Week 2 HW : Benchling and In-silico Gel Art
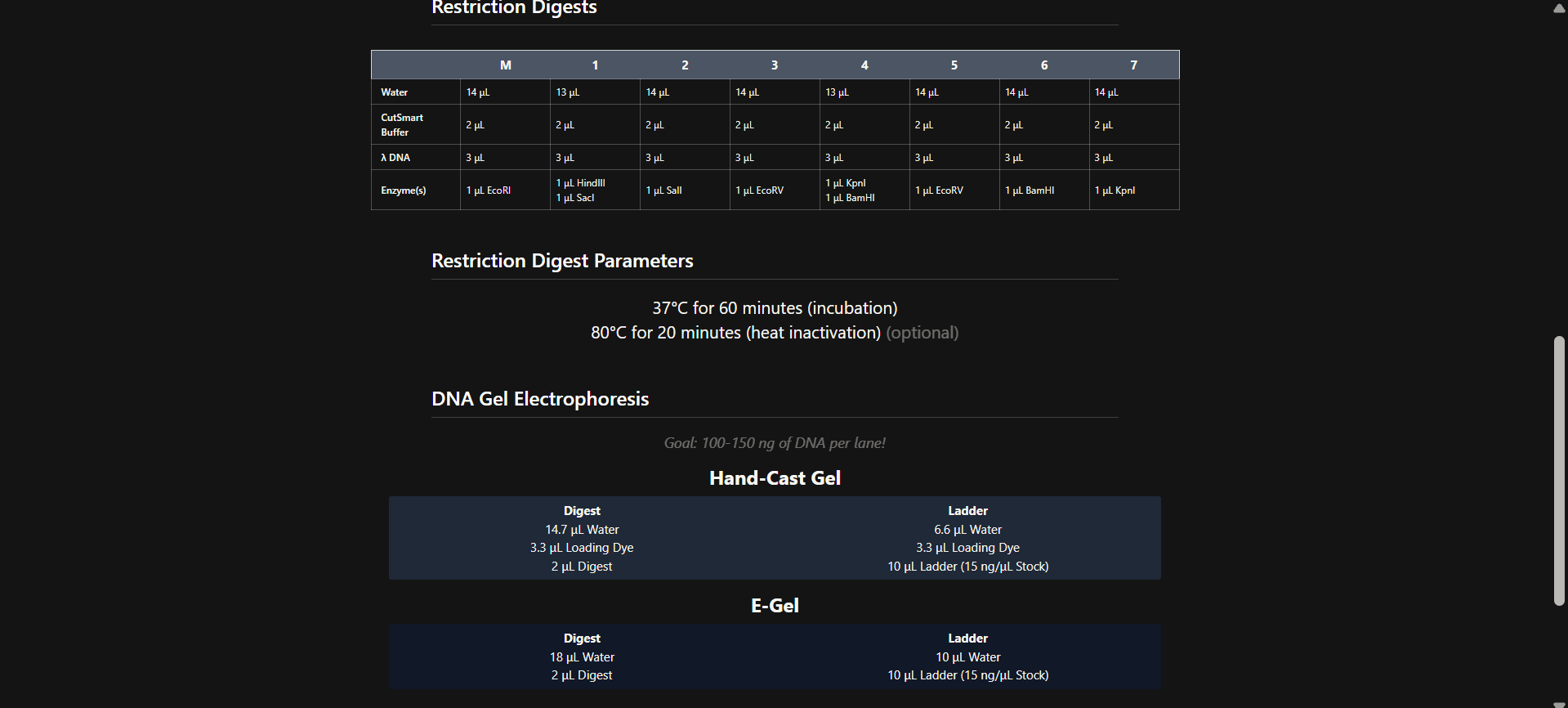
Part 1: Benchling & In-silico Gel Art
step 01 : I have already made my benchling account

step 02 : copy lambda_NEB sequence and import the sequence in my benchling
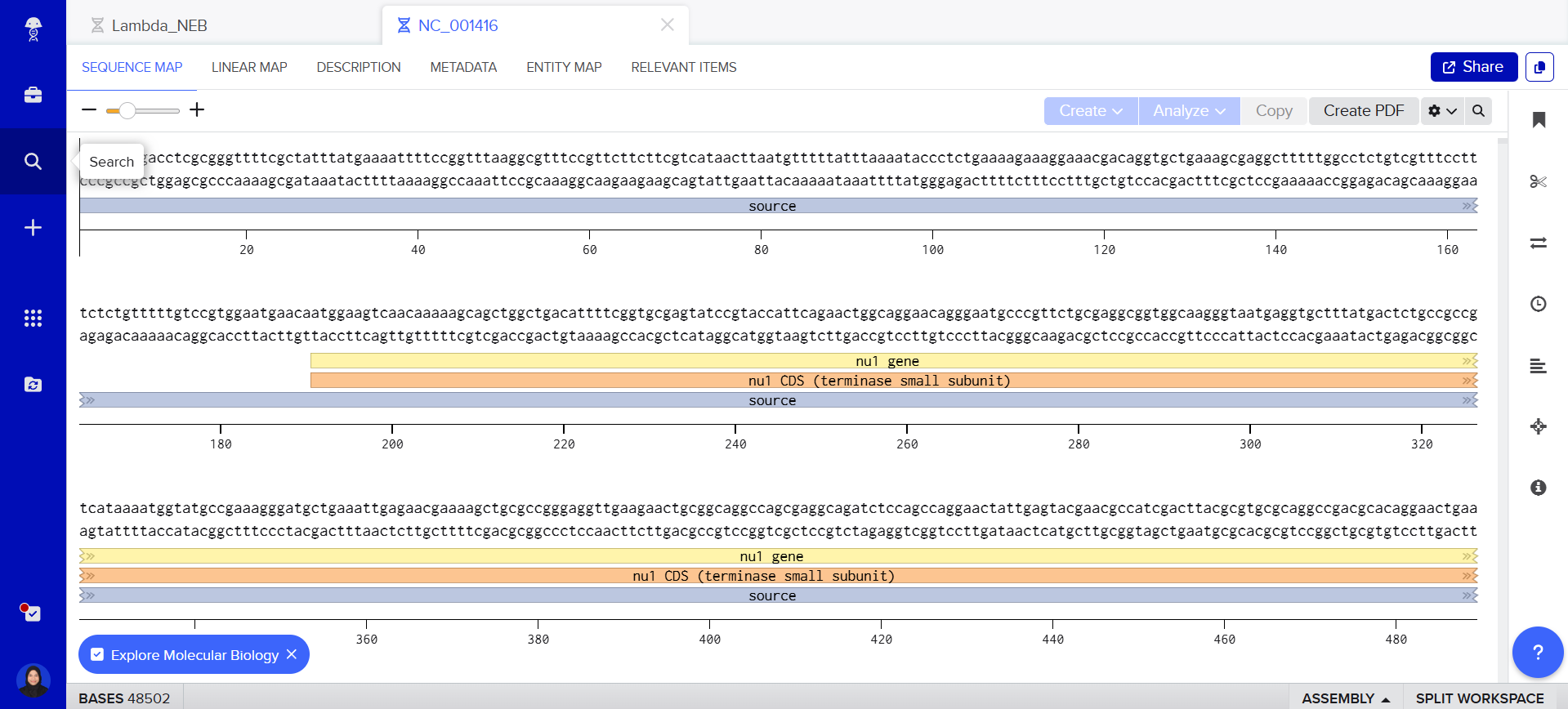

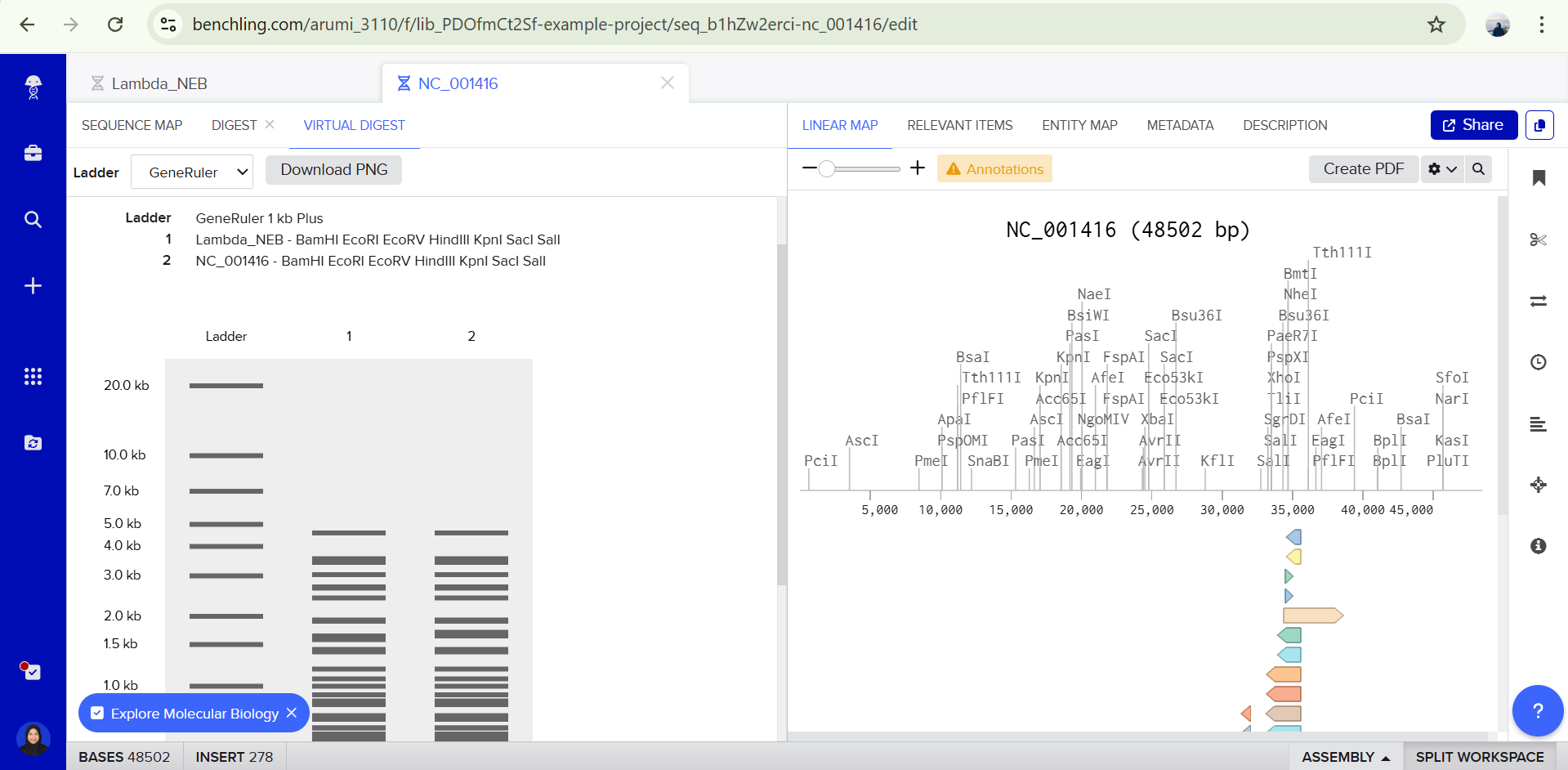

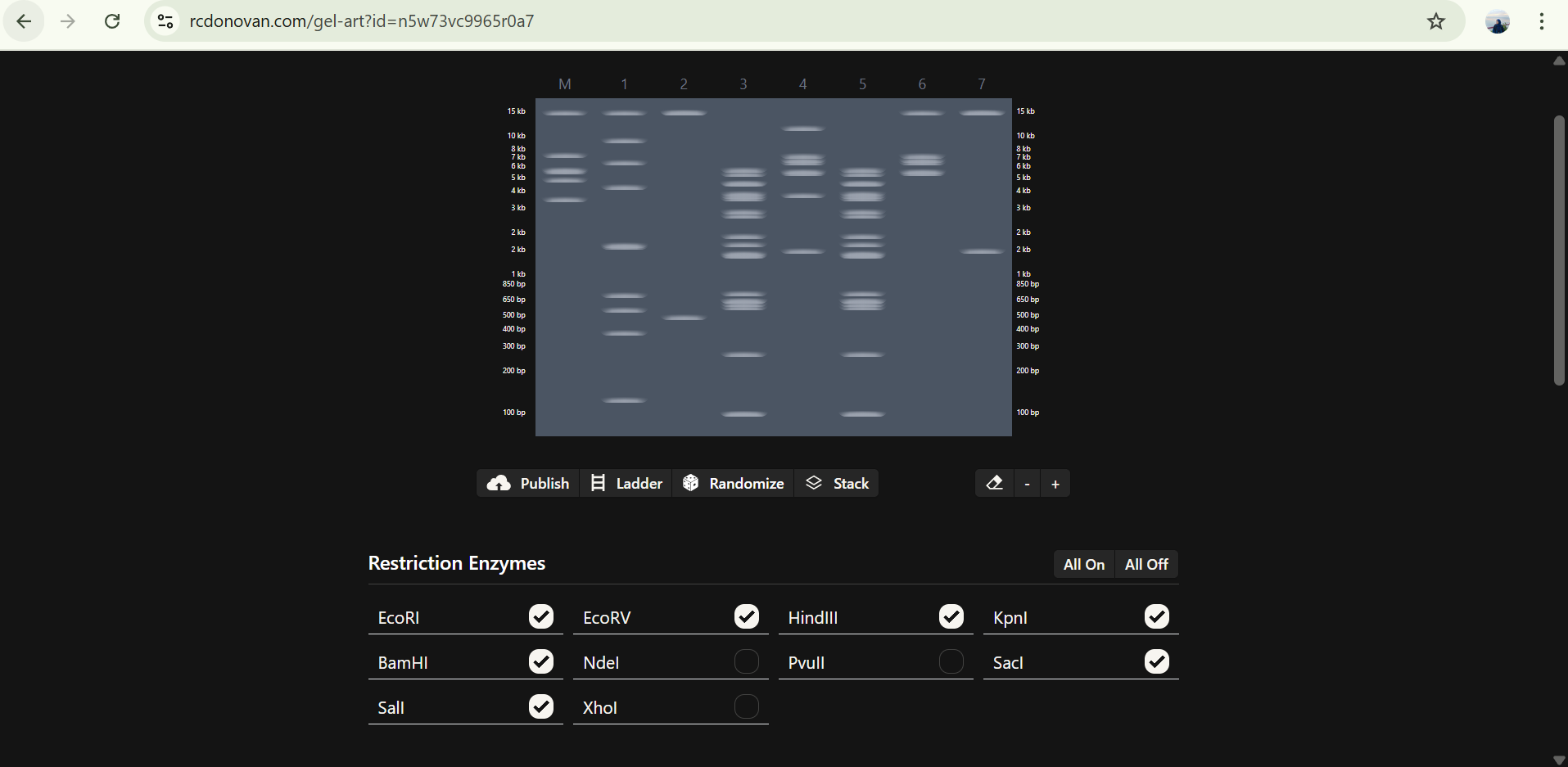


Part 3: DNA Design Challenge
my protein data from –> https://www.uniprot.org 498 AA | EC=3.2.1.1 | Full=1,4-alpha-D-glucan glucanohydrolase A | amyA | Aspergillus awamori (Black koji mold) | UniProtKB:P0C1B3
MMVAWWSLFL YGLQVAAPAL AATPADWRSQ SIYFLLTDRF ARTDGSTTAT CNTADQKYCG
GTWQGIIDKL DYIQGMGFTA IWITPVTAQL PQTTAYGDAY HGYWQQDIYS LNENYGTADD
LKALSSALHE RGMYLMVDVV ANHMGYDGAG SSVDYSVFKP FSSQDYFHPF CFIQNYEDQT
QVEDCWLGDN TVSLPDLDTT KDVVKNEWYD WVGSLVSNYS IDGLRIDTVK HVQKDFWPGY
NKAAGVYCIG EVLDGDPAYT CPYQNVMDGV LNYPIYYPLL NAFKSTSGSM DDLYNMINTV
KSDCPDSTLL GTFVENHDNP RFASYTNDIA LAKNVAAFII LNDGIPIIYA GQEQHYAGGN
DPANREATWL SGYPTDSELY KLIASRNAIR NYAISKDTGF VTYKNWPIYK DDTTIPMRKG
TDGSQIVTIL SNKGASGDSY TLSLSGAGYT AGQQLTEVIG CTTVTVGSDG NVPVPMAGGL
