Week 2 HW: DNA Read, Write, & Edit
Part 0: Basics of Gel Electrophoresis
Attend or watch all lecture and recitation videos. Optionally watch bootcamp.
Done :checkmark:
Part 1: Benchling & In-silico Gel Art
See the Gel Art: Restriction Digests and Gel Electrophoresis protocol for details. Overview:
Make a free account at benchling.com
Import the Lambda DNA.
Simulate Restriction Enzyme Digestion with the following Enzymes:
EcoRI
HindIII
BamHI
KpnI
EcoRV
SacI
SalI
Create a pattern/image in the style of Paul Vanouse’s Latent Figure Protocol artworks.
You might find Ronan’s website a helpful tool for quickly iterating on designs.
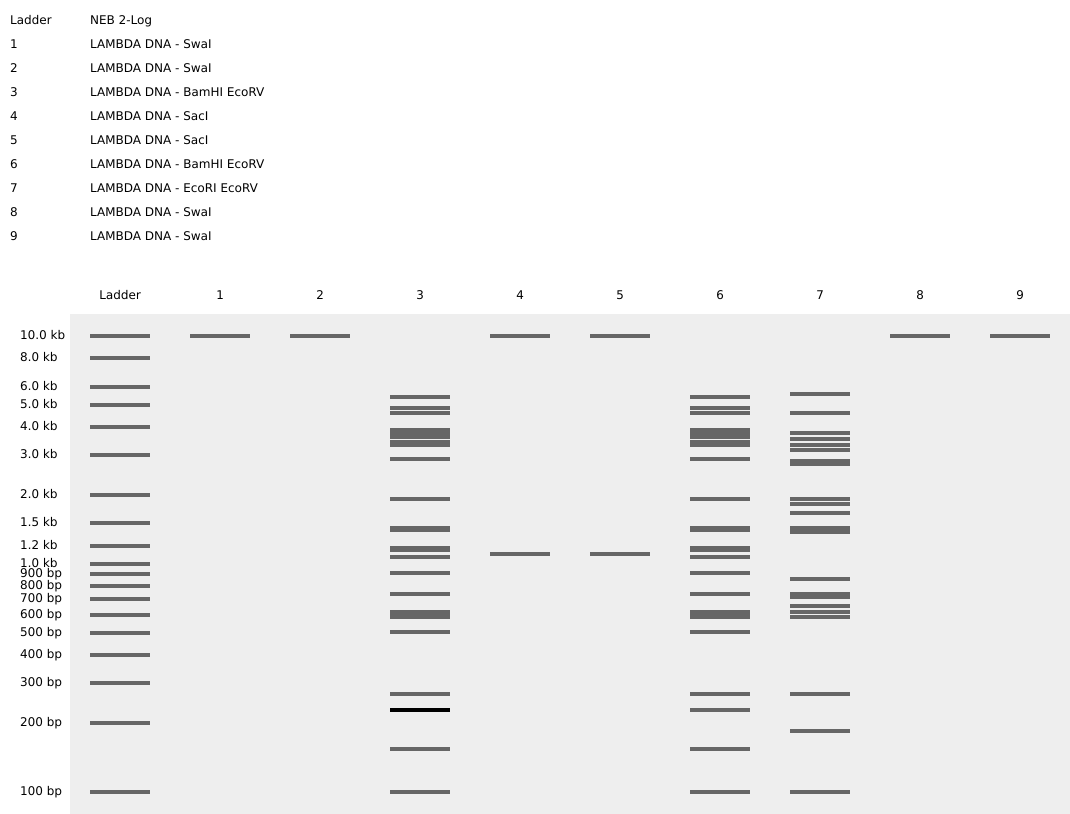

I title it “A greeting..”
Part 2: Benchling & In-silico Gel Art
Unable to do, lack the lab access.
Part 3: DNA Design Challenge
3.1. Choose your protein.
Which protein have you chosen and why?
