Week 6 — Genetic Circuits Part I: Assembly Technologies
Answer these questions about the protocol in this week’s lab:
What are some components in the Phusion High-Fidelity PCR Master Mix and what is their purpose?
As stated in the thermofisher website, “Phusion DNA Polymerase, nucleotides, and optimized reaction buffer including MgCl2” The Phusion DNA Polymerase will be the one replicating the DNA at a really high fidelity The nucleotides will be ‘inserted’ into the replicated DNA strands And the reaction buffer (Including MgCl2, which is a co-factor needed for the enzyme), which is optimized, meaning, it has the favorable conditions for the enzyme such as pH.
What are some factors that determine primer annealing temperature during PCR?
The GC content %, the amount of nucleotides, self-dimers, and homodimers
There are two methods from this class that create linear fragments of DNA: PCR, and restriction enzyme digests. Compare and contrast these two methods, both in terms of protocol as well as when one may be preferable to use over the other.
PCR uses the thermomixer, which is a machine that controls the temperature of a minitube at 3 distinct temperatures: denaturation, primer annealing, and extension. This is going to be the biggest difference in protocols compared to restriction enzyme digestions. As for when PCR might be preferable: it is when we have a physical nucleotidic sequence to amplify, as in, we don’t have enough copies for whatever we may want to do with it.
As for restriciton enzyme digests, this would only really take place with one temperature; the optimal temperature for our restriction enzyme(s). No thermomixer needed. As for when we would prefer this method; it would be when we have a lot of a nucleotidic sequence, but it is still within our sample, maybe within a plasmid. So in order to isolate the nucleotides we really care about, we use digestion enzymes.
How can you ensure that the DNA sequences that you have digested and PCR-ed will be appropriate for Gibson cloning?
The ends of fragments have to have homologous overlaps, around 20-40 bp of matching sequence (this includes primers to add those overlap regions)
How does the plasmid DNA enter the E. coli cells during transformation?
The membrane is more permeable, and this can be through different methods like electroporation, heatshock, and chemical transformation.
Describe another assembly method in detail (such as Golden Gate Assembly)
By using IIs restriction enzymes together with a ligase, multiple fragments are assembled in order and in a single reaction. This is done by having the enzyme cut fragments that generate overgangs, and complementary overhangs annel between adjacent fragments, the ligase seals them (after this, original restriction sites are removed, so they’re not digested again). This method too has its precautions, such as, fragments that have overhangs designed to be unique, so wrong annealing doesn’t happen.
Explain the other method in 5 - 7 sentences plus diagrams (either handmade or online).
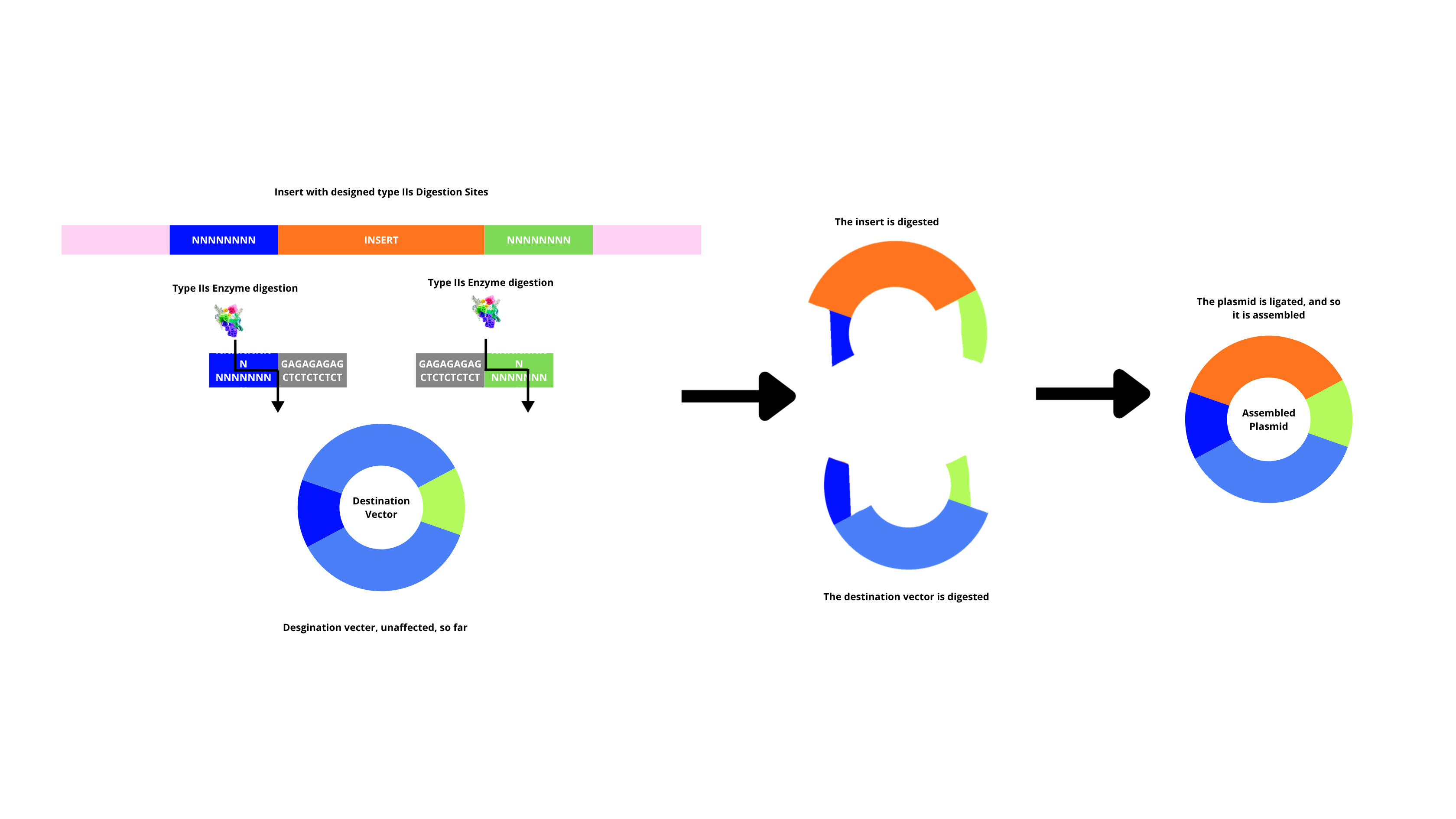

Model this assembly method with Benchling or Asimov Kernel!
Part 2 of this week’s homework
Create a Repository for your work
Create a blank Notebook entry to document the homework and save it to that Repository
Explore the devices in the Bacterial Demos Repo to understand how the parts work together by running the Simulator on various examples, following the instructions for the simulator found in the “Info” panel (click the “i” icon on the right to open the Info panel)
Create a blank Construct and save it to your Repository
Recreate the Repressilator in that empty Construct by using parts from the Characterized Bacterial Parts repository
Search the parts using the Search function in the right menu
Drag and drop the parts into the Construct
Confirm it works as expected by running the Simulator (“play” button) and compare your results with the Repressilator Construct found in the Bacterial Demos repository
Document all of this work in your Notebook entry - you can copy the glyph image and the simulator graphs, and paste them into your Notebook
Build three of your own Constructs using the parts in the Characterized Bacterials Parts Repo
Explain in the Notebook Entry how you think each of the Constructs should function
Run the simulator and share your results in the Notebook Entry
If the results don’t match your expectations, speculate on why and see if you can adjust the simulator settings to get the expected outcome