Week 4 HW: Protein Design
Part A
How many molecules of amino acids do you take with a piece of 500 grams of meat? (on average an amino acid is ~100 Daltons) Meat is 20-30% protein, so 0.25*500 g = 125 g of protein = 125 g of amino acids. 100 Daltons = 100 g/mol, so there is 125 g/100 g/mol = 1.25 moles of amino acids in the meat which is 7.53e+23 molecules of amino acids in the meat.
Why do humans eat beef but do not become a cow, eat fish but do not become fish? Because our digestive systems break down the complex molecules into the substituents that our body can use to reassemble into the molecules that we need to function. We don’t absorb the whole cow cell into our system, but we would break down the cell, grab the essential lipids, carbs, proteins, and nucleic acids from the cell, and then reassemble them into something we can use to make us function.
Why are there only 20 natural amino acids? Evolution likely didn’t need to go beyond the complexity of 20 amino acids. It was able to use the 20 amino acids to build proteins that carried a variety of tasks that allowed many different species to evolve in many different environments. Beyond this, it was likely unnecessary and a waste of resources that would deem that organism less fit if it had more than the 20 natural amino acids.
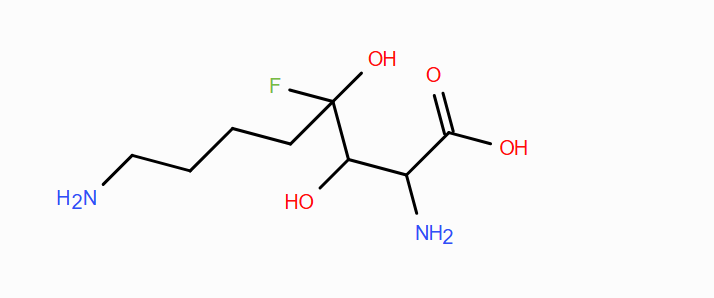
Can you make other non-natural amino acids? Design some new amino acids.
Yes, you can adjust the side chains with new structures to your liking with the right chemistry and reagents. For instance, you can design one that is able to hydrogen bond, but also vary the polar the character of the amino acid by changing the length of the carbon chain that has the hydrogen-bonding carriers on them

