Week 2 HW: DNA read, write and edit
Part One
Benchling & In-silico Gel Art!
this was a very intersing journy, starting with Ronan’s website to get some inspiration
i was very unsaure of the type of artpiece i want to make, and even with a lot of inspiration, my eyes feel on the most famous trend for 2025, the 67 meme!
so i got to work,
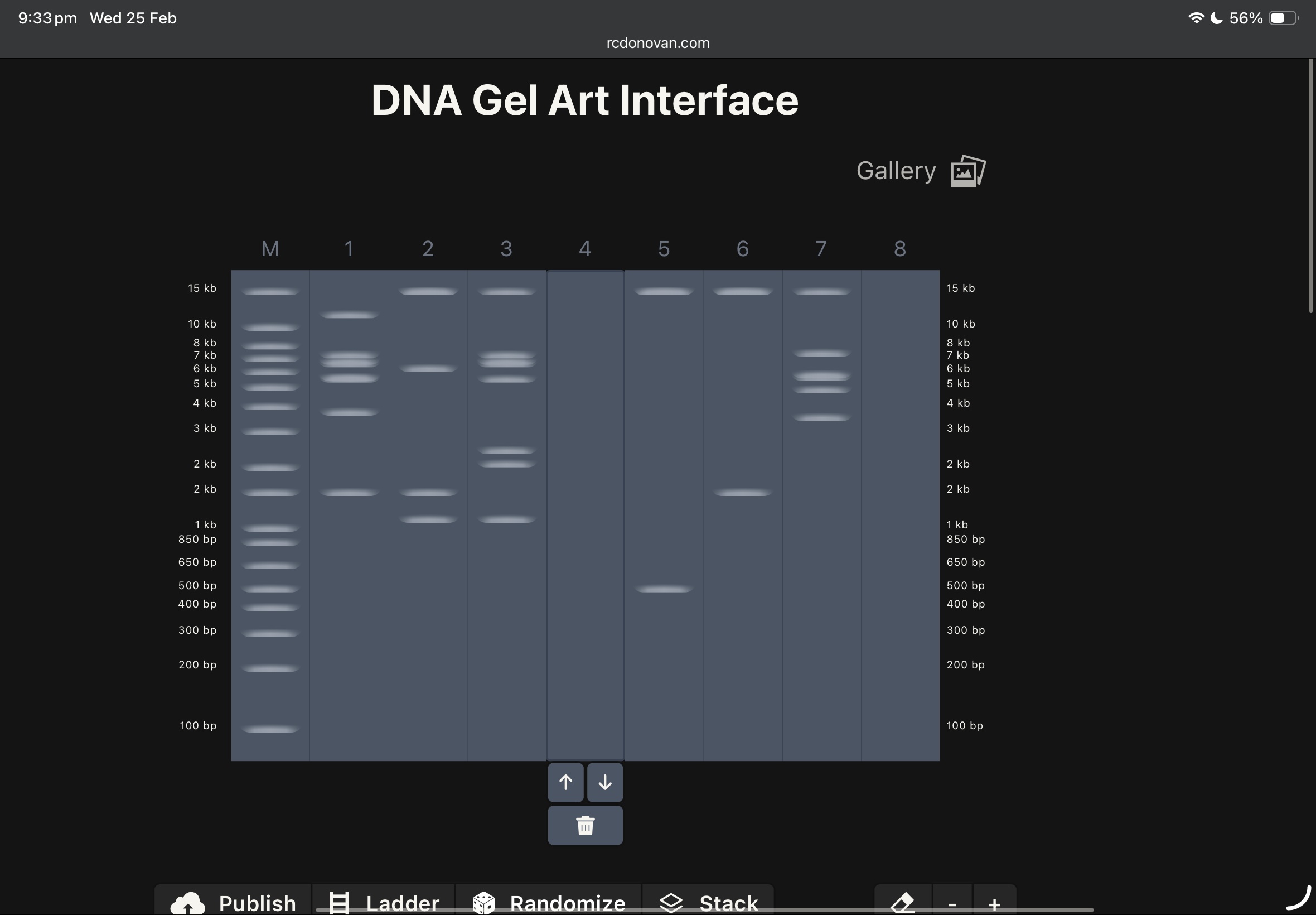
