Subsections of Homework
Week 1 HW: Principles and Practices
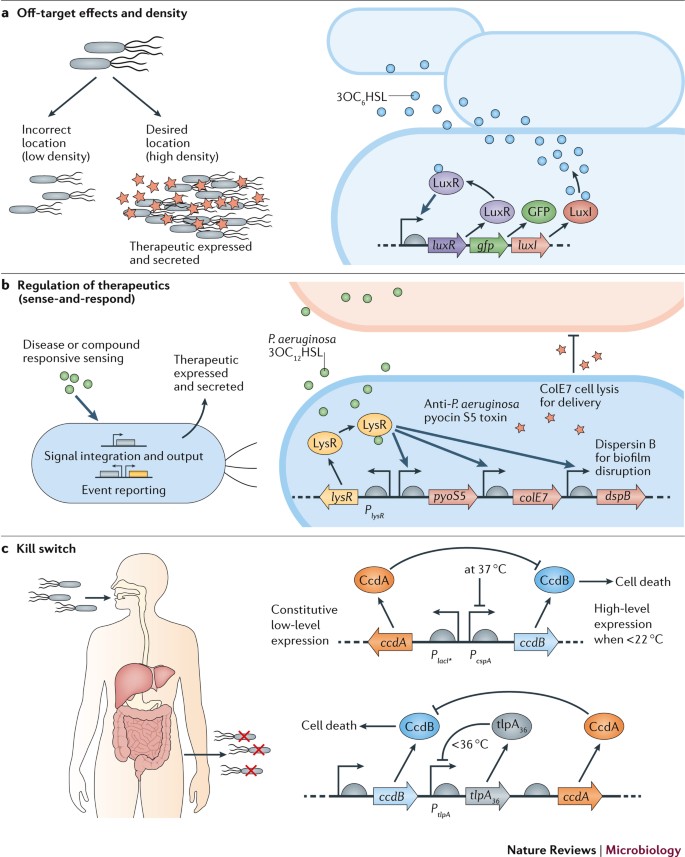

- First, describe a biological engineering application or tool you want to develop and why. This could be inspired by an idea for your HTGAA class project and/or something for which you are already doing in your research, or something you are just curious about.
Response
I’m interested in developing synthetic biology and pharmaceutical based platforms that use engineered bacteria to address major environmental and health challenges, more specifically I want to to explore the effectiveness of bacteria as therapeutic agents to prevent or treat certain type of conditions such as malaria or cancer. These engineered bacteria can live in the human as part of the normal flora but is also cheaper and less harmful as normal pharmaceutical medications.
Another part of the engineering bacteria ambitions is related to climate change and carbon footprint, as it looks synthesising bacteria that could decrease human wastes then give us oxygen and decrease carbon dioxide seems as hopeful goal and contributing to the long lasting of human civilisations and human health.
This connects my pharmaceutical field of study with synthetic biology to create solutions that are global irrelevant, beneficial and scalable.
- Next, describe one or more governance/policy goals related to ensuring that this application or tool contributes to an “ethical” future, like ensuring non-malfeasance (preventing harm). Break big goals down into two or more specific sub-goals.
Response
- Main goal: One of the primary governance goals for this application is ensuring that the engineering bacteria are used in pharmaceutical production and environmental intervention remain control safe and aligned with their public health. Throughout the life-cycle from design to testing to implementation and disposal, these engineered bacteria should have two precautions:
Subgoal Their design should be reversible and enable to persist evolving or spread outside and intended context. This include preventing the gene transfer or the uncontrolled mutations or surviving beyond defined conditions.
Subgoal Preventing that misuse risk is also important to ensure the uses of vaccines and co2 metabolism cannot be repurposed into harmful applications.
- Main goal: The second governance goal is to ensure that the benefits of the engineer bacterial system are distributed fairly and not enhancing the already existing environmental and global health inequalities.
Subgoal This can achieve by ensuring global accessibility and affordability by giving engineered bacteria locally by a lot of effect, especially for populations that are more affected with the problems of malaria or air pollution.
Subgoal Also the public trust, and transparency should be very highly valued as communication, oversight and accountability are needed to understand how and why the engineering bacteria are used in a particular environmental context.
- Next, describe at least three different potential governance “actions” by considering the four aspects below (Purpose, Design, Assumptions, Risks of Failure & “Success”).
Governance actions:
Mandatory built bio-containment for engineered bacteria:
Proposal: currently, many engineeried bacteria rely on physically engineered containment such as like labs and bio reactors, but this action focuses on requiring genetic biocontainment mechanism such as killer switches or metabolic dependencies or genetic methods for the bacteria engineered in pharmaceutical production or environmental applications.
Design: Actors: Academic researchers, biotech companies, funding agencies, regulators.
Funding agencies are regulators required by a containment future plan as a condition of approval of funding. Researchers should design organisms that cannot survive without specific laboratory or industrial conditions. Companies must document containment strategies during review and production.
Assumptions: bio-containment strategies are reliable and may remain effective overtime, and researchers can implement these mechanisms without losing functionality or effectiveness.
Risks: the containment mechanisms may fail or mutate allowing the survival and unintended environments. Or it could overly strict the requirements to produce or research or design a new innovation in academic settings.
Tiered regulatory oversight based on use case.
Proposal: engineered microbs are often regulated similar regardless of whether they are only used in contained pharmaceutical manufacturer or environmental application. This action is tiered oversize system with increasing scrutiny as a potential exposure and scale exposure.
Design: Actors: federal regulators institutional, safety committees or IBCs and the public health agencies. to lower risk, fully contained bacterial systems should face streamlined approval, other high-risk applications must require additional review testing and monitoring, and IBCs should coordinate with nation regulators for high impact deployment.
Assumptions: the risk can be reasonably categorised by application type and scale of use; regulatory agencies also have expertise and resources in such fields.
Risks: misclassification could underestimate real world risks, while complex regulatory layers may delay the deployment of urgently needed vaccines or the therapeutics.
Equity-founded access requirements:
Purpose: when novel treatments are discovered currently, the overall productions and benefits may not reach the populations most needing that treatment, this action proposes linking innovation to equitable access outcomes
Design: Actors: government global health organisations and pharmaceutical companies.
Public funding is title to avoid ability or access commitments and partnerships should be established to support global distribution and manufacturing. Also the affected communities should be supported with knowledge in order to grow and contain the bacteria within the communities.
Assumptions: the access conditions can influence how the industry and research operations can improve. Also, global partnerships can overcome the current infrastructure barriers.
Risks: access may still not be enforced overly in a fair way. or we can have lost of profits that could limit private sector participation.
| Does the option: | Mandatory built bio-containment | Tiered regulatory oversight | Equity-foundation |
|---|
| Enhance Biosecurity | 1 | 2 | 3 |
| • By preventing incidents | 1 | 2 | na |
| • By helping respond | 2 | 1 | 3 |
| Foster Lab Safety | 1 | 2 | 3 |
| • By preventing incident | 1 | 2 | na |
| • By helping respond | 2 | 1 | na |
| Protect the environment | 1 | 2 | 3 |
| • By preventing incidents | 1 | 2 | 3 |
| • By helping respond | 2 | 3 | 1 |
| Other considerations | | | |
| • Minimizing costs and burdens to stakeholders | 3 | 2 | 1 |
| • Feasibility? | 1 | 2 | 3 |
| • Not impede research | 3 | 2 | 1 |
| • Promote constructive applications | 3 | 2 | 1 |
Last, drawing upon this scoring, describe which governance option, or combination of options, you would prioritize, and why. Outline any trade-offs you considered as well as assumptions and uncertainties.