PRVLYPTEKL AGSKICYG
ATGATGATGGTTGCTTGGTGGTCTTTATTTTTATATGGTTTACAAGTTGCTGCTCCTGCTTTAGCTGCTACTCCTGCTGATTGGCGTTCTCAATCTATTTATTTTTTATTAACTGATCGTTTTGCTCGTACTGATGGTTCTACTACTGCTACTTGTAATACTGCTGATCAAAAATATTGTGGTGGTACTTGGCAAGGTATTATTGATAAATTAGATTATATTCAAGGTATGGGTTTTACTGCTATTTGGATTACTCCTGTTACTGCTCAATTACCTCAAACTACTGCTTATGGTGATGCTTATCATGGTTATTGGCAACAAGATATTTATTCTTTAAATGAAAATTATGGTACTGCTGATGATTTAAAAGCTTTATCTTCTGCTTTACATGAACGTGGTATGTATTTAATGGTTGATGTTGTTGCTAATCATATGGGTTATGATGGTGCTGGTTCTTCTGTTGATTATTCTGTTTTTAAACCTTTTTCTTCTCAAGATTATTTTCATCCTTTTTGTTTTATTCAAAATTATGAAGATCAAACTCAAGTTGAAGATTGTTGGTTAGGTGATAATACTGTTTCTTTACCTGATTTAGATACTACTAAAGATGTTGTTAAAAATGAATGGTATGATTGGGTTGGTTCTTTAGTTTCTAATTATTCTATTGATGGTTTACGTATTGATACTGTTAAACATGTTCAAAAAGATTTTTGGCCTGGTTATAATAAAGCTGCTGGTGTTTATTGTATTGGTGAAGTTTTAGATGGTGATCCTGCTTATACTTGTCCTTATCAAAATGTTATGGATGGTGTTTTAAATTATCCTATTTATTATCCTTTATTAAATGCTTTTAAATCTACTTCTGGTTCTATGGATGATTTATATAATATGATTAATACTGTTAAATCTGATTGTCCTGATTCTACTTTATTAGGTACTTTTGTTGAAAATCATGATAATCCTCGTTTTGCTTCTTATACTAATGATATTGCTTTAGCTAAAAATGTTGCTGCTTTTATTATTTTAAATGATGGTATTCCTATTATTTATGCTGGTCAAGAACAACATTATGCTGGTGGTAATGATCCTGCTAATCGTGAAGCTACTTGGTTATCTGGTTATCCTACTGATTCTGAATTATATAAATTAATTGCTTCTCGTAATGCTATTCGTAATTATGCTATTTCTAAAGATACTGGTTTTGTTACTTATAAAAATTGGCCTATTTATAAAGATGATACTACTATTCCTATGCGTAAAGGTACTGATGGTTCTCAAATTGTTACTATTTTATCTAATAAAGGTGCTTCTGGTGATTCTTATACTTTATCTTTATCTGGTGCTGGTTATACTGCTGGTCAACAATTAACTGAAGTTATTGGTTGTACTACTGTTACTGTTGGTTCTGATGGTAATGTTCCTGTTCCTATGGCTGGTGGTTTACCTCGTGTTTTATATCCTACTGAAAAATTAGCTGGTTCTAAAATTTGTTATGGTTAA
Codon optimization tools –> https://www.idtdna.com/CodonOpt
ATG ATG ATG GTT GCT TGG TGG TCC CTG TTC CTC TAT GGC TTG CAG GTA GCT GCG CCA GCG CTC GCA GCC ACC CCT GCG GAC TGG CGT TCG CAG TCT ATT TAC TTC CTT CTG ACT GAC CGG TTT GCT CGT ACT GAT GGC TCC ACA ACT GCG ACC TGT AAC ACT GCC GAC CAA AAG TAC TGC GGT GGG ACG TGG CAA GGT ATC ATT GAT AAG TTG GAT TAT ATC CAG GGA ATG GGA TTT ACT GCA ATA TGG ATT ACT CCG GTC ACC GCC CAA CTG CCT CAA ACA ACT GCC TAT GGC GAC GCA TAC CAT GGT TAC TGG CAA CAG GAT ATT TAC TCT TTG AAT GAA AAC TAC GGG ACC GCC GAT GAT CTG AAG GCA CTT TCT TCA GCT CTG CAC GAG AGA GGG ATG TAC CTC ATG GTT GAT GTT GTG GCC AAT CAC ATG GGT TAC GAC GGC GCC GGT TCG TCC GTC GAT TAC TCG GTC TTC AAG CCC TTC TCT