Week 2 HW: DNA Read Write and Edit
Question 3
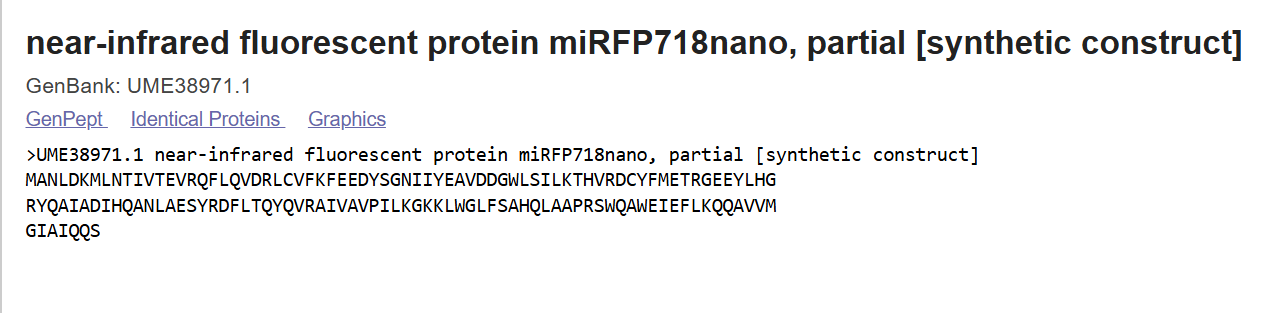

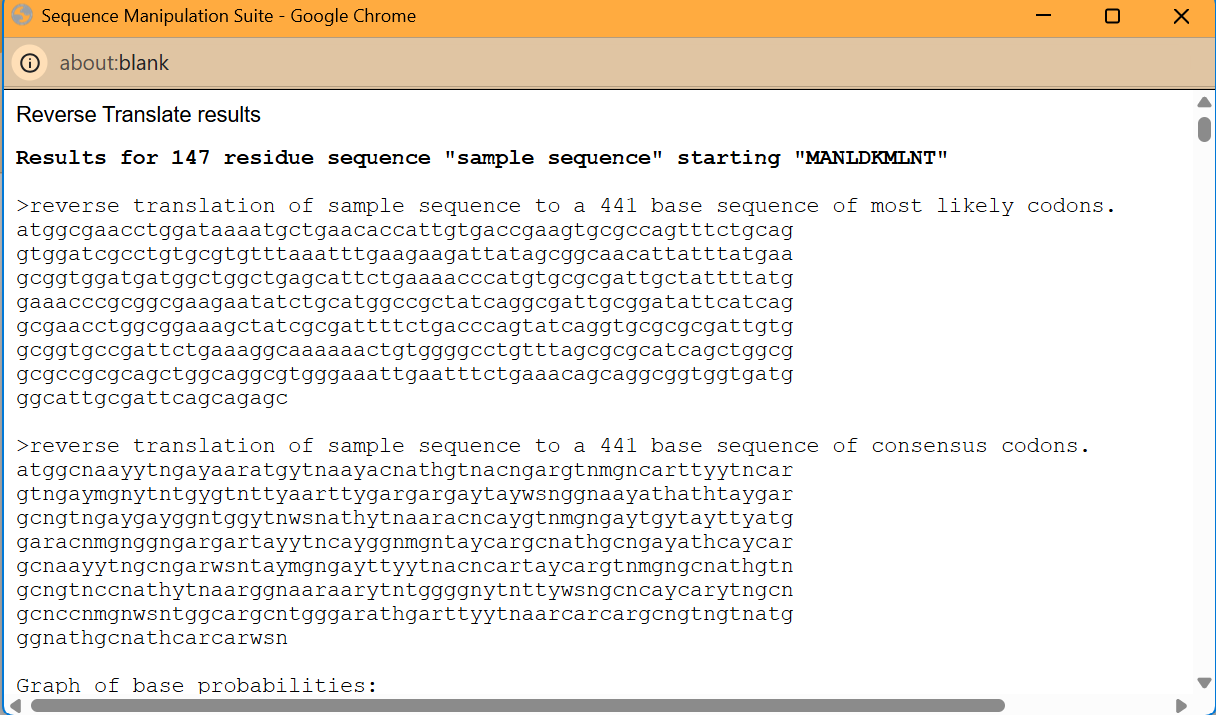

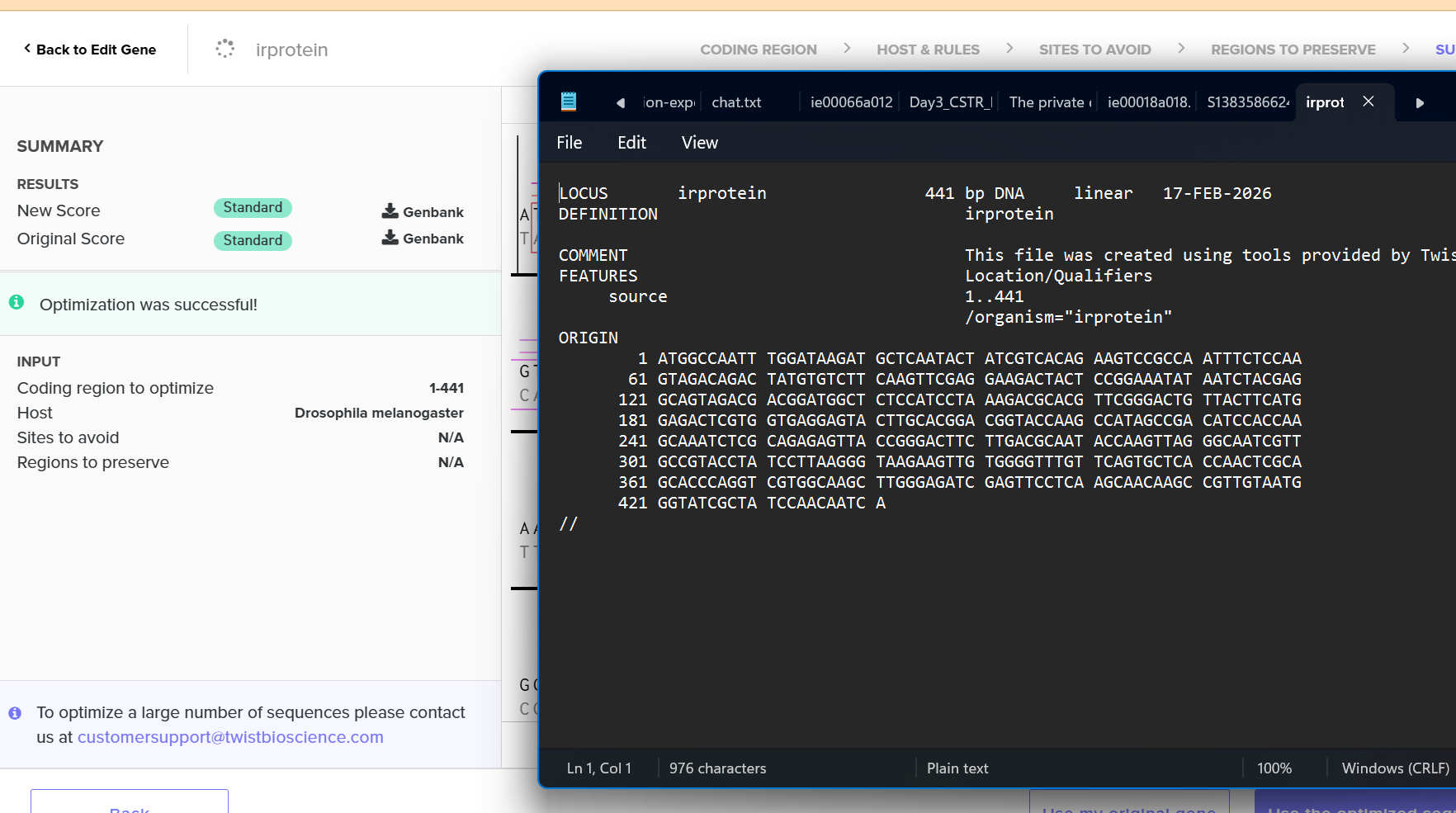
 I chose fruit fly because of their ommitidia.
3.4)
I would first amplify with PCR, and then purify with gel extraction. Make sure I have a sizeable concentration of DNA, and then I would try to use a cell free system to produce my protein. I would need to choose the right buffer that has the RNAases and ribosomes/tRNAs I need for transcription and translation, respectively.
I chose fruit fly because of their ommitidia.
3.4)
I would first amplify with PCR, and then purify with gel extraction. Make sure I have a sizeable concentration of DNA, and then I would try to use a cell free system to produce my protein. I would need to choose the right buffer that has the RNAases and ribosomes/tRNAs I need for transcription and translation, respectively.
Question 5 Read I) I want to read the gene related to oxidation reactions in human cells so that I can build effective antioxidant therapies. This could be applied from preserving the taste in foods to fighting ROS deregulation. Ii) 1. My method is third generation, as I would likely need to sequence many long mammalian DNA in parallel. 2. I would obtain mammalian cells, extract DNA, cut around the supposed regions of interest that control oxidation reactions in cells with restriction enzymes. Order primers and amplify these regions of interest with PCR. And then ligated to a plasmid so that it can be transformed with competent cells and transfected into bacterial vectors. 3. The sequencing technology works by anchoring the DNA fragments down, and then fluorescenting matching the bases while the camera captures the light that emitted. Piecing together the fragments with overlaps allows decoding of the entire sequences of interest. 4. The output will be bacteria that are able to reduce the amount of oxidative species in its environment.
Write I) I would like to write a novel antibody sequence that is able to detect receptors that show up on cells that become cancerous and circumvent the normal cell cycle. Ii) 1. I would need to write my sequence of interest using existing antibody research and my best hypothesized sequences that might work, and then order the sequences through twist, and integrate them into cancerous mammalian cells to see if the cells lose their cancerous character. 2. There is no known cure to all cancers at the moment, the ability to come up with accurate sequences is a major rate limiting step. Amplifying and transecting into mammalian cells is likely a major time and scalability limiting step.
Edit I) I would like to edit genes that allow some organisms to see infrared light to see if other wavelengths of light can be sensed by those organisms. Ii) 1. I would use CRISPR for the specificity and accuracy and ease of editing. I would need the respective guide RNAs for the genes, and the cas9-CRISPR system to edit the DNA. 2. I would need to prepare the CRISPR system to work in vivo with the organism of interest, I would deliver the editing system with virial vectors, with the respective guides. 3. Major limitations would be the delivery and measuring the output, and I would combat this by optimizing the cycle through trial and error, and have adaptive behavior testing based on stimulus from the wavelength of light of interest. I could also use this test to try to evolve their DNA as needed.