Response
Based on the scoring in the question four, I would prioritise mandatory built-in bio containment and regulatory oversight as a supporting governance mechanisms for my platform.
The bio-contanment consistently scores highest for preventing incidents across bio security, lab safety and environmental protection. This will show it as the preventative measure embedding safety directly into the engineered bacteria to reduce reliance on monitoring or human compliance. This also lowers the likelihood of irreversible harm before it occurs.
The regulatory oversight compliments this approach by scoring strongest in helping to respond to accidents while it’s less effective to prevention. It provides a good responsibility and adaptive oversight especially in higher risk contexts.
Together, these two approaches balances the front-end prevention and the backend response capacity.
However, the equity focused approach scored the lowest in safety and prevention, but I view it’s importance comes later after all the safety and security measures are already in a place that’s it’s a secondary action in promoting constructive applications.
My prioritisation assumes that genetic bio-containment is a very possible measure and that is robust and the regulators have an idea of risk tiers. The the trade-offs come in increased cost and potential friction for researchers however giving the potential consequences of failure these costs are justifiable.
These recommendations are direct toward the national funding agencies and the federal biotechnology regulators, and also the high universities who are inspiring to work in both policy making and research, especially in my country Iraq. This requires safety by design while coordination to enable creative ideas..
Week 2 pre-Lecture HW: DNA Read, Write and Edit
Questions from Professor Jacobson:
- What is the error rate of polymerase? How does this compare to the length of the human genome. How does biology deal with that discrepancy?
Response
DNA polymerases make about 1 error per 10⁶ bases during DNA synthesis when proofreading is included (≈10⁻⁶ per base). Given that the human genome is ~3.2 × 10⁹ base pairs (haploid), If replication relied only on a 10⁻⁶ error rate, each cell division would cause thousands of mutations (≈3,200 errors per replication), which would be biologically dangerous.
However, cells have multiple error reduction mechanisms: such as Polymerase proofreading (3′→5′ exonuclease activity) that removes most misincorporated bases during synthesis. And post-replicative mismatch repair (MMR) detects and fixes remaining mismatches after replication.
Also, it’s important to remember that DNA have intersting properties such diploidy, noncoding DNA and kill switches
- How many different ways are there to code (DNA nucleotide code) for an average human protein? In practice what are some of the reasons that all of these different codes don’t work to code for the protein of interest?
Response