AGT CAA GAC TAT TTC CAC CCC TTC TGC TTC ATT CAA AAC TAT GAG GAT CAG ACA CAA GTC GAA GAC TGC TGG CTT GGC GAC AAC ACC GTT TCA TTG CCT GAC CTT GAC ACG ACT AAA GAT GTA GTG AAG AAC GAG TGG TAT GAT TGG GTC GGC TCT CTG GTC TCT AAC TAT AGT ATC GAT GGT CTT CGG ATT GAT ACG GTG AAA CAC GTT CAG AAG GAC TTC TGG CCT GGT TAT AAC AAG GCA GCT GGC GTT TAC TGC ATC GGC GAA GTG CTC GAT GGA GAC CCC GCC TAT ACC TGC CCC TAT CAG AAC GTC ATG GAC GGG GTA CTC AAT TAC CCG ATA TAC TAC CCG TTG CTC AAC GCT TTC AAA TCG ACG TCC GGC TCC ATG GAC GAT CTT TAC AAT ATG ATA AAC ACA GTT AAG TCG GAT TGC CCA GAC TCA ACT CTG CTG GGG ACT TTC GTG GAG AAC CAT GAC AAC CCT CGC TTC GCT TCG TAC ACA AAC GAC ATC GCA CTG GCG AAA AAT GTC GCT GCC TTT ATC ATC CTC AAT GAC GGA ATC CCT ATT ATC TAC GCA GGC CAG GAA CAG CAT TAT GCC GGG GGC AAC GAC CCC GCT AAC AGG GAA GCT ACA TGG TTG TCA GGC TAT CCT ACT GAT AGC GAA CTC TAT AAA CTT ATC GCT TCT CGT AAC GCA ATC CGG AAC TAC GCT ATA TCT AAA GAT ACC GGG TTC GTT ACC TAT AAA AAT TGG CCC ATC TAT AAG GAT GAT ACA ACA ATC CCA ATG CGC AAA GGT ACA GAC GGT AGC CAG ATT GTG ACC ATT TTG TCC AAC AAA GGT GCA TCG GGC GAC TCG TAC