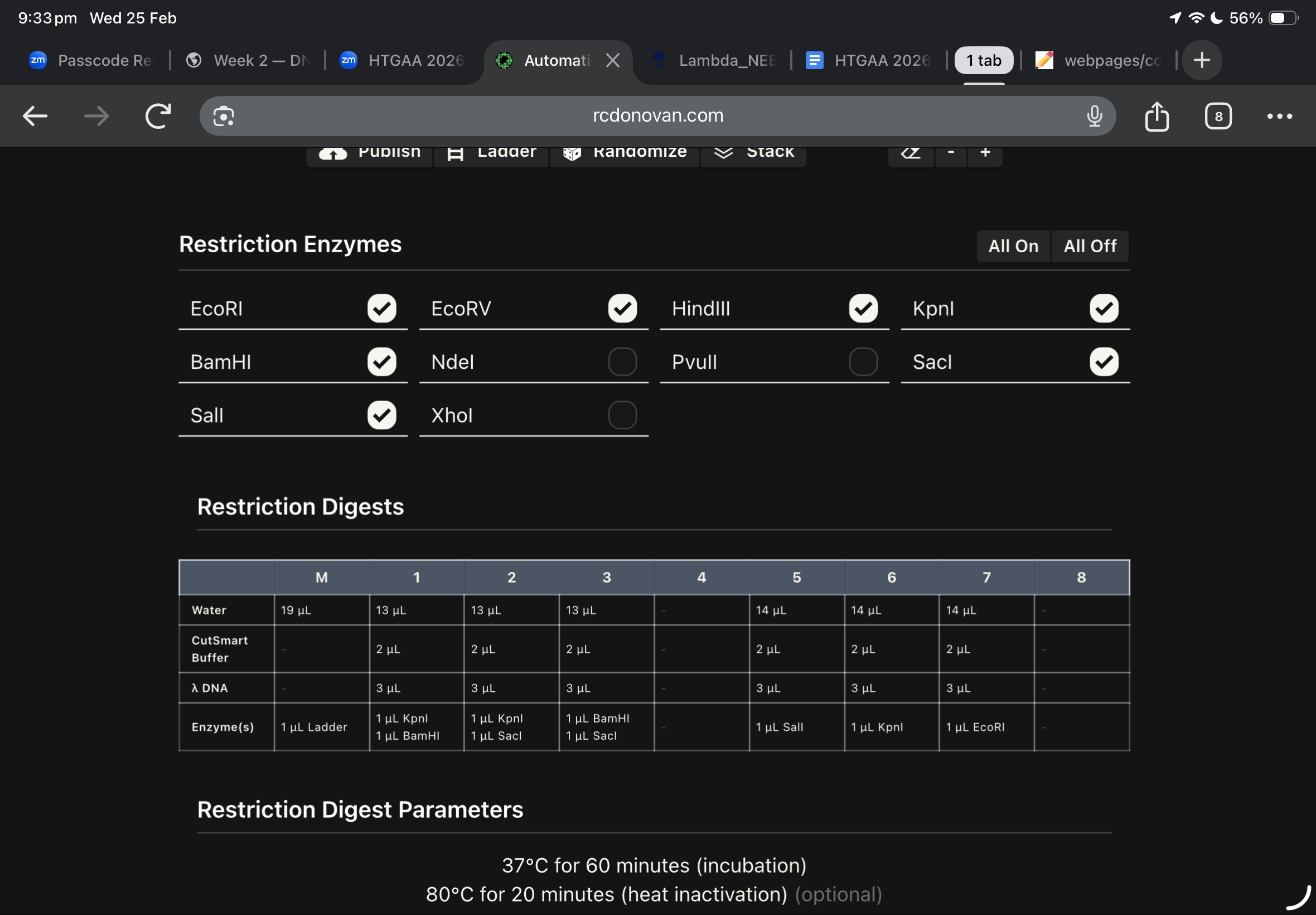
 this was my first draft using ronans website, and after checking the website enzymes used, i found these below
this was my first draft using ronans website, and after checking the website enzymes used, i found these below

Then, i moved to benchling to start a virtual digest, and i added each based on what i learned from ronans website, the final result were amazing!