An average human protein has about 350 amino acids, can be encoded in roughly ~10¹⁸⁰–10¹⁹⁰ different sequences of DNA, due to codon degeneracy.
Why don’t most work in practice? Mainly due to: Codon bias or tRNA availability causing inefficient translation. Could also be mRNA structure because of poor ribosome loading or elongation. Can also be GC content extremes leading to instability, synthesis and amplification problems
Questions from Dr. LeProust:
- Most commonly used oligo synthesis method?
Solid phase phosphoramidite chemical synthesis
- Why it’s hard to make oligos >200 nt directly?
Chemical synthesis is open-loop with an error rate of 1 per 10² bases. Each coupling step is less than 100% efficient, by the time we cross 200nt the yield and accuracy drop dramatically.
- Why you can’t make a 2000 bp gene by direct oligo synthesis?
Over thousands of cycles, cumulative coupling failures and errors reduce the full-length product to almost zero. Long base genes must be made by assembling many short oligos with enzymatic error correction like Gibson assembly.
Question from George Church:
- What are the 10 essential amino acids in all animals and how does this affect your view of the “Lysine Contingency”?
The 10 essential amino acids in animals that can not be synthesized by their cells and must come from diet are: Histidine, Isoleucine, Leucine, Lysine, Methionine, Phenylalanine, Threonine, Tryptophan, Valine, and Arginine all universally essential during growth and functionally essential across animals. Humans included!
Lysine is both essential and commonly limiting, especially in cereal-based and plant-heavy diets. This means that protein synthesis, growth, and health can be constrained by lysine availability even when total calories or protein intake is adequate. As a result, lysine availability disproportionately shapes nutrition, agriculture, and evolution, supporting the idea of a “lysine contingency” in which access to lysine strongly influences biological and societal outcomes.
Sources / prompts used:
Google search: “essential amino acids animals”; Lehninger Principles of Biochemistry 6th edition, introduction; Prof. George Church, HTGAA slides (#4).
Week 2 HW: DNA read, write and edit
Part One
Benchling & In-silico Gel Art!
this was a very intersing journy, starting with Ronan’s website to get some inspiration
i was very unsaure of the type of artpiece i want to make, and even with a lot of inspiration, my eyes feel on the most famous trend for 2025, the 67 meme!
so i got to work,
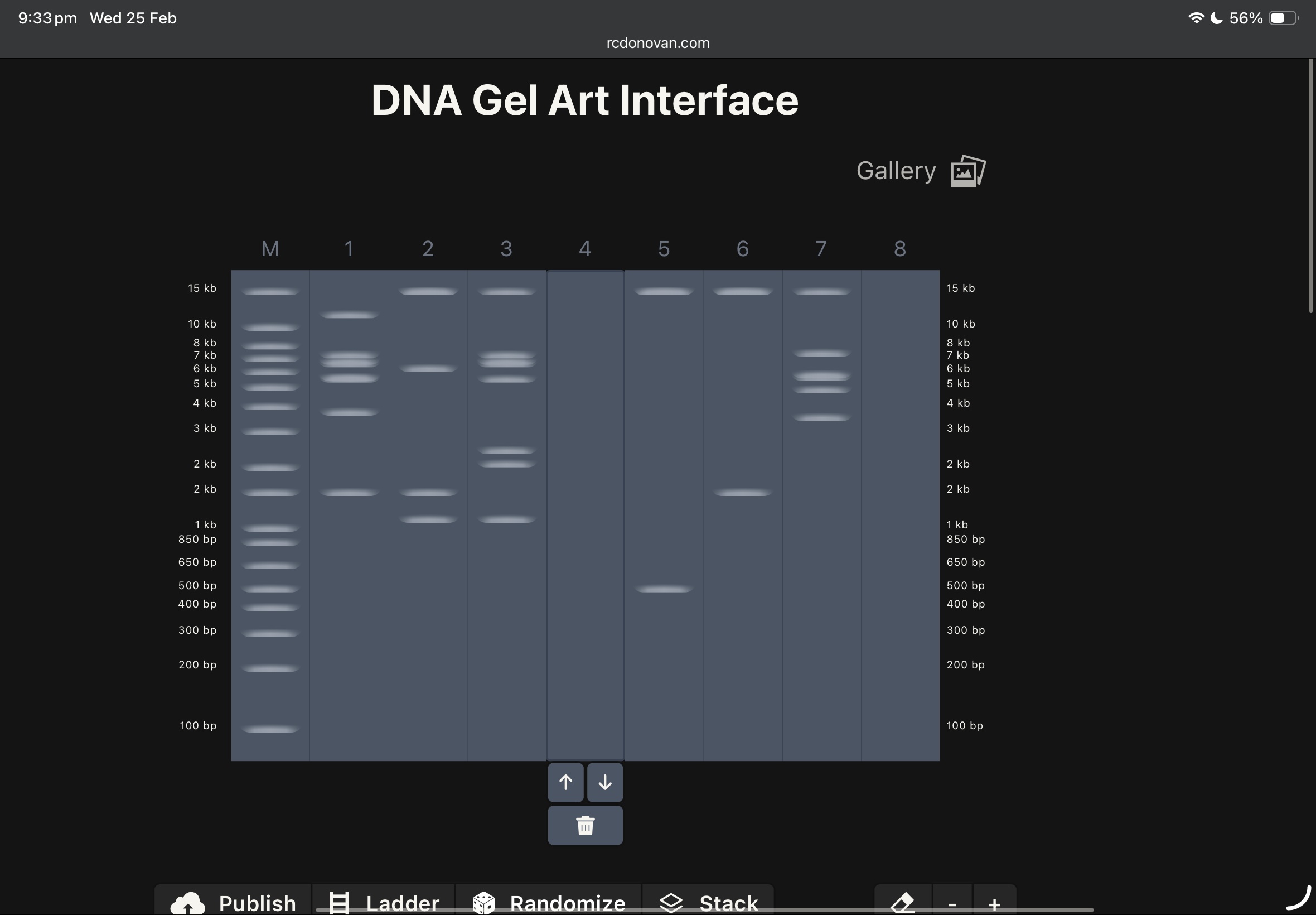
 this was my first draft using ronans website, and after checking the website enzymes used, i found these below
this was my first draft using ronans website, and after checking the website enzymes used, i found these below
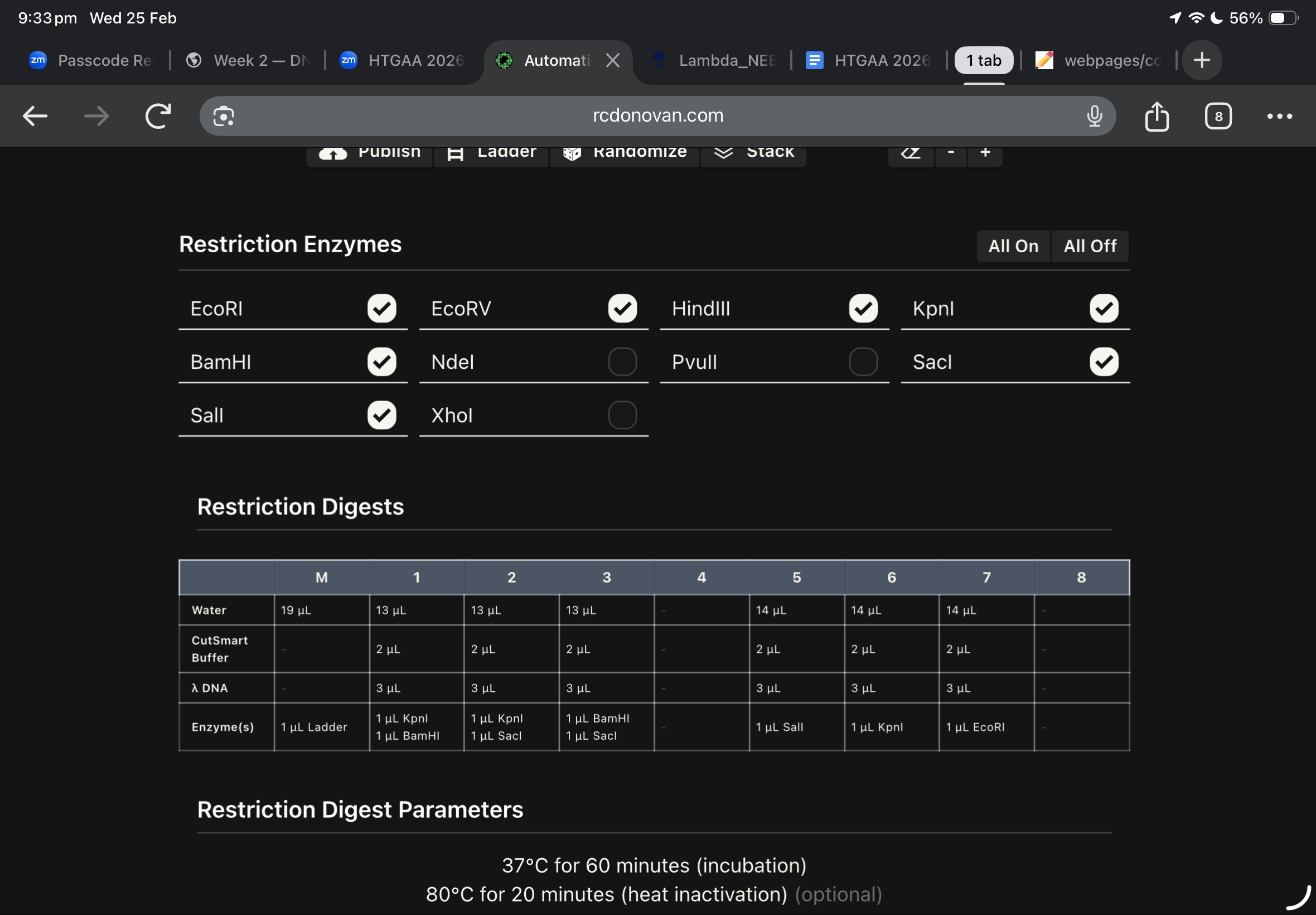

Then, i moved to benchling to start a virtual digest, and i added each based on what i learned from ronans website, the final result were amazing!