Where did amino acids come from before enzymes that make them, and before life started? Before life, carbon rich carriers from space and natural resources on Earth may have provided the basis for chemical reactions with carbon and other life-essentials atoms to react together with the energy provided by the sun, specifically UV light. From here, a base structure for amino acids may have been made excessively and, then this base would react with other elements to make the variation we see with amino acid side chains.
If you make an α-helix using D-amino acids, what handedness (right or left) would you expect? I would expect a left handedness helices because the chirality would switch from the typical L-amino acid to the D-amino acid, and this would then favor a left handed helice
Can you discover additional helices in proteins? Yes, if you can design new amino acids, then nature can too. With new side chains, new interactions that we haven’t documented yet may arise, and that is how we may discover additional helices in proteins.
Why are most molecular helices right-handed? The right handedness comes from the prevalent L - amino acids, because helices are stabilized by the intermolecular forces provided when they twist with right handedness.
Why do β-sheets tend to aggregate? They aggregate because the sheets have hydrogen bond carriers and acceptors that get paired, and the layers start stacking together as a result.
Part B
My protein is Ole E 1, its a major allergen (pollen) and I selected it because I want to look into the structural components of this protiens and what the key sites could be that allow us to denature the protein to reduce the amount of pollen allergen in the air.
SEQUENCE 184 Amino acids; 19969 Molecular Weight; MASTGAATNL LLLAMVVVVA TADYYAQPQP YVPKPTTTYT SPVKTPYLPK SNPDIAIEGF ILCKSGYKTY PIQGGKVKVV CPVVDSYGKL VAKVTISSYP TDLKGYFYFI TYGLSHKVNN ISSCKVKLES SPVFTCKTPT NVNKGVTGAP LSPDNSKFLS HDNLTLYTLE PFYFSSPVAP KPVY
The most common amino acid is V, appearing 20 times.
