Week 1 HW: Principles and Practices

Governance means different things to different people, but what truly matters to me is building systems that share power rather than seize it. I imagine resilient structures that adapt, nurture well-being in many forms, and thrive on transparency and shared commitment.
First, describe a biological engineering application or tool you want to develop and why
Plants can be reciprocal and help each other. For example, ectomycorrhizal (ECM) fungi, which form sheaths around roots, help restore soil and make plants more resilient, especially in the face of climate change. I want to create a system inspired by the ectomycorrhizal (ECM) fungi living model of resilience. By mimicking their metabolism and distribution, we could recover and manage resources more wisely, whether in organizations, food systems, or conservation efforts. Imagine a biological pump that naturally generates, transports, and processes information, strengthening our collective resilience.
Arbuscular mycorrhizae (AM), a type of endomycorrhizal fungi, associate with 80% of terrestrial plants. AM supports soil stabilisation, nutrient cycling, microbial diversity, and soil organic matter (SOM) formation.
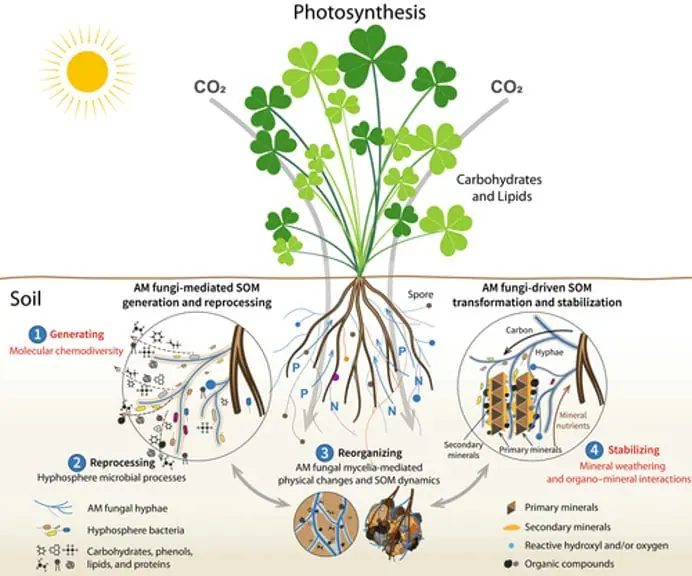

Figure 1. “Diagram showing the updated conceptual framework of arbuscular mycorrhizal (AM) fungi-mediated soil organic matter (SOM) dynamics. Plants fix carbon through photosynthesis, which is then delivered to AM fungi. Arbuscular mycorrhizal fungi influence SOM dynamics through four pathways classified as (1) generating, (2) reprocessing, (3) reorganizing, and (4) stabilizing” Wu et al. 2024. (Image credits: https://doi.org/10.1111/nph.19178)reference: https://cid-inc.com/blog/2024-research-insights-how-mycorrhizal-fungi-benefits-agriculture/
Wu et al. (2024) introduced a framework explaining AM’s role in soil organic matter formation, considering factors such as organic compound diversity, mineral weathering, chemical interactions, and hyphosphere microbial contributions. They identify four pathways: SOM generation, reprocessing, reorganisation, and stabilisation (see Figure 1).
Generating: AM fungi produce exudates, metabolites, mucilage, and necromass. Wu et al. refer to the diversity, composition, and properties of these compounds as chemodiversity. They argue that chemodiversity, rather than individual compounds, is key to SOM composition, microbial biodegradation rates, and persistence.
Reprocessing: AM fungi attract specific microbes to their hyphosphere, the soil area influenced by AM hyphal exudates. These microbes drive soil biochemistry by decomposing SOM components that AM fungi cannot process. Through internal and extracellular pathways, they break down and assimilate SOM, contributing to chemodiversity, persistence, and SOM resynthesis. This process is known as the hyphosphere “microbial carbon pump.”
Reorganising: The fungi’s mycelial growth, expansion, and colonisation change the soil’s physical porosity and hydraulic properties. While AM stabilises macro soil aggregates, the mycelial dynamics increase micro aggregate turnover, water infiltration, soil-water retention capacity, hydraulic conductivity, and redistribution of AM exudates. The changing soil conditions due to mycelial expansion also change nutrient availability, temperature, and oxygen. It results in SOM redistribution and transformation.
Stabilising: AM fungi cause mineral weathering and alter interactions that influence SOM formation and stabilisation. AM rock mineral weathering makes nitrogen, phosphorus, potassium, and magnesium available in soils that form secondary compounds with different sizes, surfaces, and reactivity. This can alter mineral absorption, catalysis, and oxidation of SOM. These processes are called the “soil mineral carbon pump”.