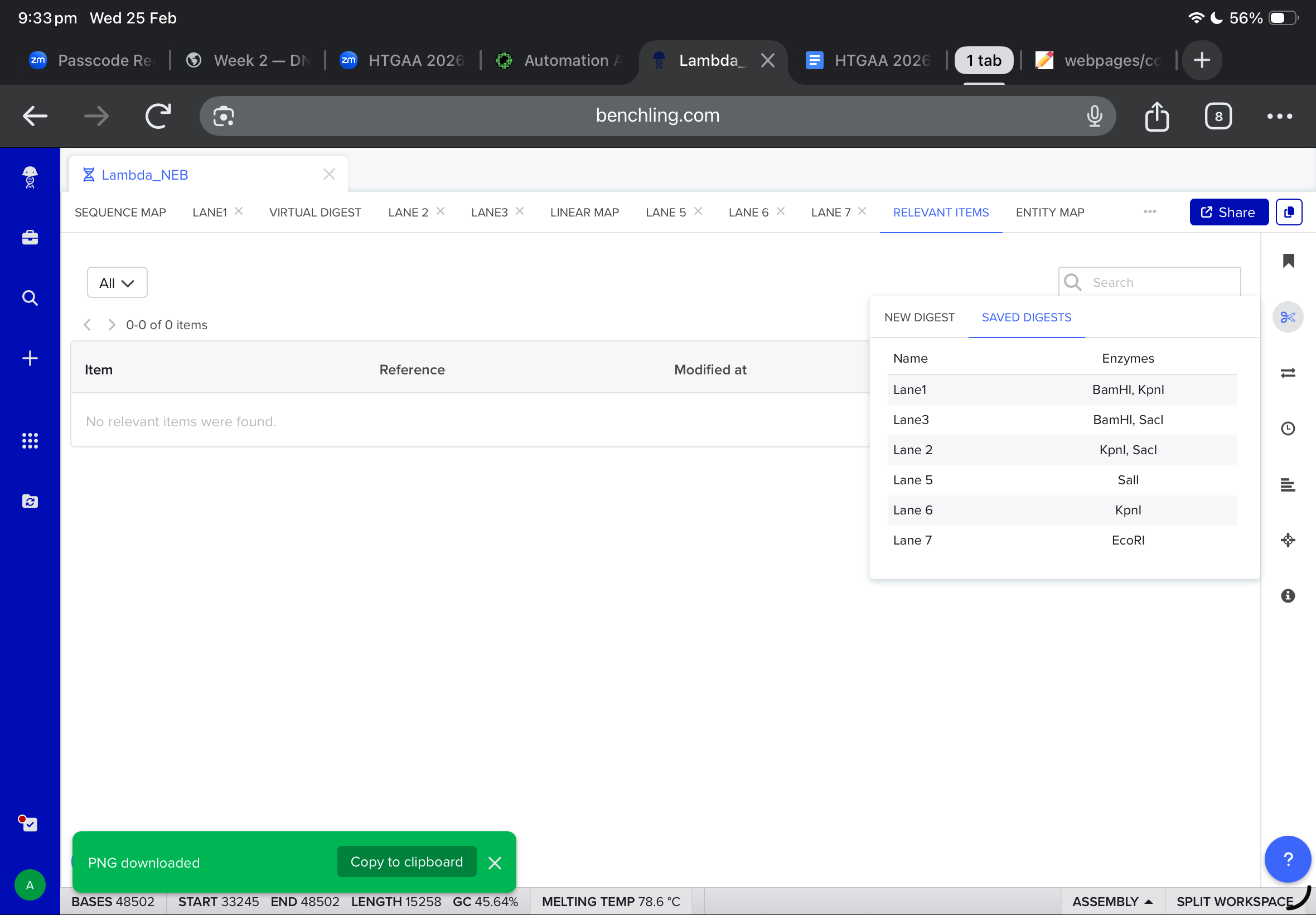

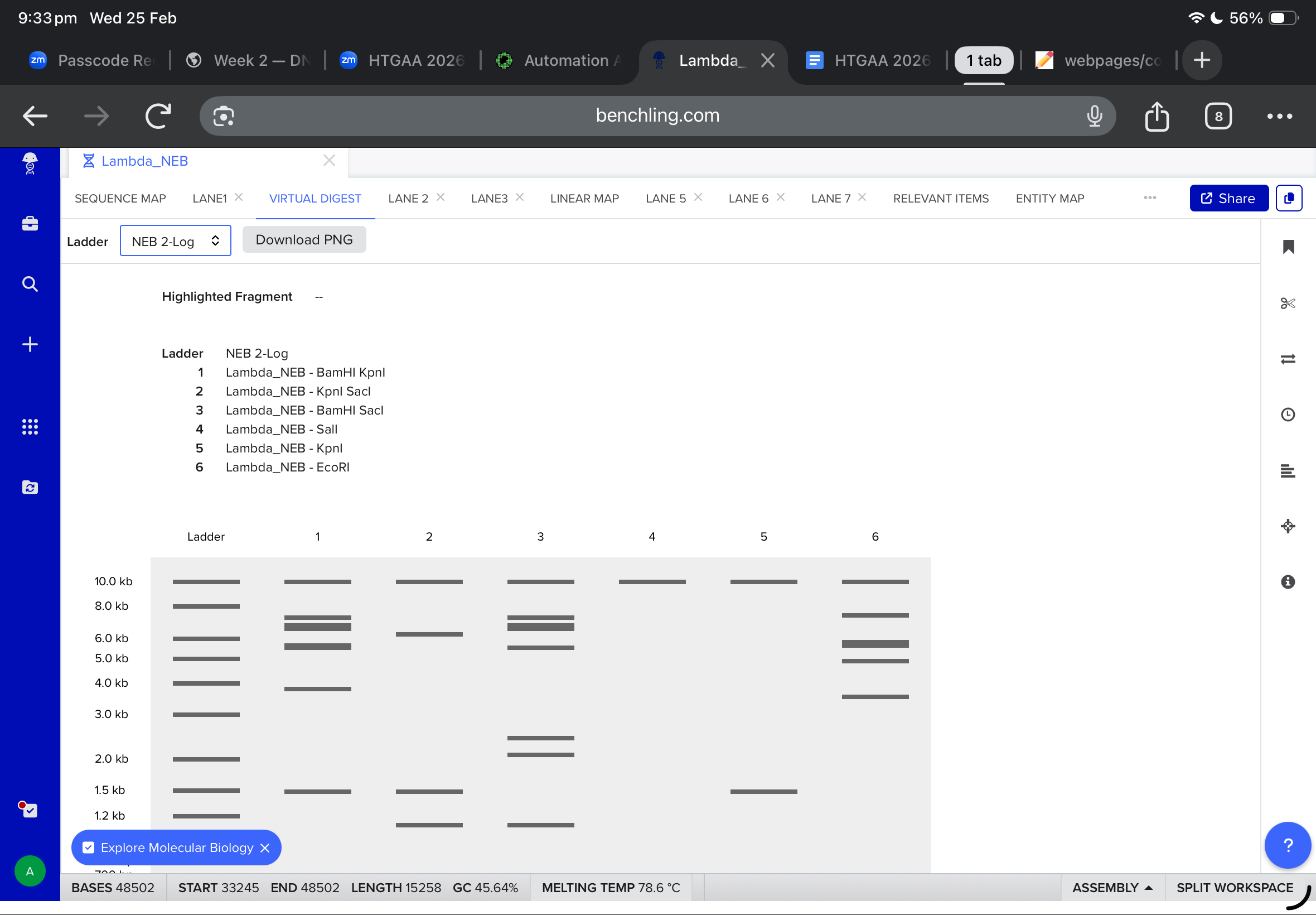

Part Two
The protein i chose was lactones synthase. Which produces nice smelling compounds by converting lipids into lactones. It is used in bacteria to produces biofilms that protect them from the enviroment.
The amino acids sequences i got after googling was the following:
sp|P74945|VANI_VIBAN Acyl-homoserine-lactone synthase OS=Vibrio anguillarum OX=55601 GN=vanI PE=3 SV=1
MTISIYSHTFQSVPQADYVSLLKLRYKVFSQRLQWELKTNRGMETDEYDVPEAHYLYAKEEQGHLVGCWRILPTTSRYMLKDTFSELLGVQQAPKAKEIYELSRFAVDKDHSAQLGGVSNVTLQMFQSLYHHAQQYHINAYVTVTSASVEKLIKRMGIPCERLGDKKVHLLGSTRSVALHIPMNEAYRASVNA
This is the amino acid sequence from gram negative bacteria species.