ACC CTT AGT TTG AGT GGA GCA GGG TAC ACC GCC GGA CAG CAA CTG ACT GAG GTG ATC GGA TGC ACC ACC GTG ACC GTT GGA TCA GAT GGT AAC GTC CCA GTT CCT ATG GCT GGG GGA CTG CCC CGA GTT CTT TAT CCA ACA GAA AAG CTC GCT GGC TCC AAG ATT TGT TAT GGT TAA
Alpha-amylase can be produced from DNA using recombinant DNA technology in either living cells (cell-dependent systems) or cell-free protein synthesis systems. In a cell-dependent method, the gene encoding alpha-amylase is first isolated—often from organisms such as Aspergillus species—and amplified using PCR. The gene is then inserted into a plasmid vector containing regulatory elements like a promoter, ribosome binding site, and terminator. This recombinant plasmid is introduced into a host organism such as Escherichia coli, Bacillus subtilis, or yeast. Once inside the host cell, the gene is expressed when the promoter is activated. The cells are grown in culture, often in a bioreactor, where they produce the alpha-amylase protein. The enzyme can then be harvested and purified. Alternatively, in a cell-free system, the DNA template is added directly into a reaction mixture containing ribosomes, enzymes, nucleotides, amino acids, and energy sources extracted from cells. In this system, transcription and translation occur in vitro without the need for living cells, allowing rapid protein production.
At the molecular level, the DNA sequence encoding alpha-amylase is first transcribed into messenger RNA (mRNA) by RNA polymerase. The enzyme binds to the promoter region of the gene, unwinds the DNA, and synthesizes a complementary mRNA strand using base-pairing rules. Once transcription is complete, the mRNA molecule is used as a template for translation. During translation, a ribosome binds to the mRNA and reads its nucleotide sequence in groups of three bases called codons. Each codon corresponds to a specific amino acid, which is delivered by transfer RNA (tRNA). The ribosome links these amino acids together through peptide bonds, forming a growing polypeptide chain. When a stop codon is reached, translation ends and the newly synthesized polypeptide is released. The chain then folds into its functional three-dimensional structure, forming active alpha-amylase.