Blast was able to find 250 homologs to this protein sequence.
Yes, it belongs to Pollen Ole e 1 allergen and extensin family protein.
I couldn’t find this protein on the RCSB database, so I searched for hemoglobin subunit alpha with MVLSPADKTNVKAAWGKVGAHAGEYGAEALERMFLSFPTTKTYFPHFDLSHGSAQVKGHG KKVADALTNAVAHVDDMPNALSALSDLHAHKLRVDPVNFKLLSHCLLVTLAAHLPAEFTP AVHASLDKFLASVSTVLTSKYR
and it was solved in 1998-11-11 with good resolution (1.59 Å). There are other portions of the hemoglobin such as the Fe holding parts, and the multiple other subunits that make up the entire structure but the solved structure is only showing the amino acid sequence in the 3D structure. In general, it belongs to domain 2DN3 A:2-141 SCOP ID: 8027457 Hemoglobin subunit alpha Species: Homo sapiens
I am visualizing my protein on pymol:
ball and stick:
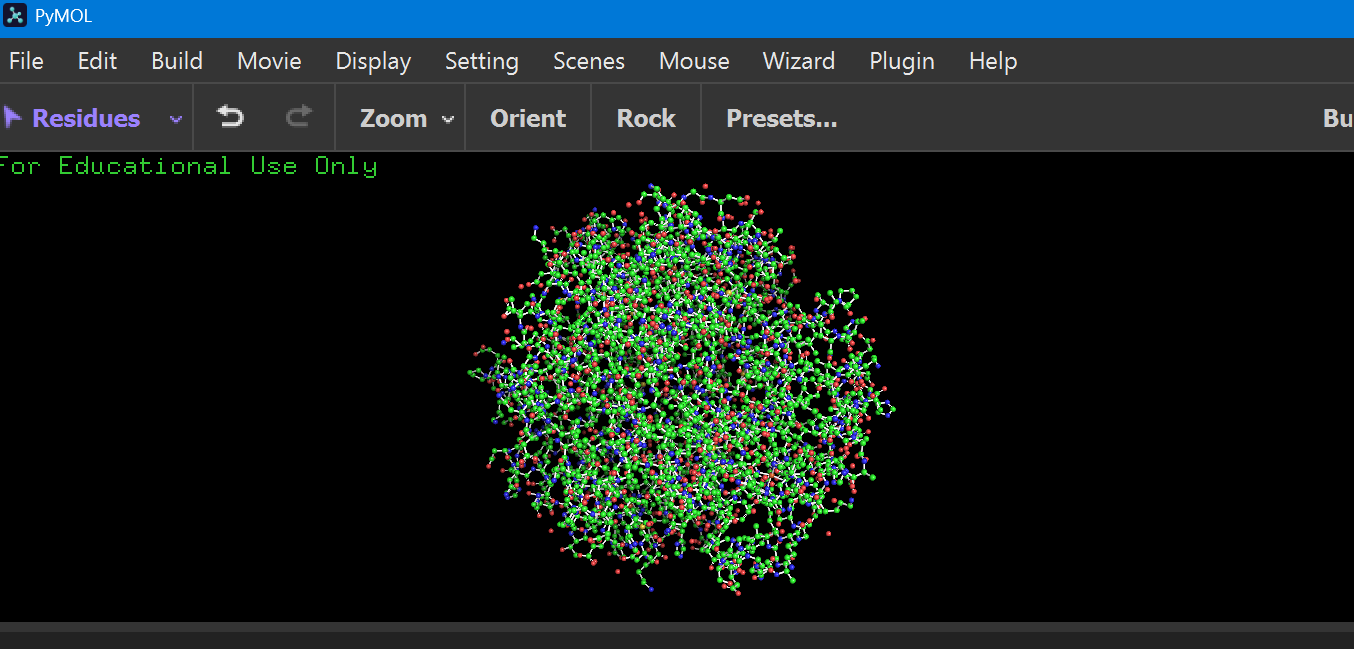
 cartoon:
cartoon:
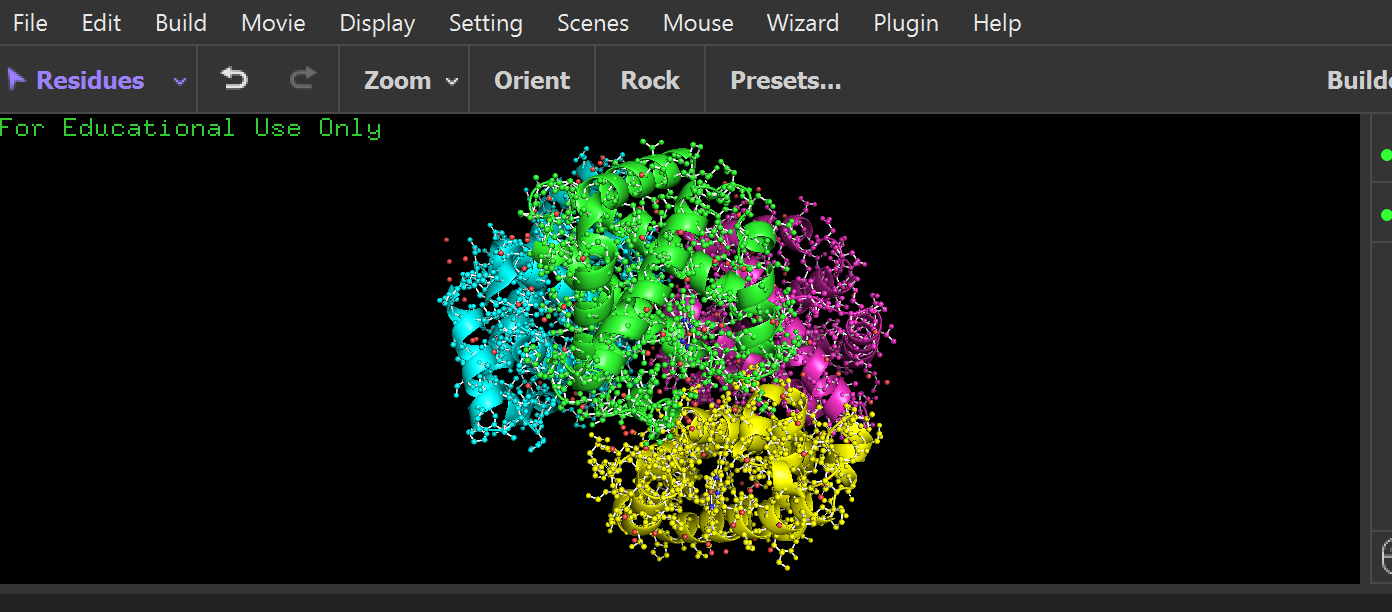