After reverse translation using bioinformatics website, the DNA sequence was obtained
ATGACCATTAGCATTTATAGCCATACCTTTCAGAGCGTGCCGCAGGCGGATTATGTGAGCCTGCTGAAACTGCGCTATAAAGTGTTTAGCCAGCGCCTGCAGTGGGAACTGAAAACCAACCGCGGCATGGAAACCGATGAATATGATGTGCCGGAAGCGCATTATCTGTATGCGAAAGAAGAACAGGGCCATCTGGTGGGCTGCTGGCGCATTCTGCCGACCACCAGCCGCTATATGCTGAAAGATACCTTTAGCGAACTGCTGGGCGTGCAGCAGGCGCCGAAAGCGAAAGAAATTTATGAACTGAGCCGCTTTGCGGTGGATAAAGATCATAGCGCGCAGCTGGGCGGCGTGAGCAACGTGACCCTGCAGATGTTTCAGAGCCTGTATCATCATGCGCAGCAGTATCATATTAACGCGTATGTGACCGTGACCAGCGCGAGCGTGGAAAAACTGATTAAACGCATGGGCATTCCGTGCGAACGCCTGGGCGATAAAAAAGTGCATCTGCTGGGCAGCACCCGCAGCGTGGCGCTGCATATTCCGATGAACGAAGCGTATCGCGCGAGCGTGAACGCG
This is the DNA Sequence of the Enzyme.
Codon optimisation is crucial to insure higher yield of the protein of interest in a specific organism. Specific codons are more available and more likely to be translated in certain organisms than others, with multiple codons coding for the same amino acids, it is better to ensure you have the Most predominated sequence.
The optimised sequence for E. Coli was acquired and i got this
ATGACCATTTCAATCTACAGCCATACTTTTCAGAGCGTGCCGCAGGCCGATTATGTGAGCCTGCTGAAACTGCGCTACAAAGTGTTTAGCCAGCGCCTGCAGTGGGAGCTGAAAACCAACCGCGGCATGGAAACCGATGAATACGACGTGCCGGAAGCGCATTATCTGTACGCGAAAGAAGAGCAGGGCCATCTGGTGGGTTGCTGGCGCATTCTGCCGACCACCAGCCGCTACATGCTGAAAGATACCTTCAGCGAACTGCTGGGCGTGCAGCAGGCGCCGAAAGCGAAAGAAATTTACGAACTGAGCCGTTTTGCCGTGGATAAAGATCATAGCGCCCAGCTGGGCGGCGTGAGCAATGTGACCCTGCAGATGTTTCAGAGCCTGTATCATCATGCGCAGCAGTATCACATCAACGCGTACGTGACCGTGACCAGCGCGAGCGTGGAAAAATTAATTAAACGCATGGGCATTCCGTGCGAACGTCTGGGCGATAAAAAAGTGCACCTGCTGGGCAGCACCCGCAGCGTGGCGCTGCATATTCCGATGAACGAAGCCTACCGCGCCAGCGTGAATGCC
the codon optimization was done using this website https://en.vectorbuilder.com/tool/codon-optimization.html
After all the work on the benchling, all annotations were added and all the promoters, terminators and tags were inserted.
the final linear map is shown below
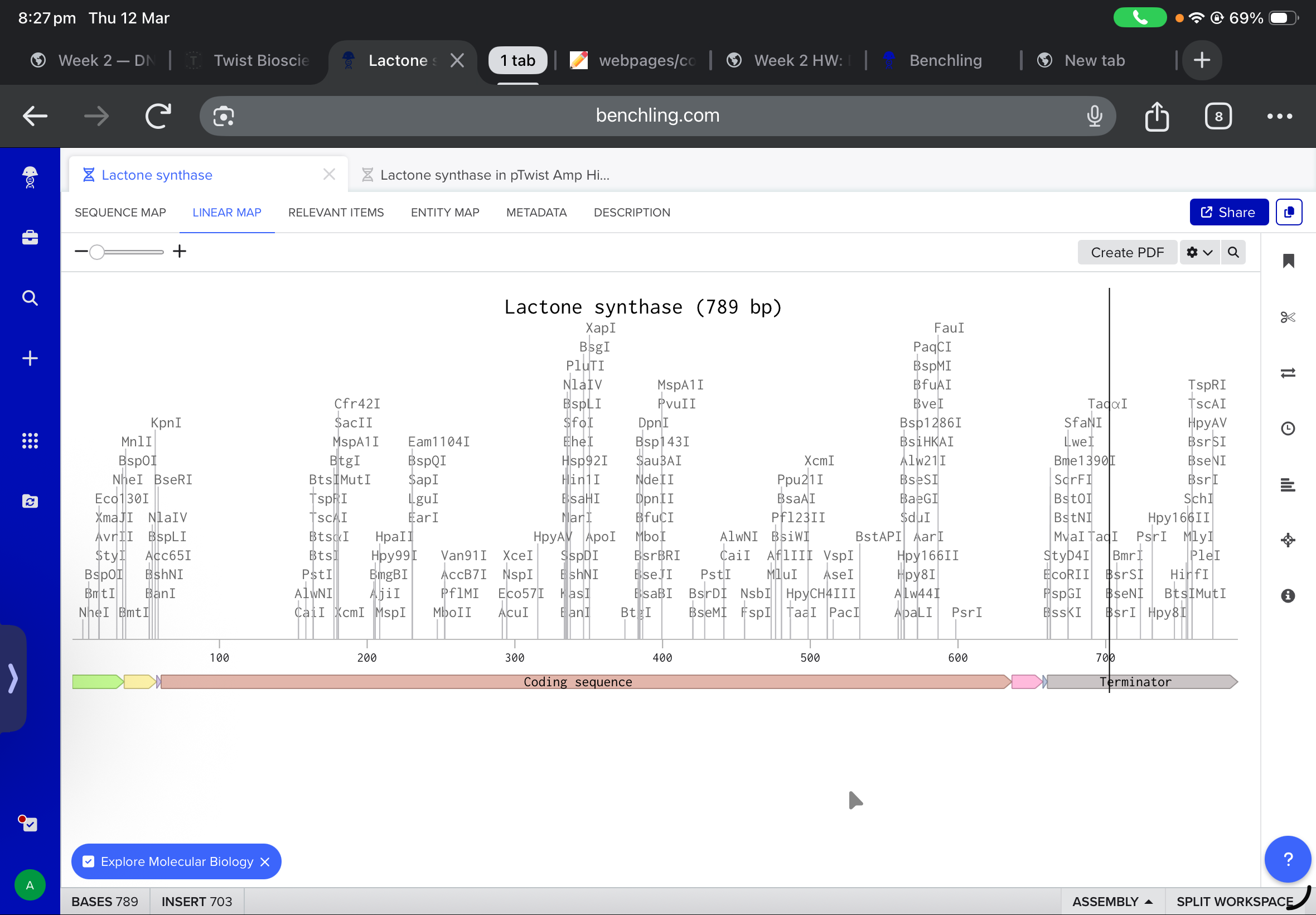

and after working with twist, and downlowing the genbank file to benchling, the final insertion plasmid is also shown below:
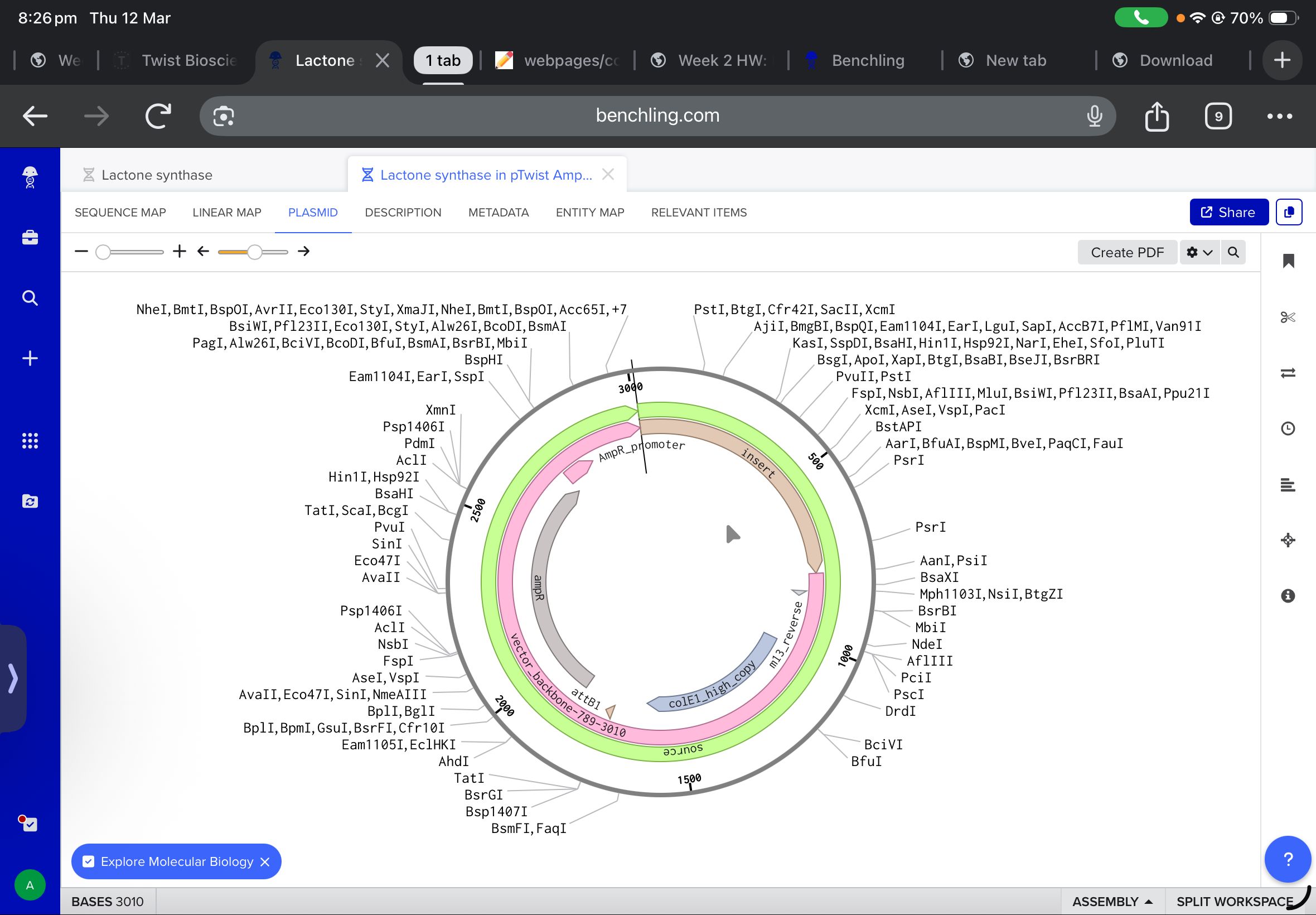

5.1 DNA READ
The DNA I would sequence is the gene responsible for telomerase activity, specifically the TERT gene. This gene encodes the catalytic subunit of telomerase, an enzyme that maintains telomere length at the ends of chromosomes. Telomeres protect chromosomes from degradation during cell division.
The sequencing method used would be Sanger sequencing, which is a first-generation sequencing technology. It determines DNA sequences using chain-terminating dideoxynucleotides during DNA synthesis.