Part 4: Prepare a Twist DNA Synthesis Order
I have already made my twist and benchling account, and then I build my DNA insert sequence to benchling.
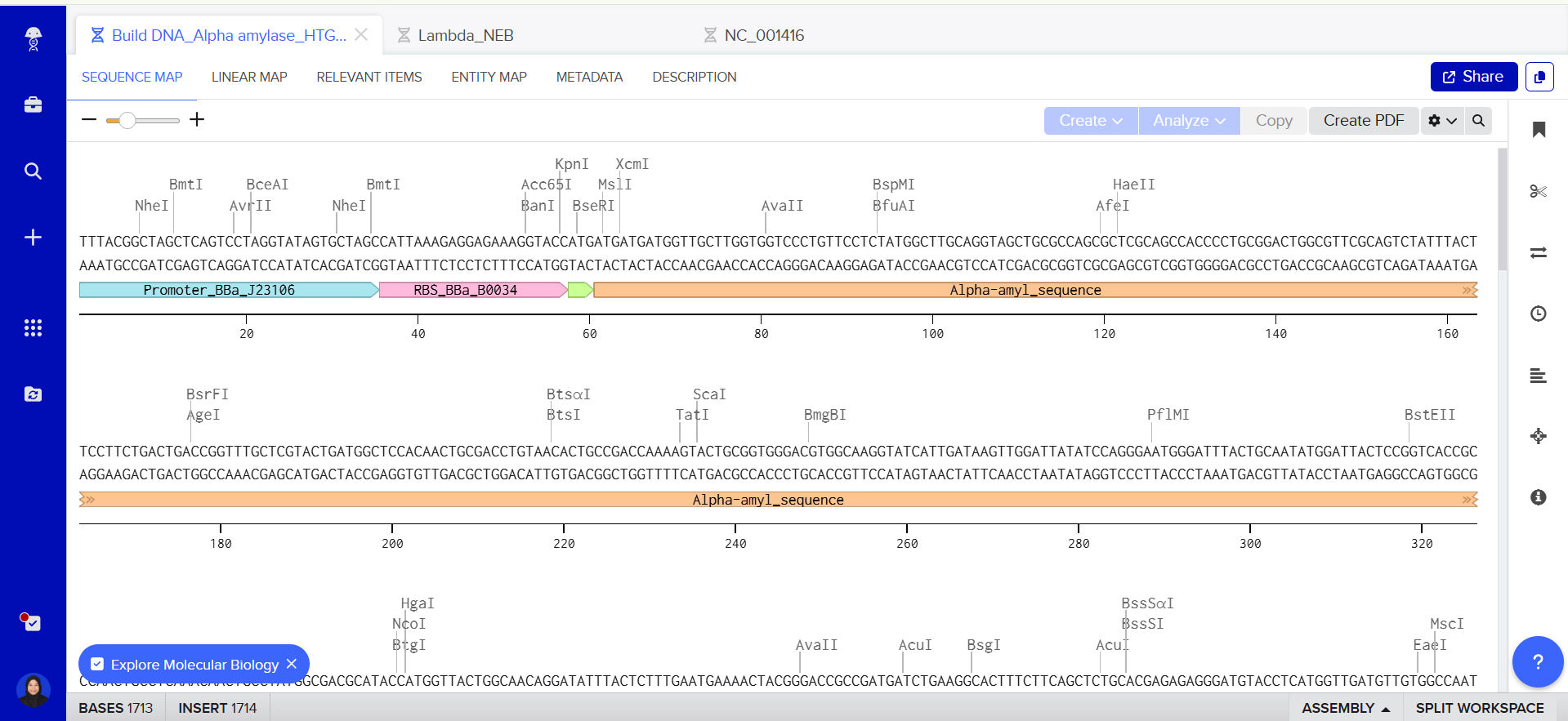

https://benchling.com/s/seq-mdL31ybjutsY3FzOCXG2?m=slm-g7IcxDwcORdvoq2oilFT
for visualize DNA I can use this website too –> https://sbolcanvas.org/
This is my plasmid, I just built with my expression cassette included! <3
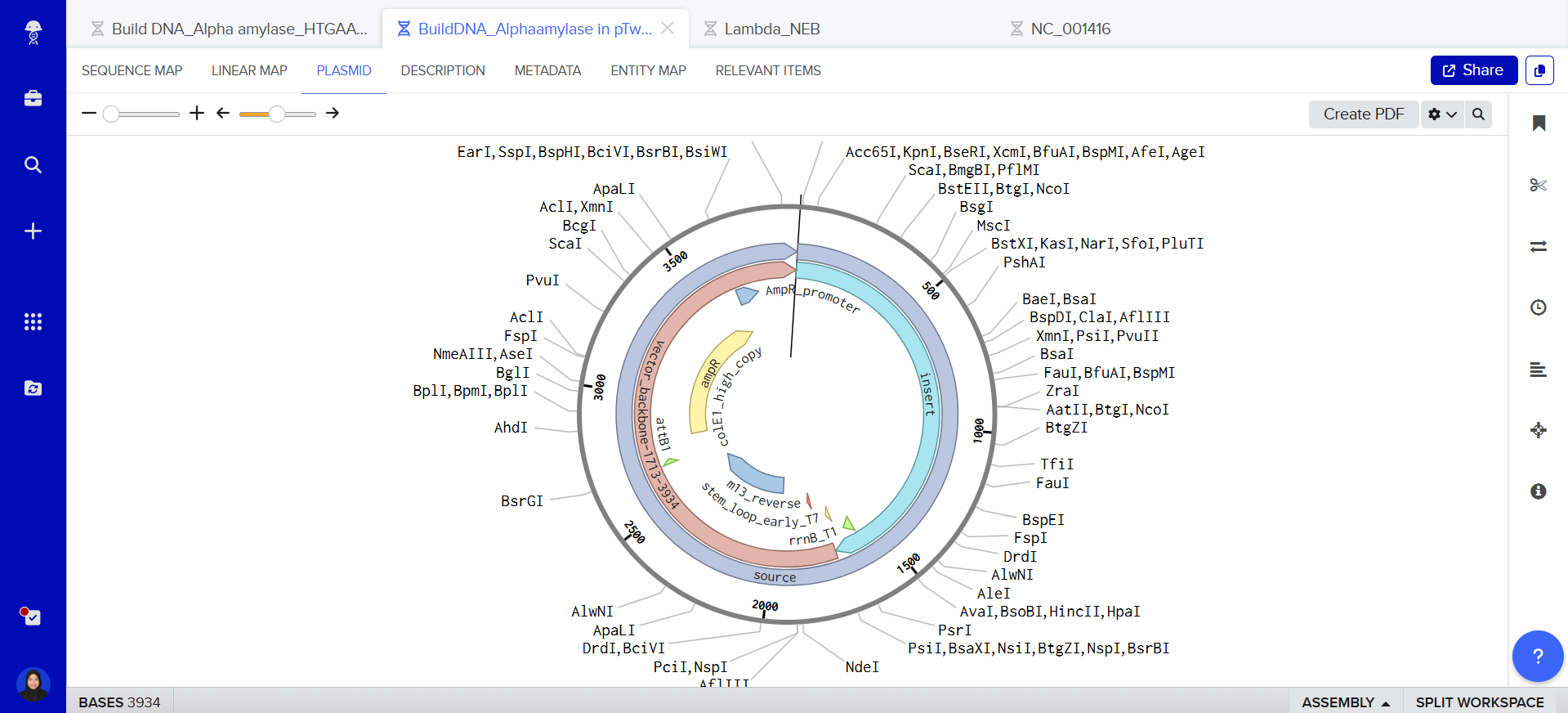

Part 5: DNA Read/Write/Edit
DNA READ
(i) What DNA would you want to sequence (e.g., read) and why?
(ii) In lecture, a variety of sequencing technologies were mentioned. What technology or technologies would you use to perform sequencing on your DNA and why?
DNA WRITE
(i) What DNA would you want to synthesize (e.g., write) and why?
(ii) What technology or technologies would you use to perform this DNA synthesis and why? Also answer the following questions:
What are the essential steps of your chosen sequencing methods?
What are the limitations of your sequencing method (if any) in terms of speed, accuracy, scalability?
DNA EDIT
(i) What DNA would you want to edit and why?
(ii) What technology or technologies would you use to perform these DNA edits and why?