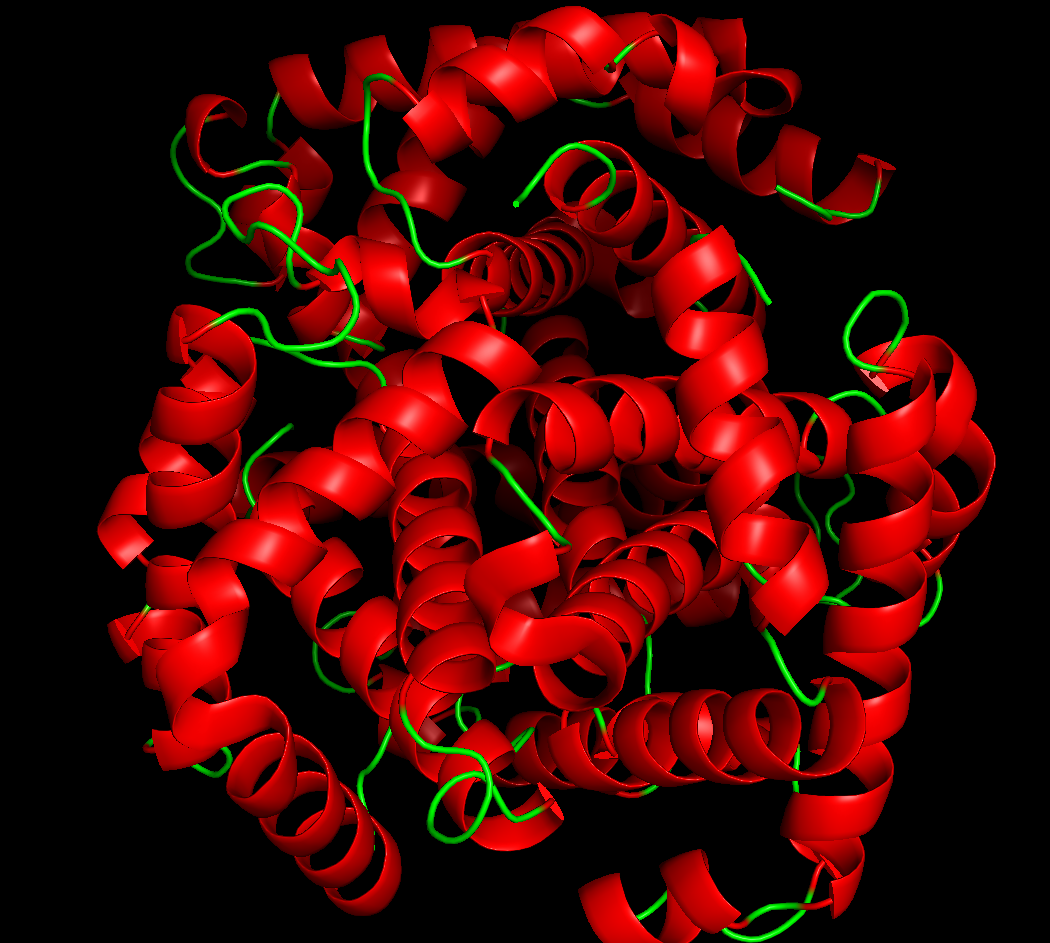
 ribbon:
ribbon:
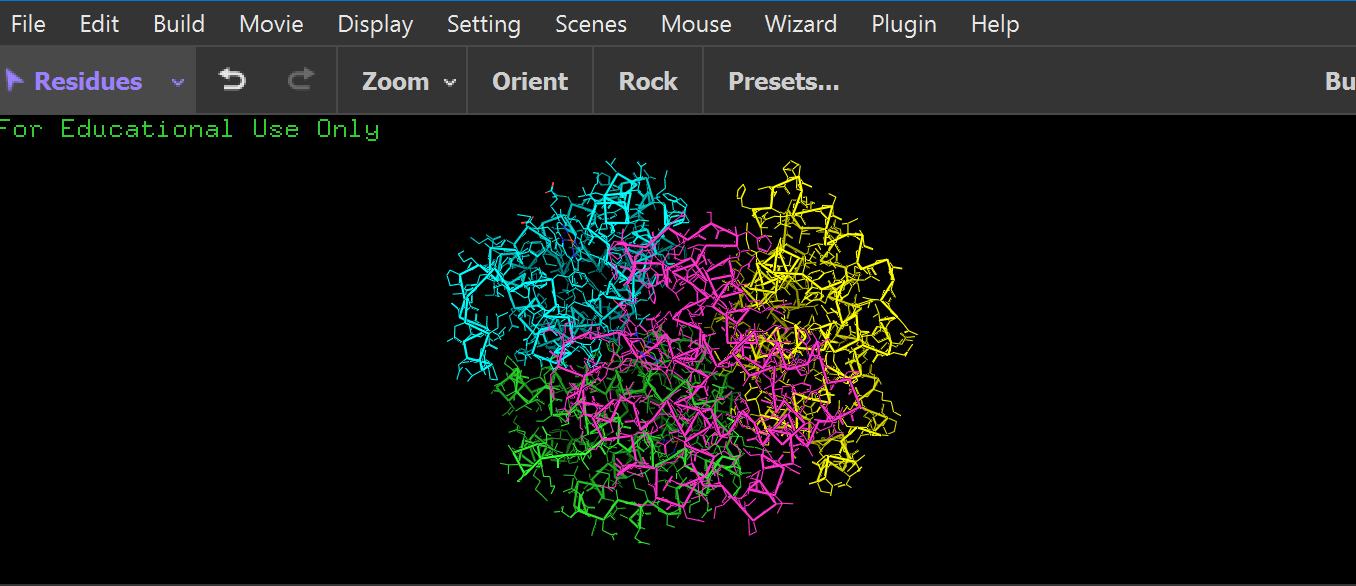
 it has more helices:
it has more helices:
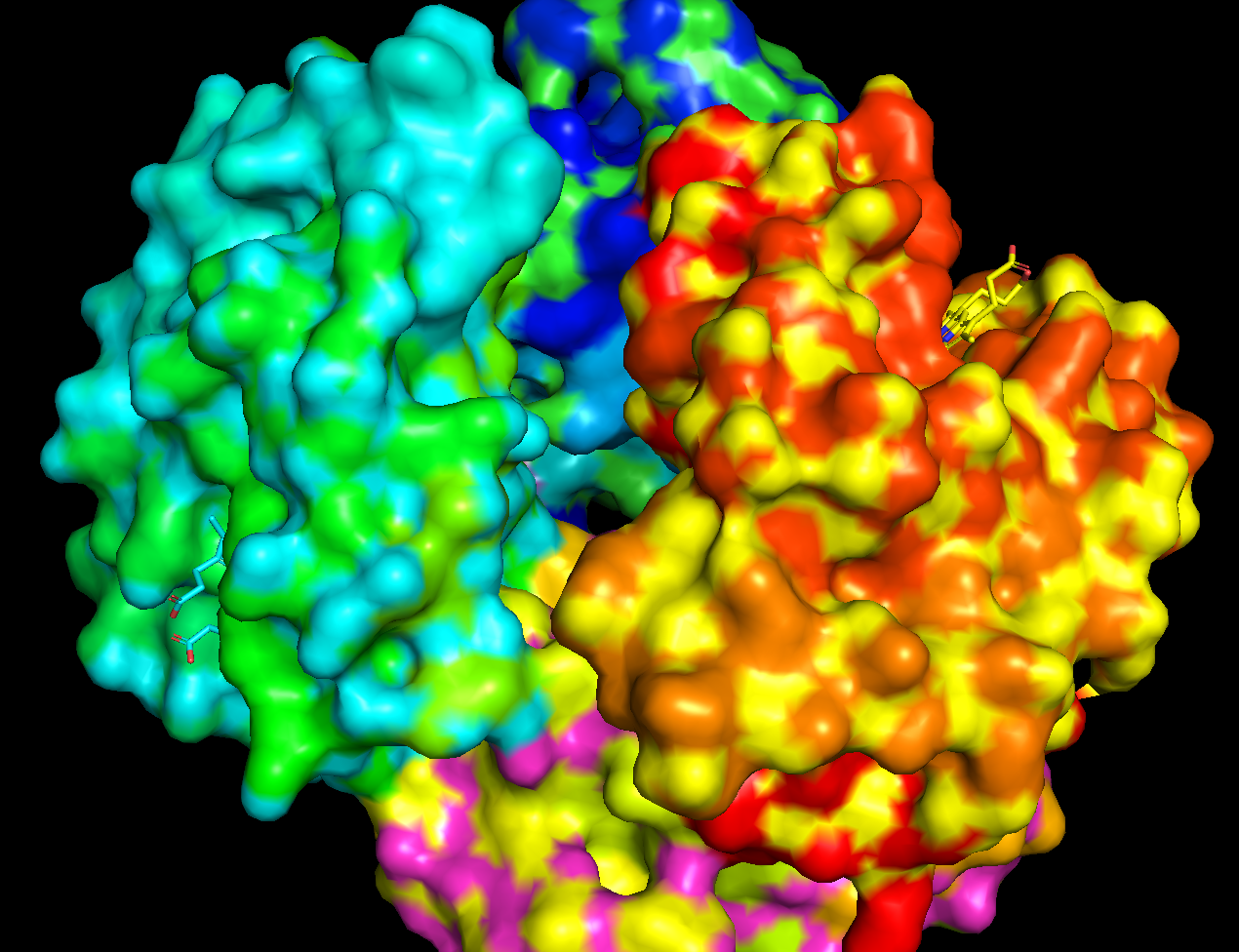
 Residue type, the hydrophyllic residues are gathered internal to the protein while the hydrophyllic sites (for Fe attachment I am guessing) are around surface typically
Residue type, the hydrophyllic residues are gathered internal to the protein while the hydrophyllic sites (for Fe attachment I am guessing) are around surface typically
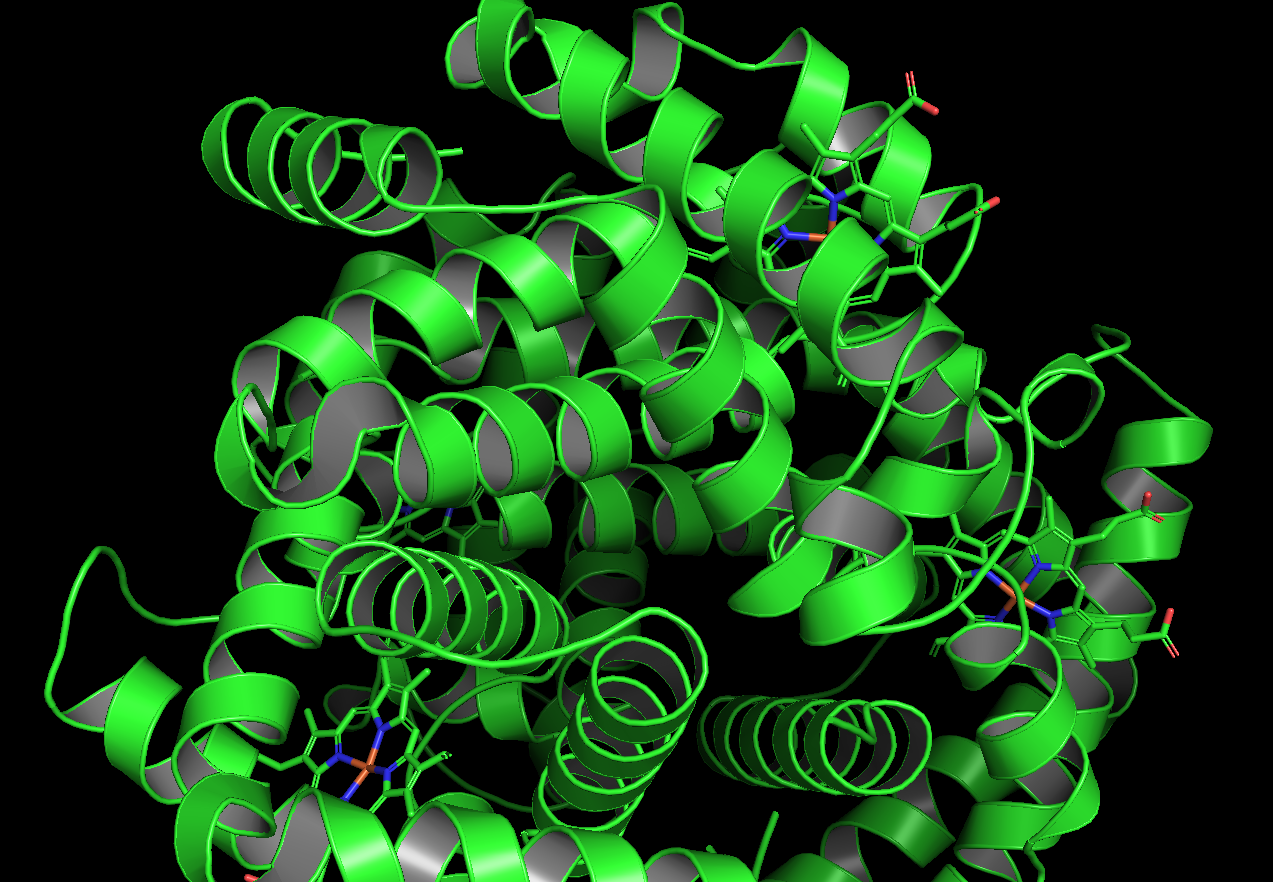

 It does have holes a few large holes on the surface
It does have holes a few large holes on the surface

Part D
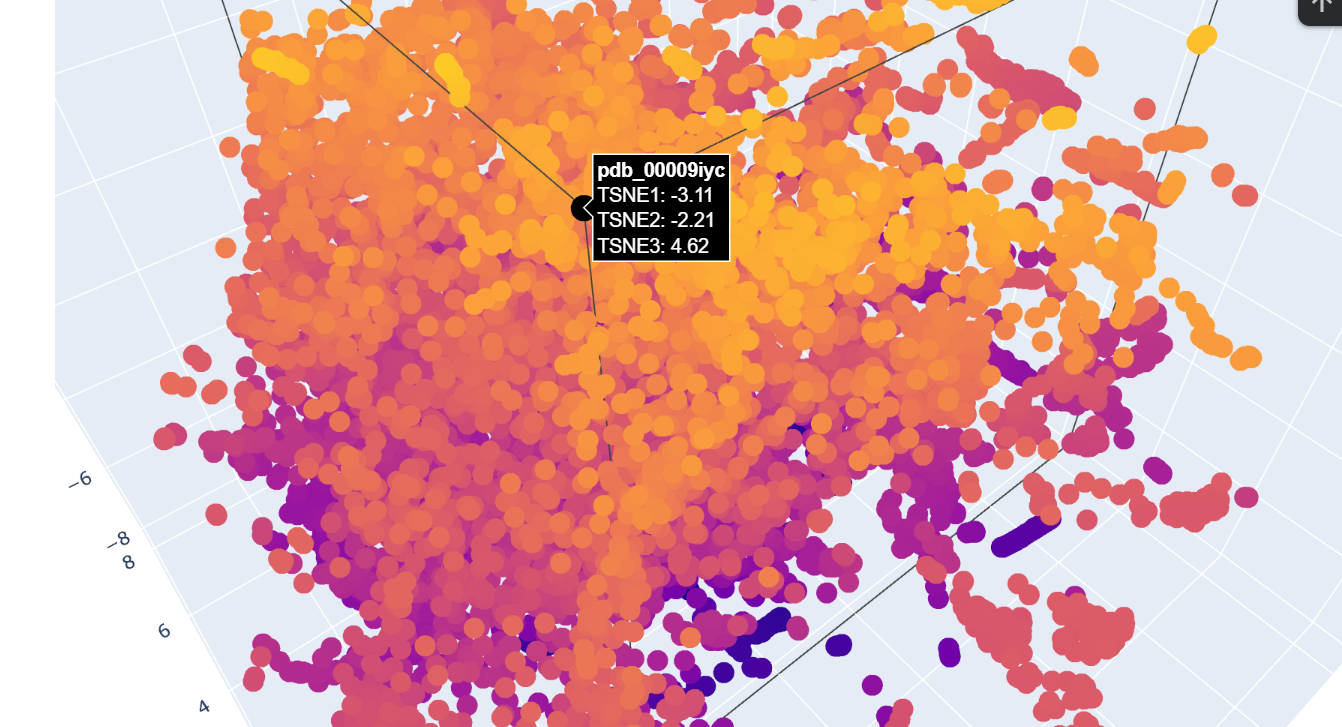
I chose this most recent uploaded protein on RCSB
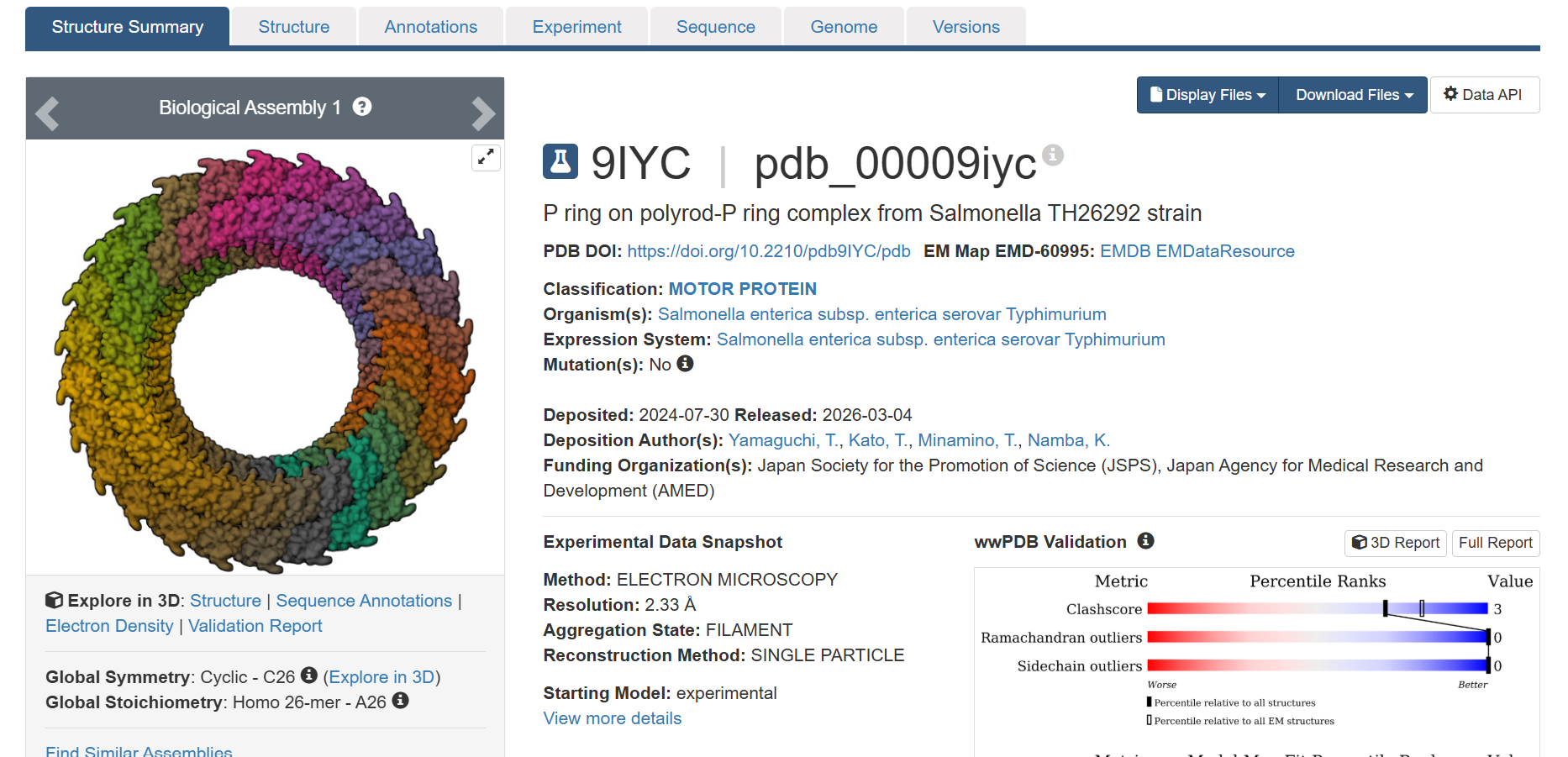