The input for this method is genomic DNA containing the TERT gene extracted from human cells. The main preparation steps include:
- DNA extraction from the biological sample (e.g., blood, saliva, or hair follicle cells).
- PCR amplification of the TERT gene using specific primers to generate many copies of the target DNA fragment.
- PCR product purification to remove excess primers, nucleotides, and enzymes.
- Sequencing reaction preparation, where the purified DNA is mixed with DNA polymerase, normal nucleotides (dNTPs), fluorescently labeled chain-terminating nucleotides (ddNTPs), and sequencing primers.
- During sequencing, DNA polymerase extends the new DNA strand by adding nucleotides. Occasionally, a fluorescently labeled ddNTP is incorporated, which terminates the DNA strand because it lacks the 3′-OH group required for further extension. This produces DNA fragments of different lengths ending with labeled bases. The fragments are then separated using capillary electrophoresis, and a laser detects the fluorescent signals corresponding to each nucleotide.
The final output is a chromatogram (electropherogram) showing colored peaks for each base (A, T, G, and C), which is converted by the sequencing software into the digital DNA sequence of the analyzed TERT gene fragment.
5.2 DNA Write
- I would synthesize a DNA construct that blocks the androgen receptor (AR) when its levels get too high. It would include an AR-responsive promoter, a coding sequence for an AR inhibitor (like a peptide or siRNA), and a terminator. This could help prevent hair loss caused by high AR activity.
2.Synthesis technology
I would use Phosphoramidite DNA Synthesis (used by Twist Bioscience). It’s accurate, fast for short sequences, and allows assembly of full genes or small circuits.
- Essential steps
- Build DNA on a solid support.
- Add nucleotides in order (coupling).
- Cap incomplete strands.
- Oxidize and remove protecting groups.
- Assemble fragments into the full construct.
- Limitations
- Works best for short sequences; longer genes need assembly.
- Can be slow or expensive for large constructs.
- Small chance of errors, so verification by sequencing is needed.
5.3 DNA Edit
- I would use CRISPR‑Cas9 to edit DNA because it can target specific genes precisely. Cas9 is guided by a gRNA to a DNA sequence, then makes a cut. The cell repairs it either by:
NHEJ → introduces small mutations, or
HDR → uses a DNA template for exact changes.
- Preparation & input: design the gRNA, prepare Cas9 protein or plasmid, optionally include a donor DNA template, and deliver them into cells.
- Limitations: may cause off-target edits, not all cells are edited, and precise HDR edits are less efficient.