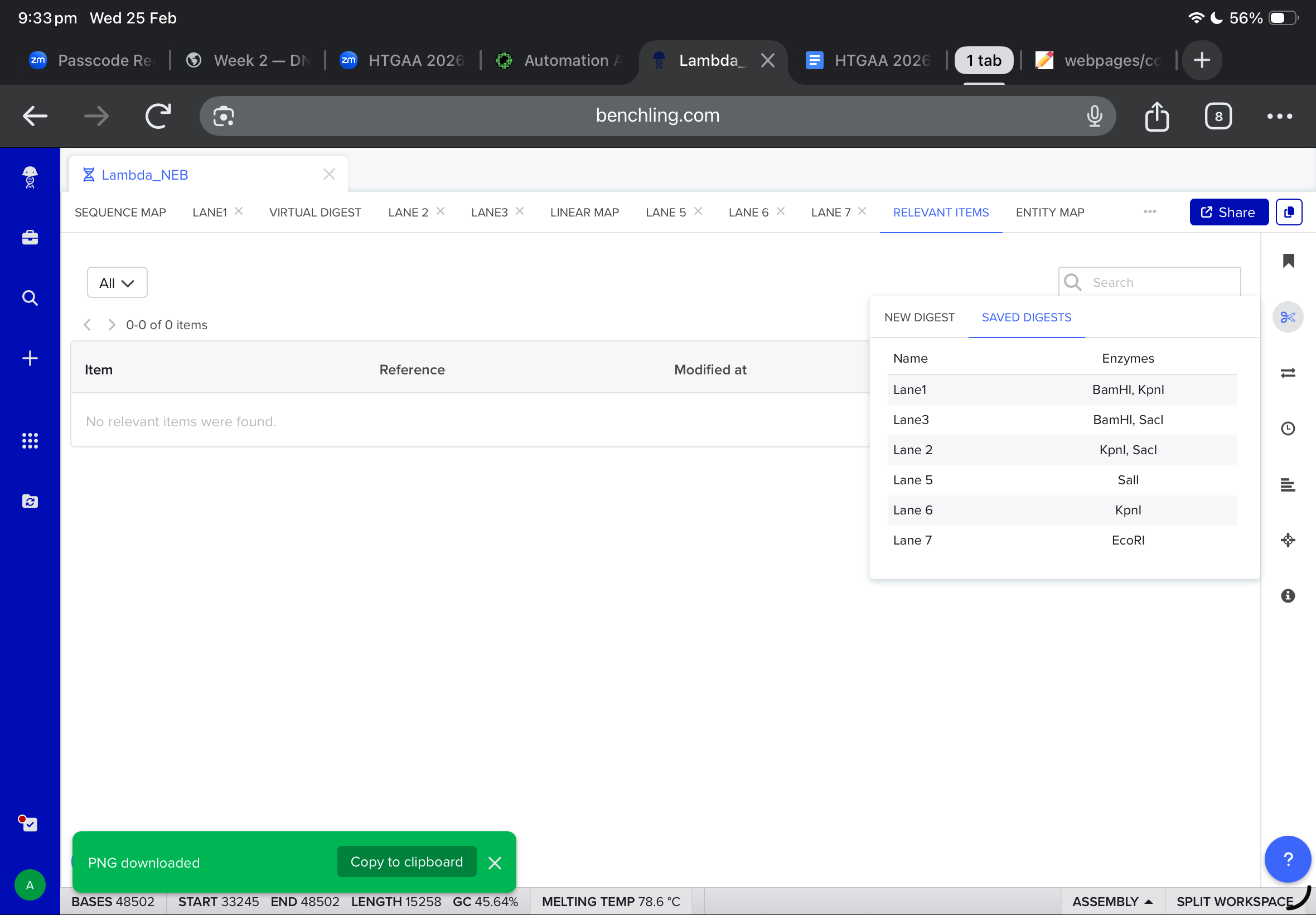

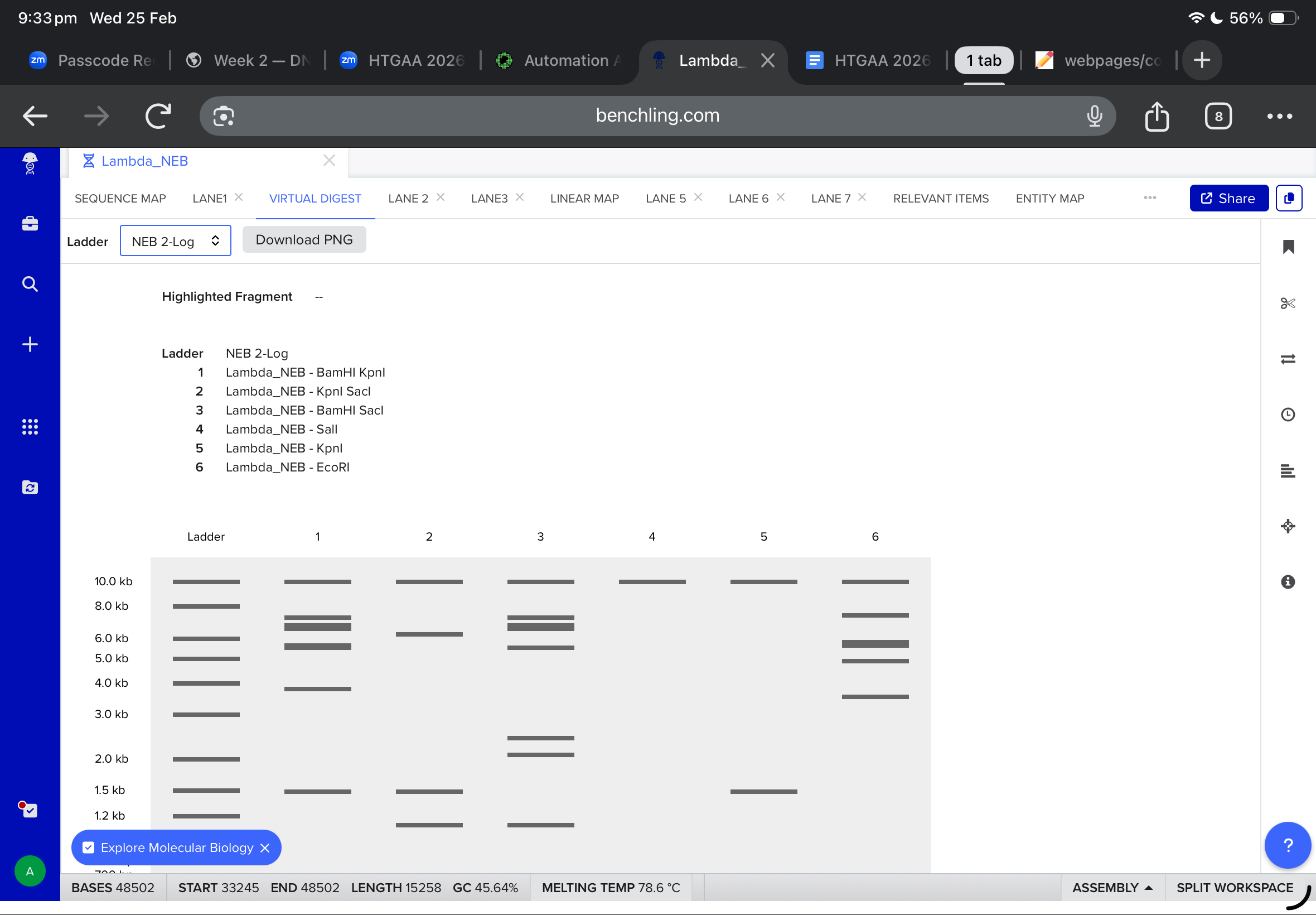

Part Two The protein i chose was lactones synthase. Which produces nice smelling compounds by converting lipids into lactones. It is used in bacteria to produces biofilms that protect them from the enviroment. The amino acids sequences i got after googling was the following:
sp|P74945|VANI_VIBAN Acyl-homoserine-lactone synthase OS=Vibrio anguillarum OX=55601 GN=vanI PE=3 SV=1 MTISIYSHTFQSVPQADYVSLLKLRYKVFSQRLQWELKTNRGMETDEYDVPEAHYLYAKEEQGHLVGCWRILPTTSRYMLKDTFSELLGVQQAPKAKEIYELSRFAVDKDHSAQLGGVSNVTLQMFQSLYHHAQQYHINAYVTVTSASVEKLIKRMGIPCERLGDKKVHLLGSTRSVALHIPMNEAYRASVNA