It gives the MSA. It looks like there are places in the sequence where the model scores high on a variety of mutations, such as around position 250. W and C mutations scored low across most of the positions in this protein sequence, meaning there is likely less di-sulfide bonds present in this protein
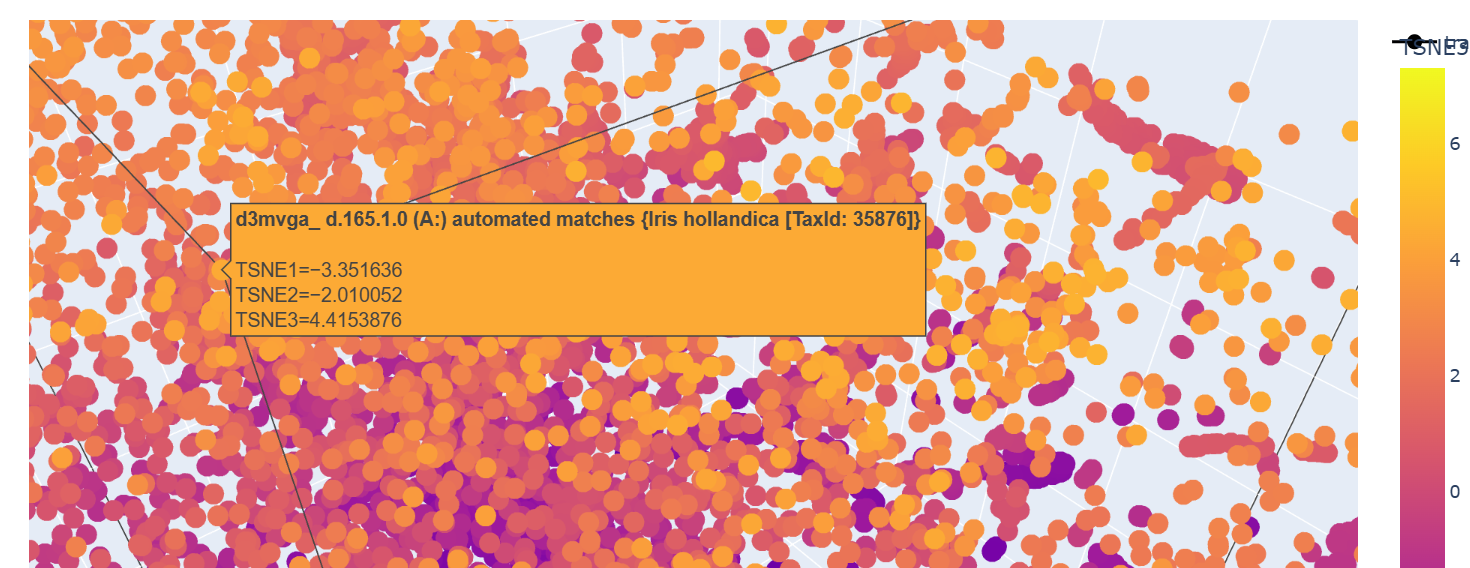
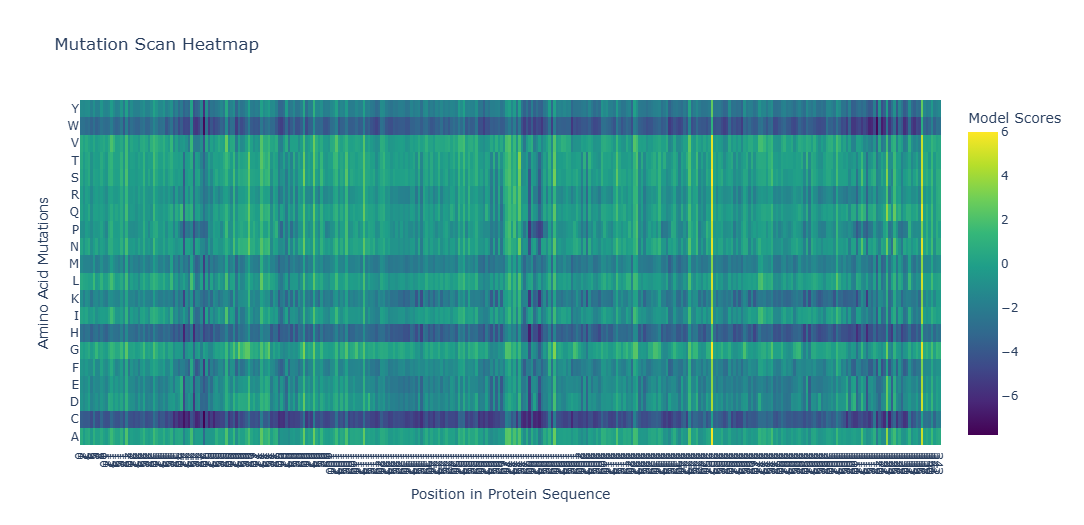

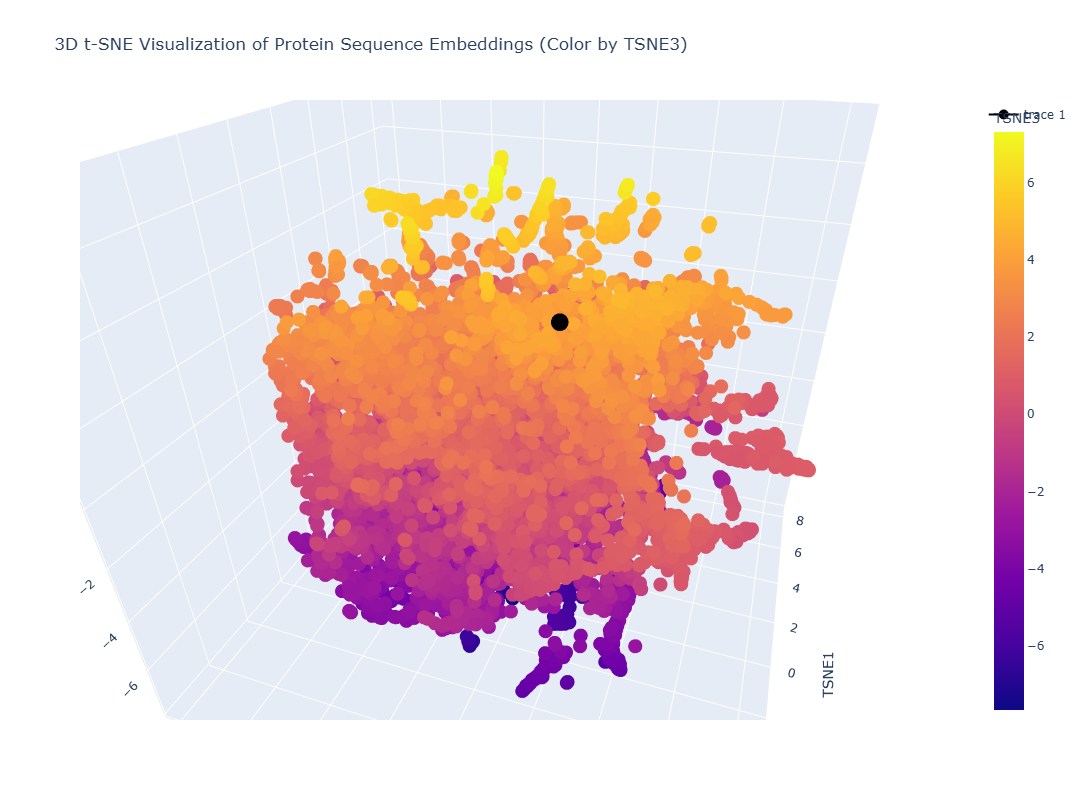

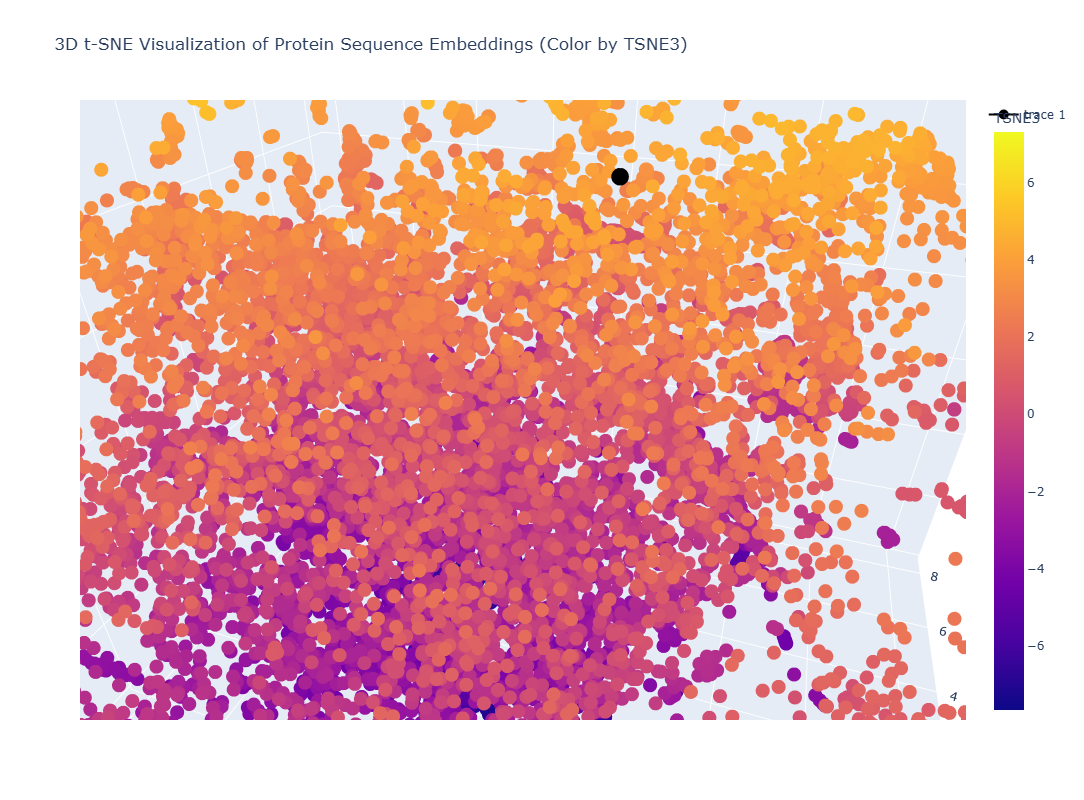



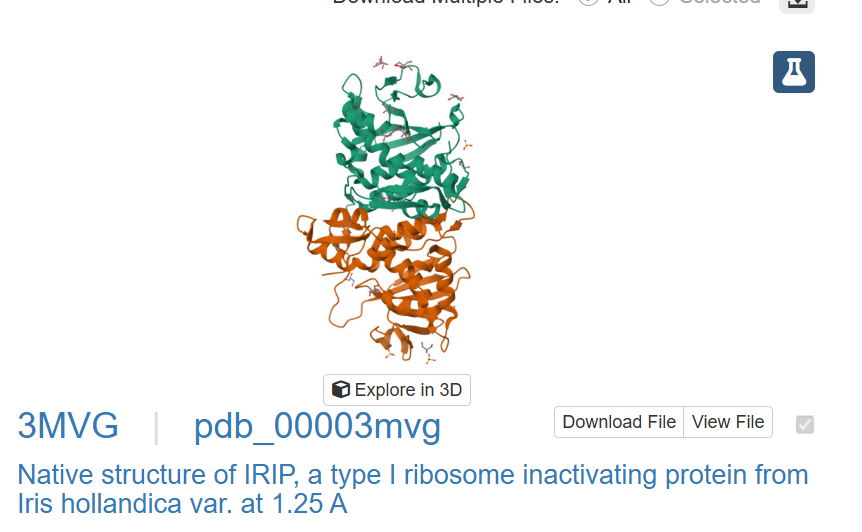
 This one could be similar for when two domains intersect and the portein I chose uses similar mechanism to attract the two domains together.
This one could be similar for when two domains intersect and the portein I chose uses similar mechanism to attract the two domains together.
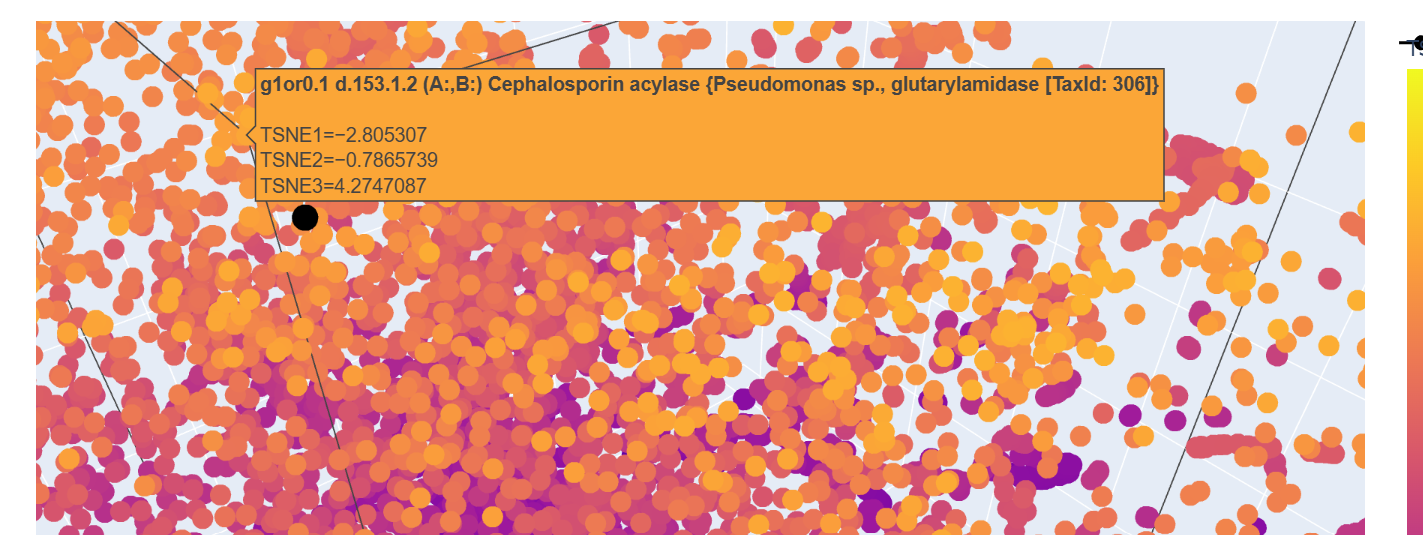


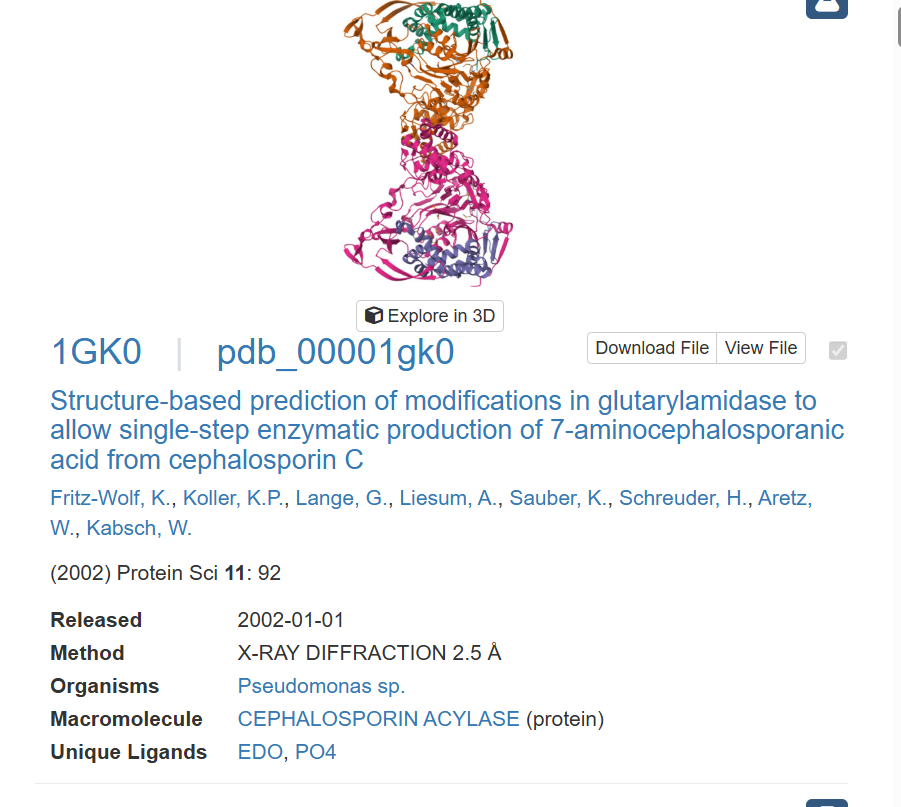
 This one looks like if you have multiple of this protein, you could rebuild the protein I chose, so this may provide the basis for which the protein I chose is built on.