Week 3 HW: LAB AUTOMATION
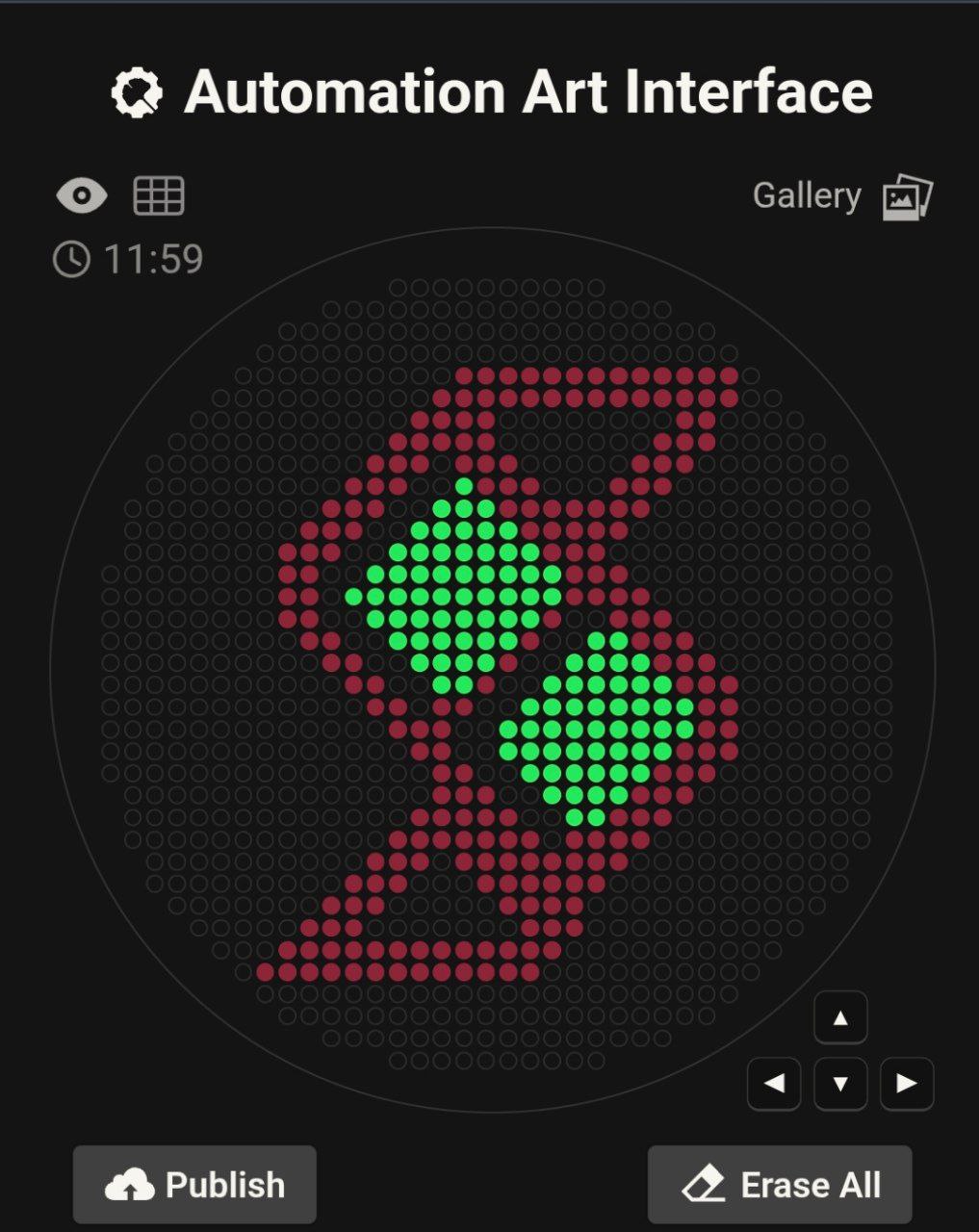

How cool it is to desgin the logo of SAB at AUIB?
the python scripit is uplouded
for the questions?
Q1: a paper that utilizes Openrons for biological purposes:?
I’ve came across this paper Titled: Automation of protein crystallization scale-up via Opentrons-2 (2025)
In Summary: it Uses an Opentrons OT‑2 robot to automate protein crystallization experiments, which are normally manual and laborious. The robot handles precise reagent dispensing, gradient formulation, and 96-well plate setup, increasing throughput and reproducibility. This is rare because low-cost robots are rarely used in structural biology workflows.
Q2:.
With my final project including Adding plasmid to E. coli and culturing them.
An automation Plan can be the following:
Use Opentrons OT‑2 to pipette prepared plasmids into E. coli wells.
Mix gently and adding growth medium.
Before finally Incubating and grow colonies.
The Python Example:
pipette.distribute(5, plasmid, ecoli_plate.rows()[0])
What else would be needed?
Extras: 3D-printed tube holders, use Nebula for protocol tracking.
Week 4 HW: Protien Design Part I

HW Questions
Assuming that the meat is ~20% protein, thus for 500 g meat ≈ 100 g protein. Average amino acid ≈ 100 Da = 100 g/mol, we can find the Moles of amino acids by ≈ 100 g (protein) ÷ 100 g/mol = 1 mol. According to Avogadro’s Number of molecules ≈ 6.02 × 10²³ amino acids in 1 mol.
Why eating beef doesn’t turn me into a cow? Proteins are digested into amino acids in the stomach and intestine and broken down before absorption, then transferred in the blood and reassembled in each cell based on our DNA and mRNA commands into different proteins. food’s structure is always broken down, that’s why insulin can’t be taken orally.
Why only 20 natural amino acids exist naturally? Many reasons, including Evolution induced minimal but sufficient set that already cover diverse properties: polar, nonpolar, charged, aromatic. Also, Genetic code constraints (triplet codons) stabilized this set and prevented increasing error rates without major benefit.
Non-natural amino acids? Yes, we can design them. For example: Bulky hydrophobic amino acid: extended aromatic rings with a side chain of 3 phenol substitutions. Or Metal-binding amino acid: includes extra histidine-like rings that bind to metals and increase stability or provide a catalytic function.
Origin of amino acids before life
• Formed via prebiotic chemistry (e.g., Miller–Urey-type reactions)
• Sources: lightning, volcanic gases, meteorites
• Found in space (e.g., meteorites like Murchison meteorite)
• Accumulated before enzymes existed
Why most helices are right-handed
Because proteins use L-amino acids, whose stereochemistry favors right-handed packing with low steric effects .
Why β-sheets aggregate
• Backbone forms strong hydrogen bonds
• Flat structure allows stacking
Why amyloid diseases come form β-sheets?
Misfolded proteins reorganize into stable β-sheet-rich fibrils
Examples include Alzheimer’s disease
Can they be used as materials?
Yes, Extremely strong (like nanofibers) or Used in biomaterials, nanotechnology, and tissue scaffolds
Designing a β-sheet motif
Key principles: we Alternate hydrophobic (H) and polar (P) residues, Example pattern:
H–P–H–P–H–P–H–P. The machanisims were a Hydrophobic side chains align on one face that drives stacking, Adding turns can happen with a Gly–Pro to stabilize sheet folding . The Result: ordered, self-assembling β-sheet structure.
PART A
I choose lactone synthase, an enzyme responsible for the synthesis of compounds containing lactone which are bio adhesive, protective and good smelling.
2. Its 230 aa, the most frequent one is Leucine. like shown below
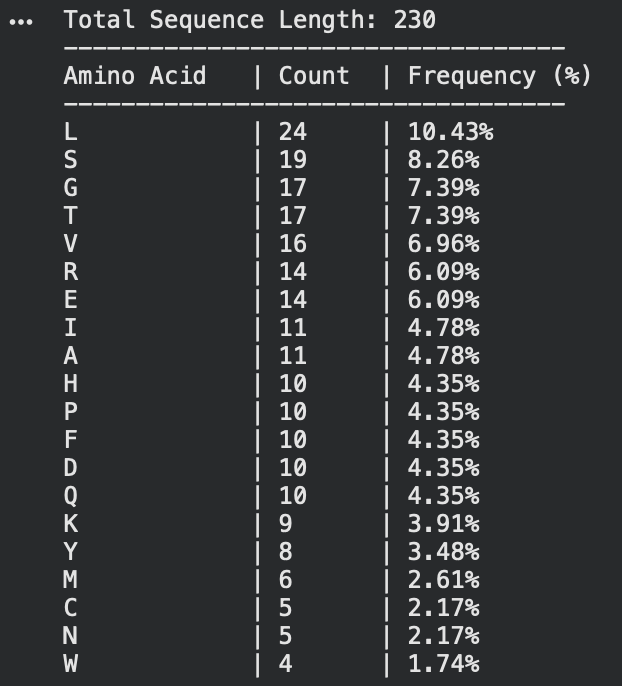
 3. there was around 250 homologue sequences there for it.