I have chosen Lactose. I wanted to do Lactase instead, but it seemed to big.
sp|P00709|LALBA_HUMAN Alpha-lactalbumin OS=Homo sapiens OX=9606 GN=LALBA PE=1 SV=1 MRFFVPLFLVGILFPAILAKQFTKCELSQLLKDIDGYGGIALPELICTMFHTSGYDTQAI VENNESTEYGLFQISNKLWCKSSQVPQSRNICDISCDKFLDDDITDDIMCAKKILDIKGI DYWLAHKALCTEKLEQWLCEKL
3.2. Reverse Translate: Protein (amino acid) sequence to DNA (nucleotide) sequence.
Alpha-lactalbumin DNA sequence
atgcgcttttttgtgccgctgtttctggtgggcattctgtttccggcgattctggcgaaa cagtttaccaaatgcgaactgagccagctgctgaaagatattgatggctatggcggcatt gcgctgccggaactgatttgcaccatgtttcataccagcggctatgatacccaggcgatt gtggaaaacaacgaaagcaccgaatatggcctgtttcagattagcaacaaactgtggtgc aaaagcagccaggtgccgcagagccgcaacatttgcgatattagctgcgataaatttctg gatgatgatattaccgatgatattatgtgcgcgaaaaaaattctggatattaaaggcatt gattattggctggcgcataaagcgctgtgcaccgaaaaactggaacagtggctgtgcgaa aaactg
3.3. Codon optimization.
Alpha-lactalbumin DNA sequence Optimized
ATGCGATTCTTCGTCCCCCTGTTTCTGGTCGGTATTCTGTTCCCCGCCATCCTCGCCAAGCAGTTCACCAAGTGCGAGCTGTCCCAGCTGCTGAAGGACATCGACGGATACGGTGGTATCGCCCTGCCCGAGCTGATTTGCACCATGTTCCACACCTCTGGATACGACACCCAGGCCATCGTCGAGAACAACGAGTCCACTGAGTACGGCCTGTTCCAGATTTCCAACAAGCTGTGGTGCAAGTCTTCTCAGGTTCCTCAGTCCCGAAACATCTGCGACATTTCTTGCGACAAGTTCCTGGACGACGACATCACCGACGACATCATGTGCGCCAAGAAGATTCTGGACATCAAGGGTATCGACTACTGGCTGGCCCACAAGGCCCTGTGCACCGAGAAGCTGGAGCAGTGGCTGTGCGAGAAGCTG
In your own words, describe why you need to optimize codon usage. Which organism have you chosen to optimize the codon sequence for and why?
Codon optimization is needed in case a sequence contains too many rare tRNAs (which, may be common from the original organism), which could mess up with protein translation; it also may impact the protein’s stability depending on the cellular context, specifically, the pH, so having compatible codon’s for once the protein’s done is also something to keep in mind.
3.4. You have a sequence! Now what?
What technologies could be used to produce this protein from your DNA? Describe in your words the DNA sequence can be transcribed and translated into your protein. You may describe either cell-dependent or cell-free methods, or both.
First, it needs a promoter, and a terminator; once that’s edited into the sequence, it can be inserted into a yeast (I chose yeast and not a bacteria because there are some post-translational-modifications that bacteria is unable to do!)
Oh, also, it would be a plasmid. In order to insert this plasmid, a method that can be used is Heat Shock of Yarrowia lipolytica (forgive me if I forget to name the organism in italics). Would just need to follow the protocol, but also design a selection method, such as the introduction of an antibiotic-resistant gene, or the usage of a strain that lacks the capability of producing a metabolite that is essential for metabolism, transform, and then pass it onto a medium that also does not have said metabolite.
Part 4: Prepare a Twist DNA Synthesis Order
4.1. Create a Twist account and a Benchling account
Done!
4.2. Build Your DNA Insert Sequence
https://benchling.com/s/seq-JUNnre8HZRwtYLbAV7ER?m=slm-NLI7GfaPtF5eARNkdXdJ
Done also!
4.3. On Twist, Select The “Genes” Option
Mhm
4.4. Select “Clonal Genes” option
Yep
4.5. Import your sequence
Done!