This is the amino acid sequence from gram negative bacteria species. After reverse translation using bioinformatics website, the DNA sequence was obtained ATGACCATTAGCATTTATAGCCATACCTTTCAGAGCGTGCCGCAGGCGGATTATGTGAGCCTGCTGAAACTGCGCTATAAAGTGTTTAGCCAGCGCCTGCAGTGGGAACTGAAAACCAACCGCGGCATGGAAACCGATGAATATGATGTGCCGGAAGCGCATTATCTGTATGCGAAAGAAGAACAGGGCCATCTGGTGGGCTGCTGGCGCATTCTGCCGACCACCAGCCGCTATATGCTGAAAGATACCTTTAGCGAACTGCTGGGCGTGCAGCAGGCGCCGAAAGCGAAAGAAATTTATGAACTGAGCCGCTTTGCGGTGGATAAAGATCATAGCGCGCAGCTGGGCGGCGTGAGCAACGTGACCCTGCAGATGTTTCAGAGCCTGTATCATCATGCGCAGCAGTATCATATTAACGCGTATGTGACCGTGACCAGCGCGAGCGTGGAAAAACTGATTAAACGCATGGGCATTCCGTGCGAACGCCTGGGCGATAAAAAAGTGCATCTGCTGGGCAGCACCCGCAGCGTGGCGCTGCATATTCCGATGAACGAAGCGTATCGCGCGAGCGTGAACGCG This is the DNA Sequence of the Enzyme.
Codon optimisation is crucial to insure higher yield of the protein of interest in a specific organism. Specific codons are more available and more likely to be translated in certain organisms than others, with multiple codons coding for the same amino acids, it is better to ensure you have the Most predominated sequence.