4. Transferase family of finctionality.
3. there was around 250 homologue sequences there for it.
4. Transferase family of finctionality.
Identifying the structure page of the protein in RCSB!
well, There is no other molecules in the protein.
and It was solved in 2002, with a good resolution of 1.8Å
It is an autoinducer synthase family.
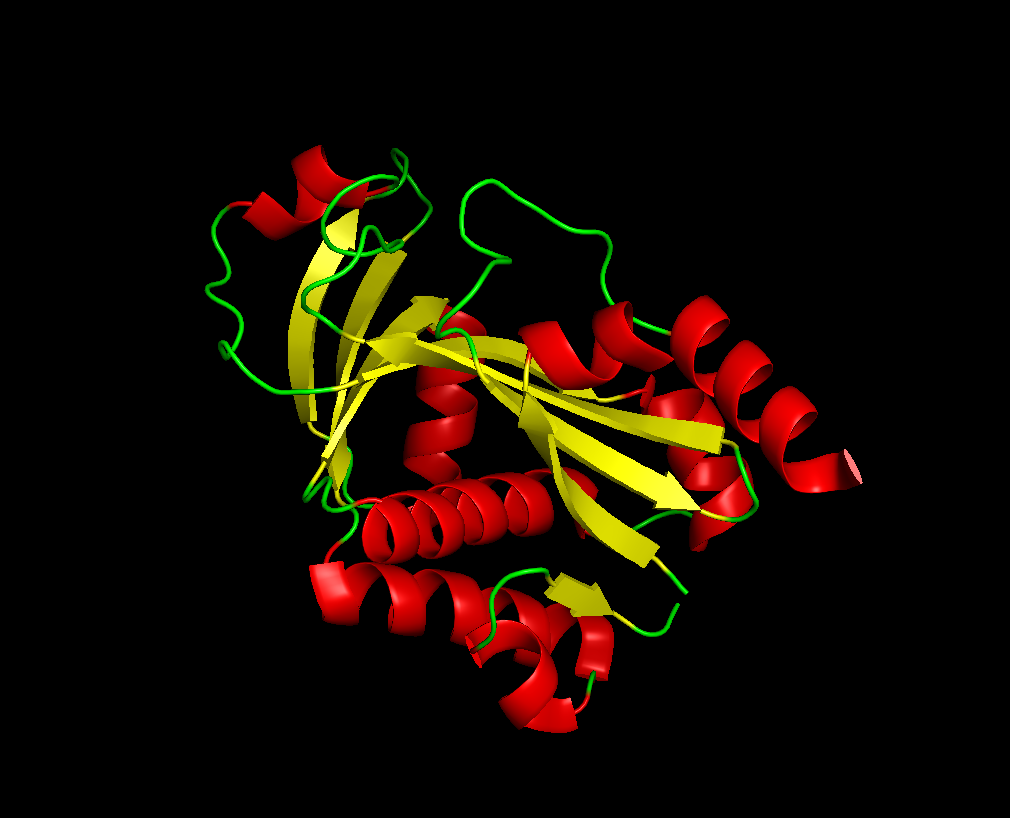
for the 3D part:
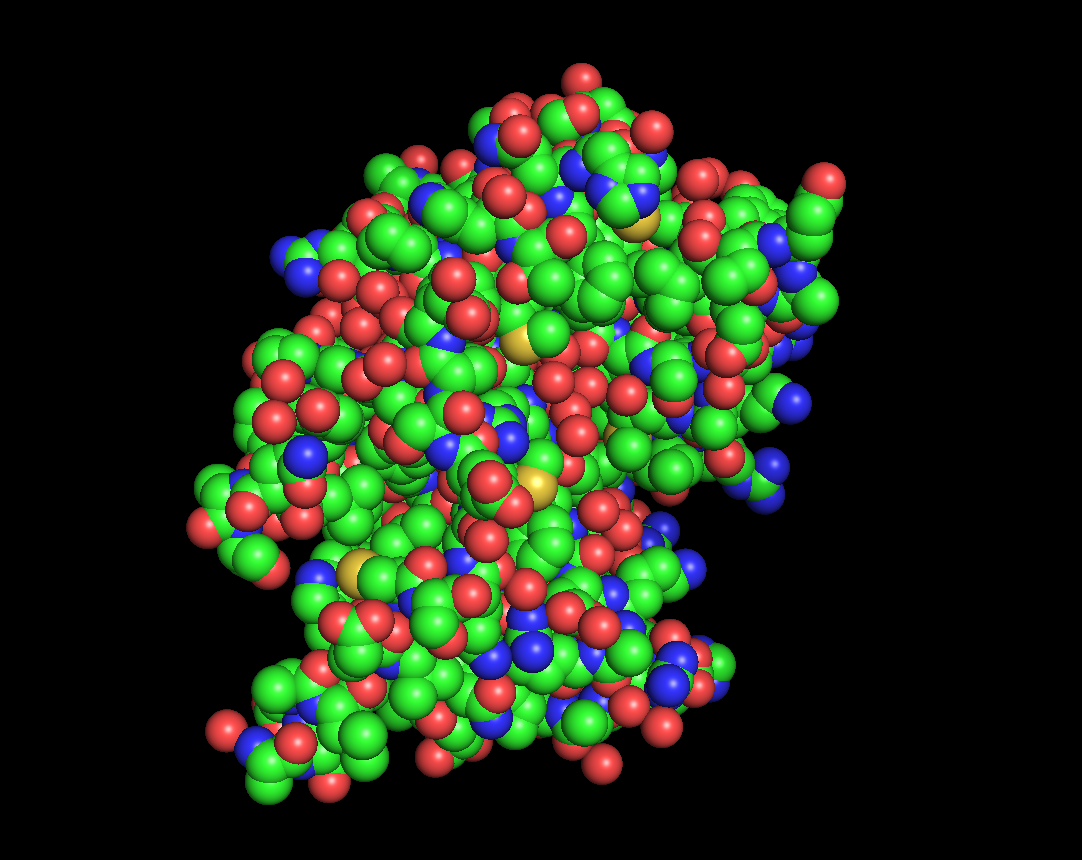


after the varoius types of visualization of my protien, I acquired the following informations:
Total main 5 alpha helix and 3 mini, while the beta sheet were 7 large and one mini. So the name of helix and sheets, with sheets larger.
The protein has many hydrophobic and hydrophilic properties, the hydrophilic amino acids are more oriented towards the outer layers of the protein.
There was around 4 cavities that might work as binding pockets