4.6. Choose Your Vector
I chose pTwist Amp High Copy - (2221bp)
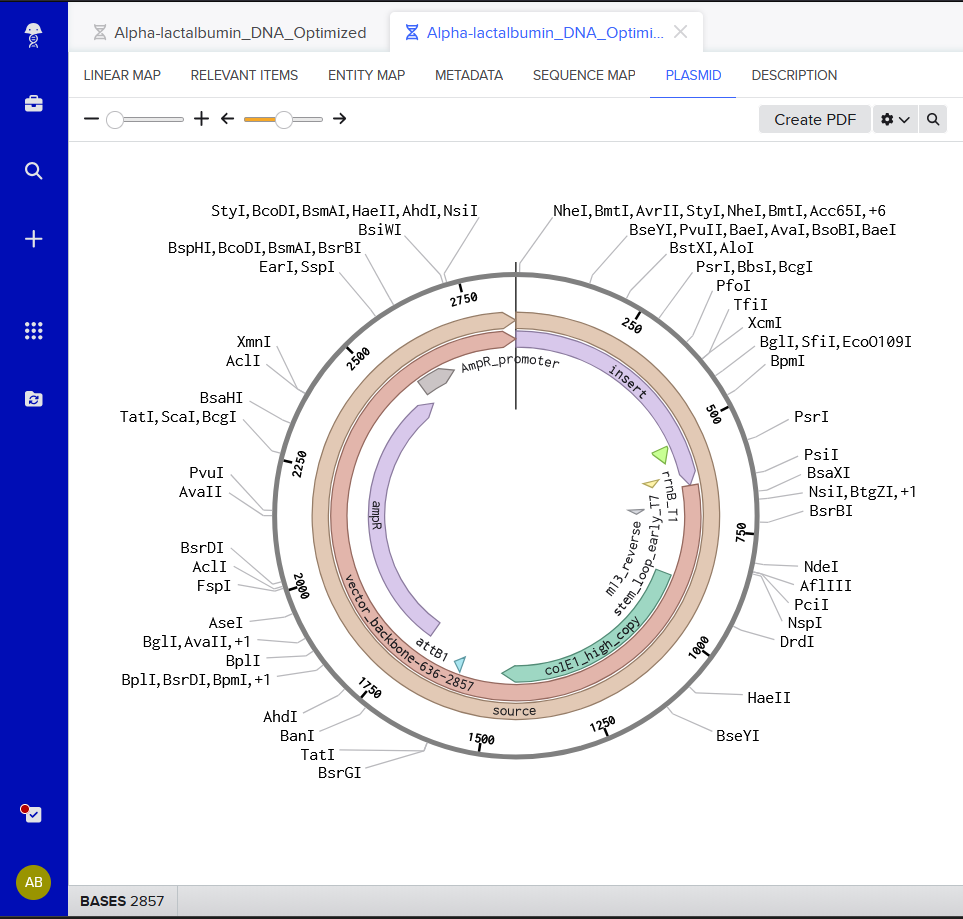

My, hopefully functional, plasmid..
Part 5: DNA Read/Write/Edit
5.1 DNA Read
(i) What DNA would you want to sequence (e.g., read) and why? This could be DNA related to human health (e.g. genes related to disease research), environmental monitoring (e.g., sewage waste water, biodiversity analysis), and beyond (e.g. DNA data storage, biobank).
I think something interesting would be the DNA of all, if not, multiple flowers that produce the color blue. Blue is quite a rare color in nature, and it would be wonderful if it could be figured out what genes are responsible for the blue of the flower… the reason why is because, with genetic engineering, surely blue cotton could be possible!
(ii) In lecture, a variety of sequencing technologies were mentioned. What technology or technologies would you use to perform sequencing on your DNA and why? Also answer the following questions:
Illumina’s sequencing-by-synthesis. It has a great output and, because plant genomes are immense, I think a technology that can do a whole-genome sequencing would be the best fit for the DNA I want to sequence as answered above.
Is your method first-, second- or third-generation or other? How so?