The optimised sequence for E. Coli was acquired and i got this ATGACCATTTCAATCTACAGCCATACTTTTCAGAGCGTGCCGCAGGCCGATTATGTGAGCCTGCTGAAACTGCGCTACAAAGTGTTTAGCCAGCGCCTGCAGTGGGAGCTGAAAACCAACCGCGGCATGGAAACCGATGAATACGACGTGCCGGAAGCGCATTATCTGTACGCGAAAGAAGAGCAGGGCCATCTGGTGGGTTGCTGGCGCATTCTGCCGACCACCAGCCGCTACATGCTGAAAGATACCTTCAGCGAACTGCTGGGCGTGCAGCAGGCGCCGAAAGCGAAAGAAATTTACGAACTGAGCCGTTTTGCCGTGGATAAAGATCATAGCGCCCAGCTGGGCGGCGTGAGCAATGTGACCCTGCAGATGTTTCAGAGCCTGTATCATCATGCGCAGCAGTATCACATCAACGCGTACGTGACCGTGACCAGCGCGAGCGTGGAAAAATTAATTAAACGCATGGGCATTCCGTGCGAACGTCTGGGCGATAAAAAAGTGCACCTGCTGGGCAGCACCCGCAGCGTGGCGCTGCATATTCCGATGAACGAAGCCTACCGCGCCAGCGTGAATGCC
the codon optimization was done using this website https://en.vectorbuilder.com/tool/codon-optimization.html
After all the work on the benchling, all annotations were added and all the promoters, terminators and tags were inserted.
the final linear map is shown below
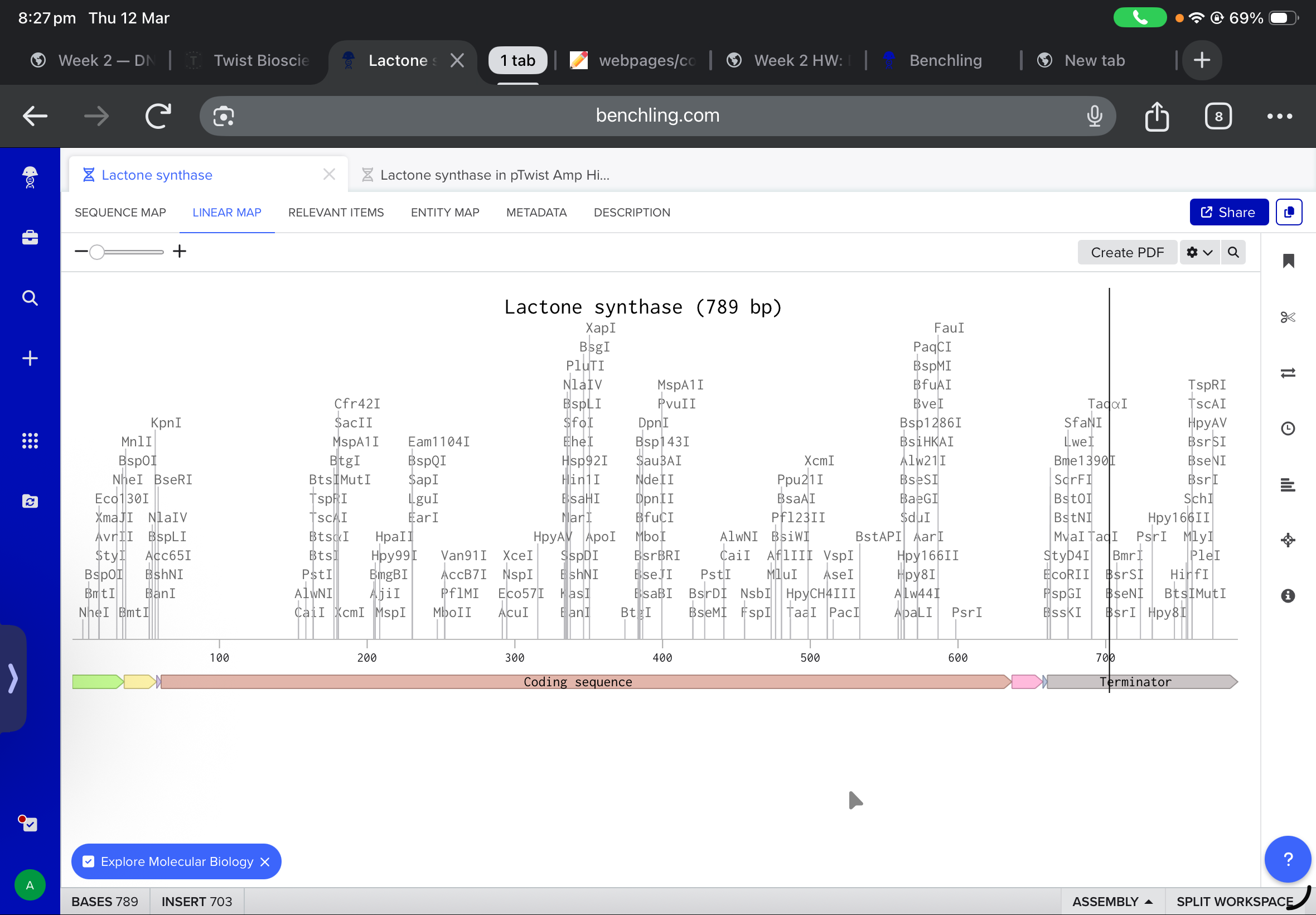

and after working with twist, and downlowing the genbank file to benchling, the final insertion plasmid is also shown below:
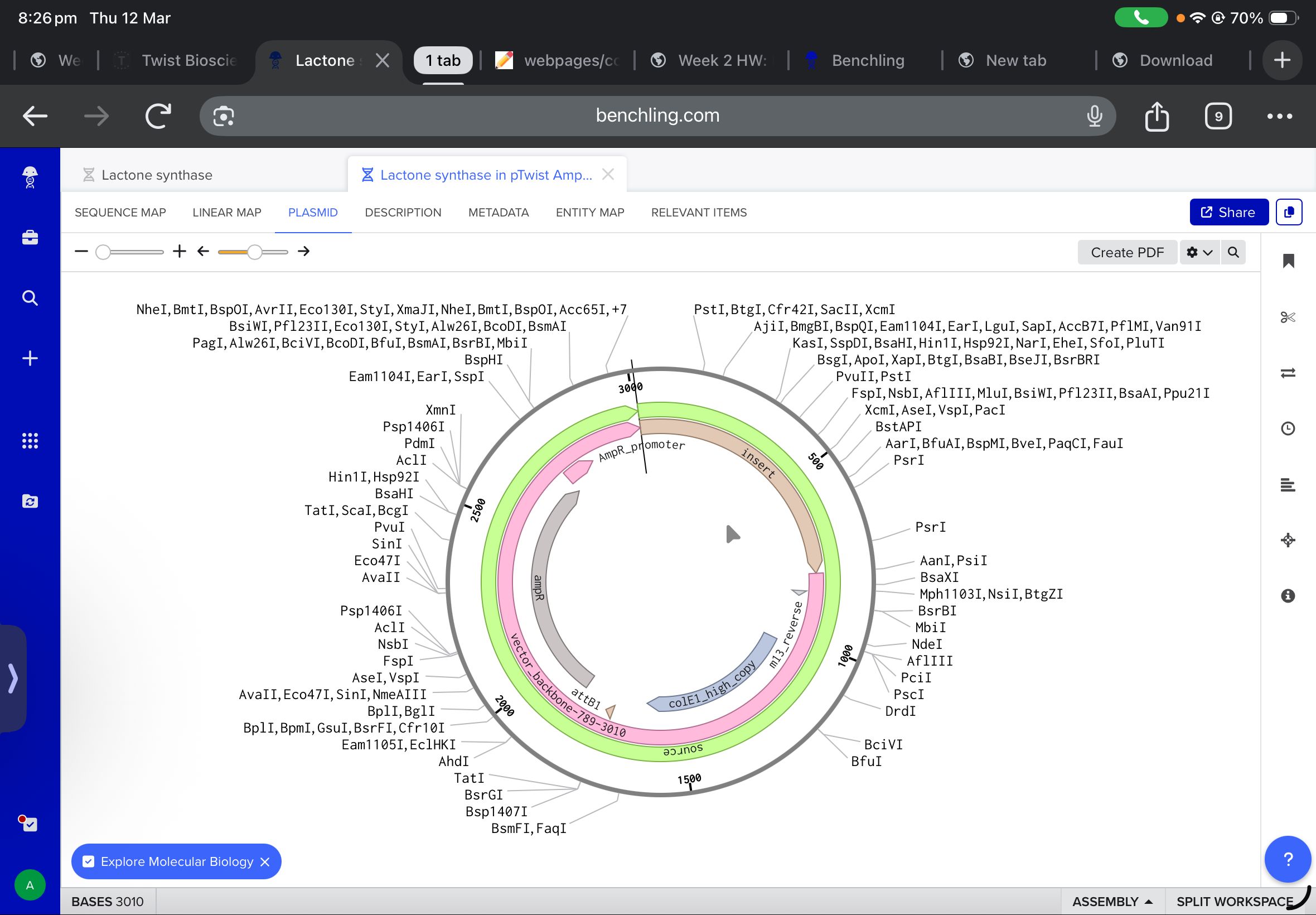

5.1 DNA READ
The DNA I would sequence is the gene responsible for telomerase activity, specifically the TERT gene. This gene encodes the catalytic subunit of telomerase, an enzyme that maintains telomere length at the ends of chromosomes. Telomeres protect chromosomes from degradation during cell division.