Takeaway: The new concept can explain AM’s role in small- to large-scale SOM dynamics, which can help develop mycorrhiza-based technologies to enhance soil health.
I am facinating of the concept of carbon cycling and the major processes and mechanisms involved in this process, and how it can create positive changes in the ecosystem:
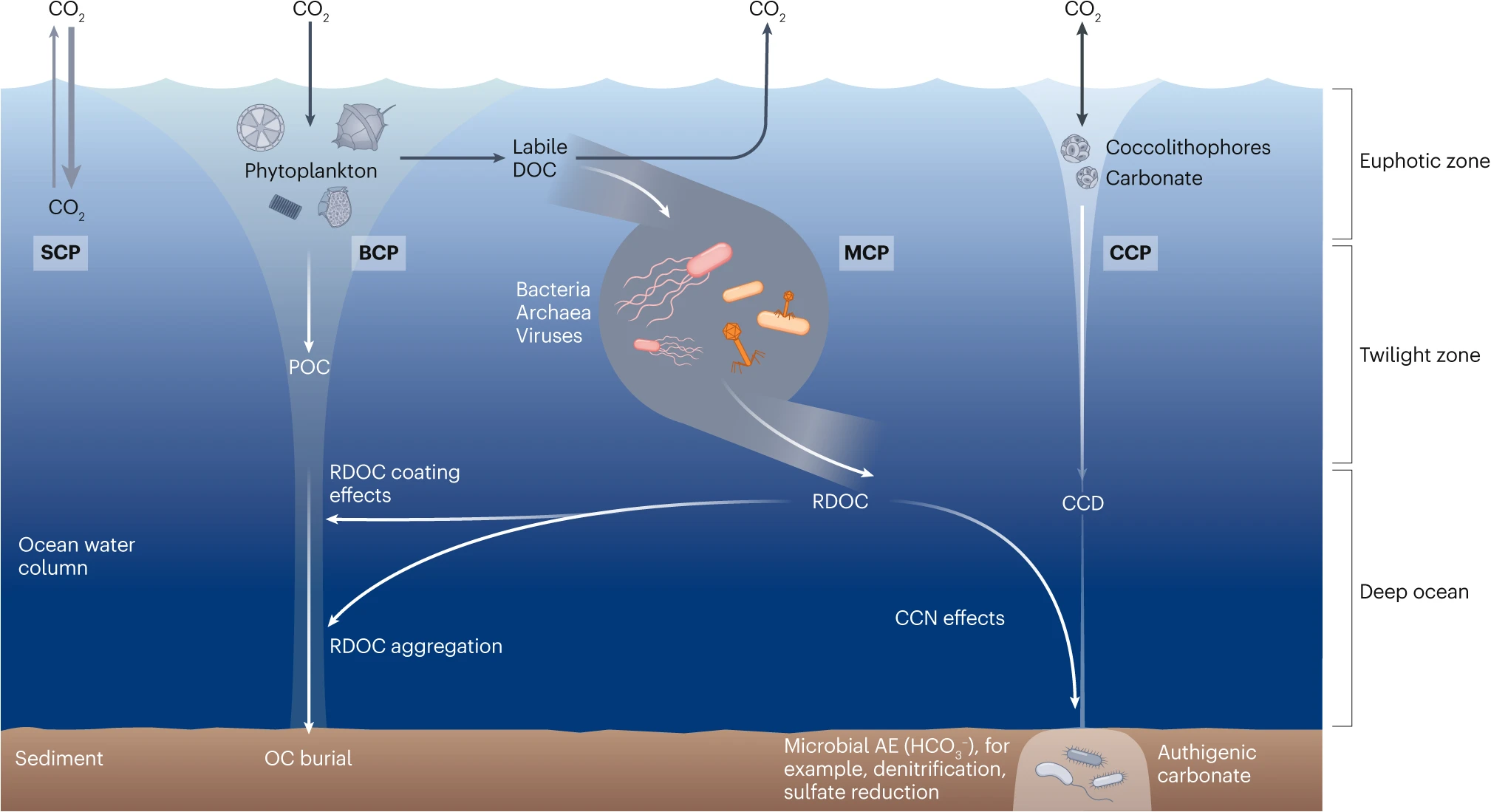
 Figure 2. The solubility carbon pump (SCP) is driven by the difference of CO2 partial pressures between the atmosphere and surface waters; exchanges of CO2 occur through dissolution into water or release into the air. Generally, the SCP refers to the pumping of CO2 from the atmosphere into the ocean driven by abiotic processes such as lowering temperature and downward mixing. The biological carbon pump (BCP) refers to a series of biogeochemical processes that transport organic carbon (mainly particulate organic carbon (POC)) from the surface to the ocean interior
Figure 2. The solubility carbon pump (SCP) is driven by the difference of CO2 partial pressures between the atmosphere and surface waters; exchanges of CO2 occur through dissolution into water or release into the air. Generally, the SCP refers to the pumping of CO2 from the atmosphere into the ocean driven by abiotic processes such as lowering temperature and downward mixing. The biological carbon pump (BCP) refers to a series of biogeochemical processes that transport organic carbon (mainly particulate organic carbon (POC)) from the surface to the ocean interior
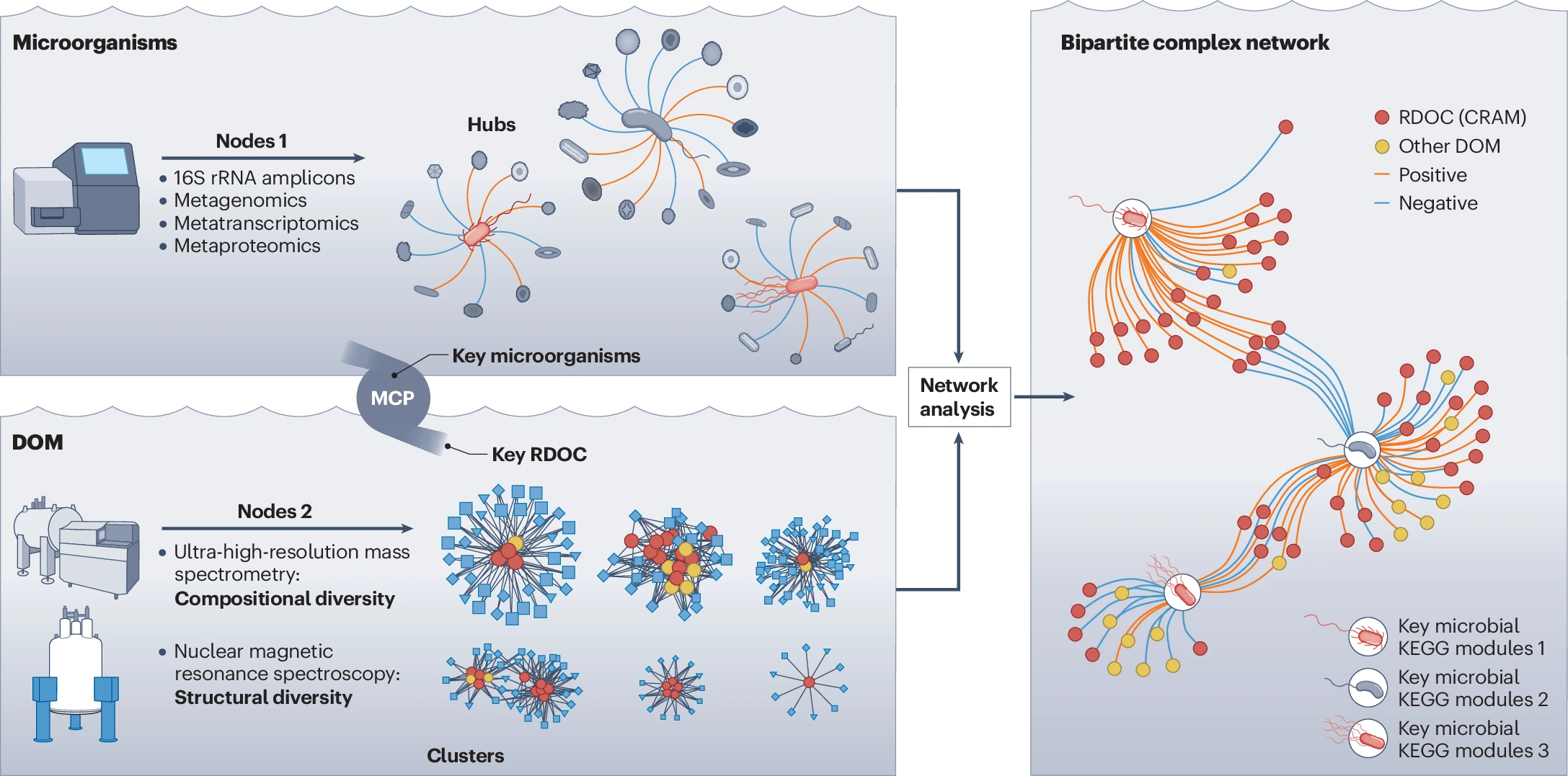
 Figure 3. Microorganism–dissolved organic matter (DOM) complex networks consist of two types of nodes: microbial and DOM. Connections are made between nodes based on correlations of data sets. Microbial diversity can be analysed using 16S rRNA amplicons, metagenomics, metatranscriptomics, and metaproteomics, as described in Jiao, N., Luo, T., Chen, Q. et al. The microbial carbon pump and climate change. Nat Rev Microbiol 22, 408–419 (2024). https://doi.org/10.1038/s41579-024-01018-0