Second generation! Because it’s a high output and massive-in-parallel sequencing method
What is your input? How do you prepare your input (e.g. fragmentation, adapter ligation, PCR)? List the essential steps.
Purified plant DNA, quantify DNA (with nanodrop), fragmentation, NGS protocol (usage of adapters and then the fragments are amplified with PCR. Then this library is ready for the sequencer).
What are the essential steps of your chosen sequencing technology, how does it decode the bases of your DNA sample (base calling)?
Using the sequencer, the DNA fragments are absorbed by the flow cell, new strands of DNA (one base at a time) are synthesized with fluorescent labeled nucleotides. Given that each nucleotide will have a different color, each new base added will be captured.
What is the output of your chosen sequencing technology?
A digital file with reads, which are these sequences that are meant to be put together to reconstruct the whole genome.
5.2 DNA Write
(i) What DNA would you want to synthesize (e.g., write) and why? These could be individual genes, clusters of genes or genetic circuits, whole genomes, and beyond. As described in class thus far, applications could range from therapeutics and drug discovery (e.g., mRNA vaccines and therapies) to novel biomaterials (e.g. structural proteins), to sensors (e.g., genetic circuits for sensing and responding to inflammation, environmental stimuli, etc.), to art (DNA origamis). If possible, include the specific genetic sequence(s) of what you would like to synthesize! You will have the opportunity to actually have Twist synthesize these DNA constructs! :)
I would want to synthesize a mRNA vaccine that uses the principle of those superadjuvant HIV vaccines, but the immunogen would be a cancer molecule. It would be interesting to see the effects of bnAbs against cancer.
(ii) What technology or technologies would you use to perform this DNA synthesis and why?
A DNA plasmid into RNA with the use of phage RNA polymerase. The reason why is because it is an mRNA vaccine, so that polymerase is very essential.
Also answer the following questions:
What are the essential steps of your chosen sequencing methods?
Design the plasmid template with compatible elements like the promoter for the phage RNA polymerase. Then, do in vitro transcription.
What are the limitations of your sequencing method (if any) in terms of speed, accuracy, scalability?
This is a method that is relatively simple and does not really require much speed given that it is a small synthesis (in terms of nucleotides), accurate-wise it can be ~70-80% because capping sometimes can be incomplete. Since it is an enzymatic in vitro process, it is suitable to be scaled up pretty quickly.
5.3 DNA Edit
(i) What DNA would you want to edit and why? In class, George shared a variety of ways to edit the genes and genomes of humans and other organisms. Such DNA editing technologies have profound implications for human health, development, and even human longevity and human augmentation. DNA editing is also already commonly leveraged for flora and fauna, for example in nature conservation efforts, (animal/plant restoration, de-extinction), or in agriculture (e.g. plant breeding, nitrogen fixation). What kinds of edits might you want to make to DNA (e.g., human genomes and beyond) and why?
I would want to edit the p53 gene. I believe that elephants do not have risky rates of cancer despite their size/number of cells, and that’s because (If I am not misremembering) they have a bunch of p53 copies. Reason I would do this is for the better health of a lot of people.
(ii) What technology or technologies would you use to perform these DNA edits and why?
CRISPR. Because I think this would be the safest option given how many gene editing therapies that use CRISPR are being approved lately.
Also answer the following questions:
How does your technology of choice edit DNA? What are the essential steps?
CRISPR-Cas9 uses a guide RNA that matches a specific DNA sequence, and it has the enzyme Cas9 which is directed by said guide to the target. This enzyme makes a double-strand break, and then the cell tries to repair this break, while it is repairing it, there can be the insertion of a corrected sequence if a repair template is provided.
What preparation do you need to do (e.g. design steps) and what is the input (e.g. DNA template, enzymes, plasmids, primers, guides, cells) for the editing?
Target DNA is identified in order to design the guide RNA. The input includes Cas9, guide RNA and donor DNA template.
What are the limitations of your editing methods (if any) in terms of efficiency or precision?
I’m not aware of any as CRISPR seems to be described as a very efficient and precise gene editing tool in literature.