The sequencing method used would be Sanger sequencing, which is a first-generation sequencing technology. It determines DNA sequences using chain-terminating dideoxynucleotides during DNA synthesis.
The input for this method is genomic DNA containing the TERT gene extracted from human cells. The main preparation steps include:
- DNA extraction from the biological sample (e.g., blood, saliva, or hair follicle cells).
- PCR amplification of the TERT gene using specific primers to generate many copies of the target DNA fragment.
- PCR product purification to remove excess primers, nucleotides, and enzymes.
- Sequencing reaction preparation, where the purified DNA is mixed with DNA polymerase, normal nucleotides (dNTPs), fluorescently labeled chain-terminating nucleotides (ddNTPs), and sequencing primers.
- During sequencing, DNA polymerase extends the new DNA strand by adding nucleotides. Occasionally, a fluorescently labeled ddNTP is incorporated, which terminates the DNA strand because it lacks the 3′-OH group required for further extension. This produces DNA fragments of different lengths ending with labeled bases. The fragments are then separated using capillary electrophoresis, and a laser detects the fluorescent signals corresponding to each nucleotide. The final output is a chromatogram (electropherogram) showing colored peaks for each base (A, T, G, and C), which is converted by the sequencing software into the digital DNA sequence of the analyzed TERT gene fragment.