Figure 3. Microorganism–dissolved organic matter (DOM) complex networks consist of two types of nodes: microbial and DOM. Connections are made between nodes based on correlations of data sets. Microbial diversity can be analysed using 16S rRNA amplicons, metagenomics, metatranscriptomics, and metaproteomics, as described in Jiao, N., Luo, T., Chen, Q. et al. The microbial carbon pump and climate change. Nat Rev Microbiol 22, 408–419 (2024). https://doi.org/10.1038/s41579-024-01018-0
Next, describe one or more governance/policy goals related to ensuring that this application or tool contributes to an “ethical” future, like ensuring non-malfeasance (preventing harm). Break big goals down into two or more specific sub-goals
Restoring governance in our ecosystems, especially within food systems and conservation, is crucial for protecting key species. By studying these organisms, we can build a deeper, more collaborative knowledge base and move toward distributed, systemic governance.
Purpose: Transform the soil by introducing this plant, aiming to regenerate and sustain a thriving, healthy food environment.
Design: Build partnerships among small businesses, farmers, and policymakers to expand these practices across diverse ecosystems.
Assumptions: it is not a law or regutlation is ian a mediation with the ecosystmen
Risks of Failure & Success: Outcomes depend on the ecosystem, but the ultimate goal is to design interventions that measure plant survival and resilience as the true indicators of success.
Although I am not entirely certain how the process functions, I believe in the potential of replicating plant-based or bioinspired systems to support regeneration and address deforestation within food systems.
First, implementing plant-based systems can address the irrigation and soil nutrition needs of various sectors. For example, cacao plantations, which are highly sensitive to extreme temperatures, could benefit from such approaches and support regenerative agriculture. Monitoring these plants could facilitate the development of sensing systems and enable the implementation of more complex networks for reforestation efforts.