This one looks like if you have multiple of this protein, you could rebuild the protein I chose, so this may provide the basis for which the protein I chose is built on.
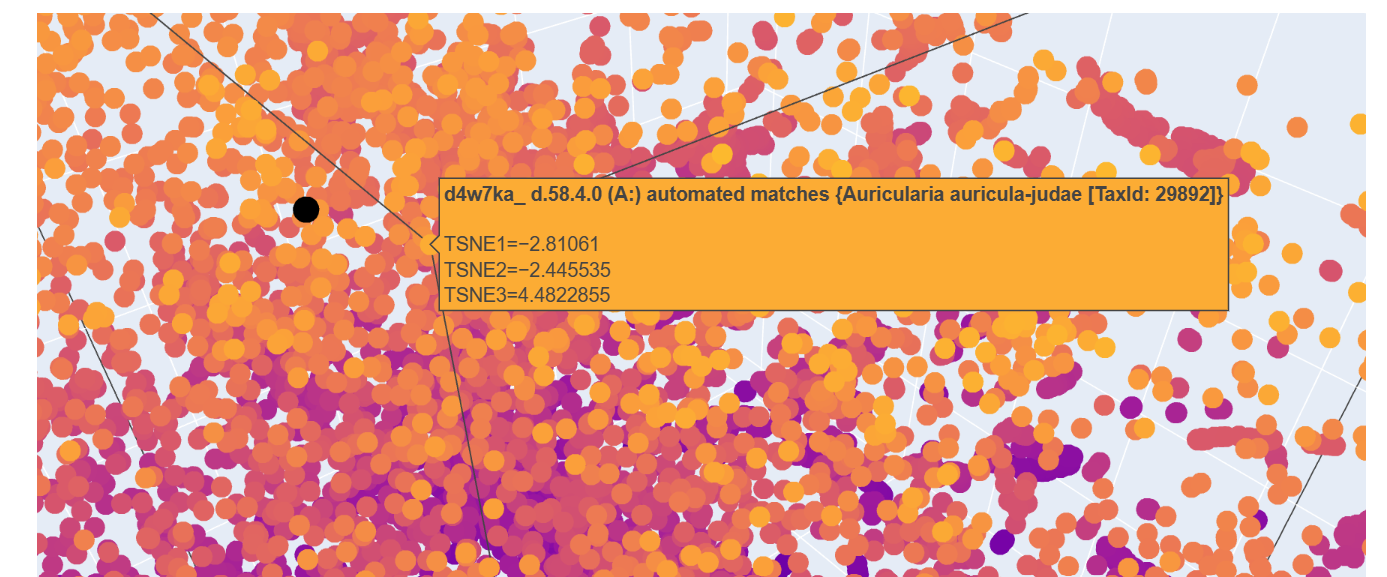



 This one seems like a bunch of regions intesecting at the middle whereas the other is a taurus shape. They might be really similar in that they have multiple regions interacting to make a specific shape but how they interact vary and that could give rise to the different shape.
This one seems like a bunch of regions intesecting at the middle whereas the other is a taurus shape. They might be really similar in that they have multiple regions interacting to make a specific shape but how they interact vary and that could give rise to the different shape.

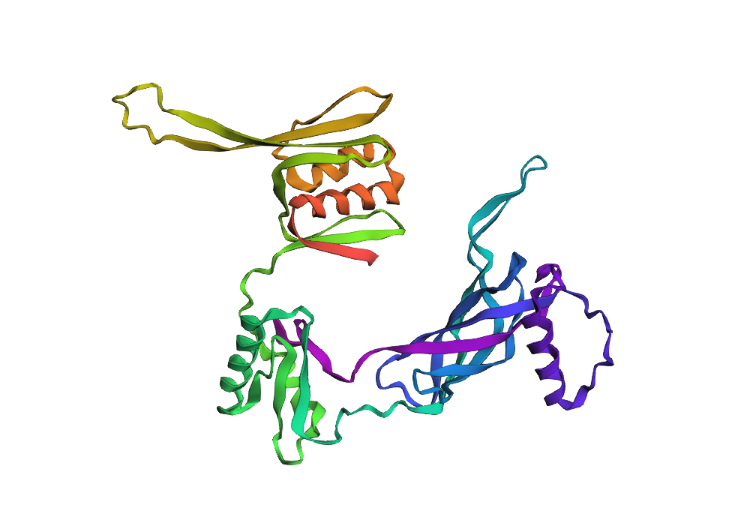
 While the model tried to fold the protien sequence I gave it, it was not too accurate to the actual structure. This protein was added to the database in March 2026 and has a relatively intricate/unique structure that the model likely wasn’t able to predict accurately nor precisely.
While the model tried to fold the protien sequence I gave it, it was not too accurate to the actual structure. This protein was added to the database in March 2026 and has a relatively intricate/unique structure that the model likely wasn’t able to predict accurately nor precisely.
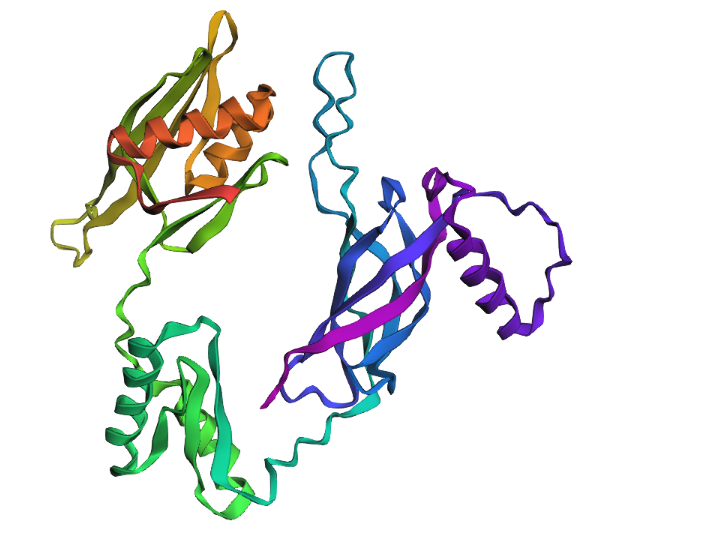
 The protein is relatively resistent to mutations, after one amino acid deletion, after many amino acids deletions, after many amino acids insertions, the structure was still able to form a relatively in tack loop.
The protein is relatively resistent to mutations, after one amino acid deletion, after many amino acids deletions, after many amino acids insertions, the structure was still able to form a relatively in tack loop.