a living cell.
3.5 Transcriptional Protein Diversity (Optional)
Skipped.
Part 4: Twist DNA Synthesis Order
Following the Twist tutorial, I built an expression cassette in Benchling containing a promoter, RBS, start codon, codon-optimized coding sequence, 7x His tag, stop codon, and terminator. The linear sequence was then uploaded to Twist to prepare a clonal gene order, and the resulting construct was imported back into Benchling to verify the plasmid design.
Step 1 — Linear expression cassette in Benchling (924 bp):
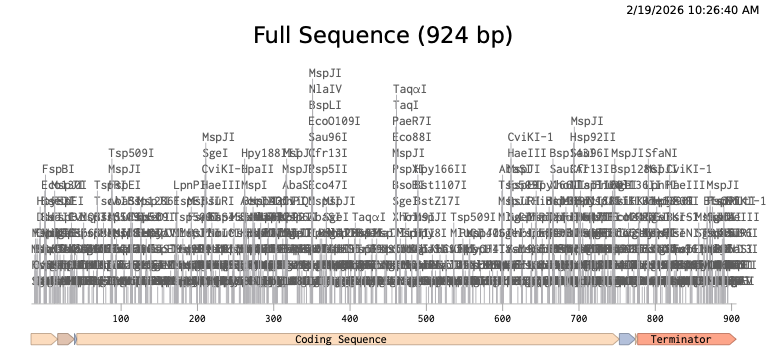


Step 2 — Uploaded to Twist (clonal gene order):
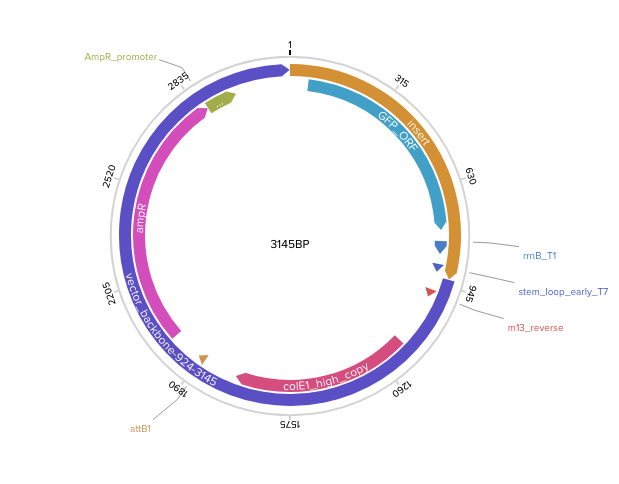

Step 3 — Final circular plasmid verified in Benchling (3145 bp, pTwist Amp High Copy):
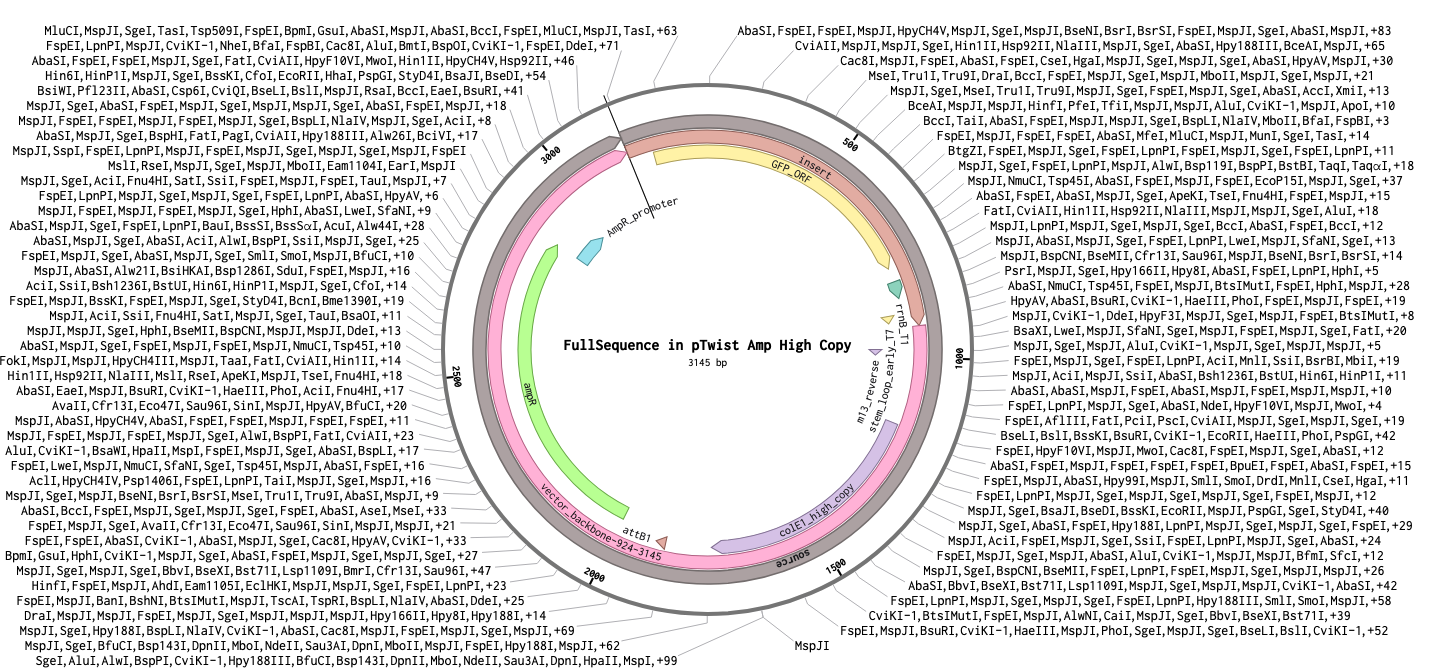

Part 5: DNA Read/Write/Edit
5.1 DNA Read
In progress.
5.2 DNA Write
In progress.
5.3 DNA Edit
In progress.