Idea 2
My family’s cacao farm in Colombia has always sparked my curiosity about our crop’s roots and how we might make it more sustainable. For years, we have wondered about the true quality of our cacao and whether it belongs to the prized Criollo variety, since its genetics remain a mystery. Now, I am exploring how synthetic biology could unlock new possibilities for our farm, strengthen local leadership, and add value by revealing and celebrating the unique DNA of our cacao. hire a vision with a longer term
Step 1: Identify and document ancestral cacao varieties, which can be the first stage of a HTGAA
- Map existing Criollo cacao varieties and their genetic diversity.
- Record sensory profiles, cultural significance, and local cultivation knowledge. Connect with the idea 1 to produce regenerative practices
- Collaborate with farmers, researchers, and local organisations. to contribut into a ecosytem DNA picture
Step 2 — Establish participatory cacao genetic biobanks
- Develop genetic repositories combining scientific methods with community participation.
- Ensure fair access, ethical data governance, and benefit sharing with producer communities.
- Integrate sustainable agricultural policies and biodiversity frameworks.
Step 3: Strengthen biodiversity conservation
- Preserve endangered cacao genetic resources.
- Promote agroecological cultivation models that maintain ecosystem health.
- Support climate resilience through genetic diversity.
Step 4: Empower local farming communities
- Recognise farmers as co-stewards of genetic heritage.
- Provide training, technical support, and participatory decision-making spaces.
- Foster economic opportunities linked to high-quality heritage cacao.
Step 5: Support sustainable certification and traceability
- Use genetic data to strengthen transparency in cacao supply chains.
- Enable certification schemes that reflect biodiversity conservation and ethical production.