5.2 DNA Write
- I would synthesize a DNA construct that blocks the androgen receptor (AR) when its levels get too high. It would include an AR-responsive promoter, a coding sequence for an AR inhibitor (like a peptide or siRNA), and a terminator. This could help prevent hair loss caused by high AR activity.
2.Synthesis technology I would use Phosphoramidite DNA Synthesis (used by Twist Bioscience). It’s accurate, fast for short sequences, and allows assembly of full genes or small circuits.
- Essential steps
- Build DNA on a solid support.
- Add nucleotides in order (coupling).
- Cap incomplete strands.
- Oxidize and remove protecting groups.
- Assemble fragments into the full construct.
- Limitations
- Works best for short sequences; longer genes need assembly.
- Can be slow or expensive for large constructs.
- Small chance of errors, so verification by sequencing is needed.
5.3 DNA Edit
- I would use CRISPR‑Cas9 to edit DNA because it can target specific genes precisely. Cas9 is guided by a gRNA to a DNA sequence, then makes a cut. The cell repairs it either by: NHEJ → introduces small mutations, or HDR → uses a DNA template for exact changes.
- Preparation & input: design the gRNA, prepare Cas9 protein or plasmid, optionally include a donor DNA template, and deliver them into cells.
- Limitations: may cause off-target edits, not all cells are edited, and precise HDR edits are less efficient.