- Improve governance mechanisms linking agriculture, sustainability, and cultural heritage.
| Does the option: | Option 1 | Option 2 | Option 3 |
|---|---|---|---|
| Enhance Biosecurity | |||
| • By preventing incidents | x | ||
| • By helping respond | |||
| Foster Lab Safety | |||
| • By preventing incident | |||
| • By helping respond | |||
| Protect the environment | x | ||
| • By preventing incidents | |||
| • By helping respond | x | ||
| Other considerations | |||
| • Minimizing costs and burdens to stakeholders | |||
| • Feasibility? | |||
| • Not impede research | x | ||
| • Promote constructive applications | x |
Bio Questions Week 1 <3 !
-> Question by J. Jacobson´s Presentation <-
Nature’s machinery for copying DNA is called polymerase. What is the error rate of polymerase? How does this compare to the length of the human genome. How does biology deal with that discrepancy?
As all concepts are new for me, I might need a bit more explanation of all concepts A polymerase is a type of enzyme, such as DNA or RNA polymerase, that synthesises long chains of nucleic acids by adding nucleotides to a template strand. These enzymes are essential for critical biological processes, including DNA replication, repair, and transcription. Polymerases are also vital in laboratory applications, most notably in the polymerase chain reaction (PCR) for amplifying DNA. https://en.wikipedia.org/wiki/Polymerase
Function: DNA polymerases catalyze the synthesis of DNA by adding nucleoside triphosphates, creating two identical DNA duplexes from one. Types: DNA polymerase (replicates DNA) and RNA polymerase (transcribes DNA into RNA) are the primary types, found in all living organisms. Mechanism: They require a primer and a template strand to function, adding nucleotides in a to direction. Accuracy: DNA polymerases often have built-in proofreading abilities ( to exonuclease activity) to ensure high-fidelity replication. PCR Application: Thermostable polymerases, such as Taq polymerase, are used in PCR to automate DNA copying, allowing for billions of copies to be made in a few hours. Structural Variation: Polymerases range from simple single proteins to complex, multi-subunit assemblies.
What is the error rate of polymerase? Throughput Error Rate Product Differential: ~10⁹ based on a consensus DNA polymerase is extremely accurate, with typical error rates around 1 mistake per 10⁵–10⁷ bases before cellular repair, and ~10⁹–10¹¹ per base per replication after all proofreading and repair.
How does this compare to the length of the human genome?
3.2 Gbp
How does biology deal with that discrepancy?
By error correcton MutS Repair System by consensus: Biology layers multiple error‑correction systems to shrink polymerase’s raw error rate down to ~1 mutation per genome copy.
How many different ways are there to code (DNA nucleotide code) for an average human protein?
I am not sure, but in this paper they present a method for constructing complex and diverse DNA sequences using DNA three-way junctions. Theoretically, because of genetic code degeneracy, an “average” human protein can be encoded by an astronomically large number of DNA sequences.
In practice, what are some of the reasons that all of these different codes don’t work to code for the protein of interest?
I guess because the fragmentation :S
-> Questions by LEPROUST´s Presentation <-
What’s the most commonly used method for oligo synthesis currently?
Oligonucleotide Synthesis
Why is it difficult to make oligos longer than 200nt via direct synthesis? I didn´t find the aswer but I found this: phosphoramidite synthesis struggles beyond ~150–200 nt because stepwise inefficiency and side reactions make long, error‑free chains very rare. Filges, S., Mouhanna, P., & Ståhlberg, A. (2021). Digital Quantification of Chemical Oligonucleotide Synthesis Errors. Clinical chemistry. https://doi.org/10.1093/clinchem/hvab136.
Why can’t you make a 2000bp gene via direct oligo synthesis?
If I understand correctly, it is more efficient to work based on the Twist Silicon Platform, which can produce 9,600 genes. but not sure
-> Question by George Church´s Presentation <-
[Using Google & Prof. Church’s slide #4] What are the 10 essential amino acids in all animals and how does this affect your view of the “Lysine Contingency”?
Across vertebrates, most mammals (including humans) require nine essential amino acids: histidine, isoleucine, leucine, lysine, methionine, phenylalanine, threonine, tryptophan, valine Biochemically, a “lysine contingency” (as in Jurassic Park) is plausible only in the trivial sense that any essential amino acid could serve as a single‑point nutritional dependency. Lysine is important and often limiting, but it is not uniquely essential compared with the rest of the conserved EAA set, and species differ enough that no single “10‑EAA rule” applies universally. Mccann, J., & Rawls, J. (2023). Essential Amino Acid Metabolites as Chemical Mediators of Host-Microbe Interaction in the Gut.. Annual review of microbiology. https://doi.org/10.1146/annurev-micro-032421-111819.