Week 7 HW: Genetic Circuits Part II: Neuromorphic Circuits
Assignment Part 1: Intracellular Artificial Neural Networks (IANNs)
What advantages do IANNs have over traditional genetic circuits, whose input/output behaviors are Boolean functions?
Scalability for one, and for second, way more outputs; comparing this to, for example, the lac operon, if I recall correctly it only has a represor, an operator, and the lactose gene. IANNs would have way more repressor, operators, and so way more outputs, and ways said outputs are regulated, by multiple inputs.
Describe a useful application for an IANN; include a detailed description of input/output behavior, as well as any limitations an IANN might face to achieve your goal.
It would be an interesting way to make a diagnosis method by using IANNs, specifically with diseases which may have slight genetic variations; let’s say, there’s a disease that has the same symptoms on a person, but there are 5 variations of it, so the 5 biomarkers are slightly different, with other biomarkers needed too, in order to confirm it really is this disease. So, set of biomarkers needed to be detected, but one of the biomarkers can have 5 variations.
With IANNs, upon receiving the input of all the biomarkers characterized, and the variable biomarker (let’s say, variation n°1), the output could be one fluorescence color. As for, now, variation n°2, same other biomarkers as input, and thanks to this v.n°2, the output will be different, could be luminescence now.
The limitations would definitely be the different responses the are in biology (not all cells will have the exact same response despite having the same genetic components), and personally, defining the threshold needed for the inputs.
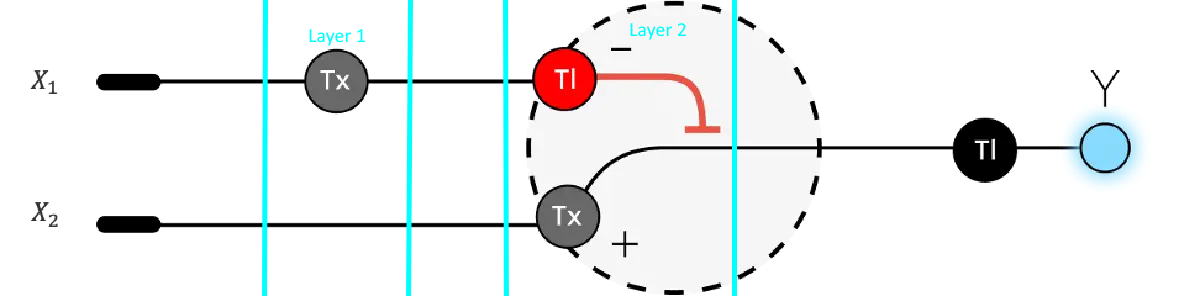
Below is a diagram depicting an intracellular single-layer perceptron where the X1 input is DNA encoding for the Csy4 endoribonuclease and the X2 input is DNA encoding for a fluorescent protein output whose mRNA is regulated by Csy4. Tx: transcription; Tl: translation. Draw a diagram for an intracellular multilayer perceptron where layer 1 outputs an endoribonuclease that regulates a fluorescent protein output in layer 2.
My edition, as to my understanding, this is already kind of happening:

Assignment Part 2: Fungal Materials
What are some examples of existing fungal materials and what are they used for? What are their advantages and disadvantages over traditional counterparts?
I know that they’re used for paints, coatings, and textile dyeing! (Venil, 2020) Some of the fungal materials produced are namely: carotenoids, melanins, flavins, phenazines, quinones, and monascins.
As for advantages, they’re not season-dependant like plants, they’re more intense, and even costs are better (Lagashetti et al., 2019)
In literature, there doesn’t seem to be much disadvantages reported.
What might you want to genetically engineer fungi to do and why? What are the advantages of doing synthetic biology in fungi as opposed to bacteria?
For one, the limitation of hyperglicosilation, which is not available to bacteria. Secondly, the exploitation of pathways that some bacteria do not have, for example, the acetyl-CoA pathway can be taken advantage from, from .S. cerevisiae to avoid competing intermediates, like Yuan and Ching did (2016).
Assignment Part 3: First DNA Twist Order
Review the Individual Final Project documentation guidelines.
Submit this Google Form with your draft Aim 1, final project summary, HTGAA industry council selections, and shared folder for DNA designs. DUE MARCH 20 FOR MIT/HARVARD/WELLESLEY STUDENTS
The first aim of my final project is to genetically engineer autologous hematopoietic stem cells to carry predefined anti-HIV Env broadly neutralizing antibody heavy- and light-chain genes, so that their B-cell progeny express an HIV-specific broadly neutralizing B-cell receptor, by utilizing CRISPR-based gene editing, donor DNA design for bnAb heavy- and light-chain insertion, ex vivo HSC culture, and downstream differentiation and molecular validation assays
The following is what I’ve submitted in the Google Form.
The steps/protocols:
Used sequence in order to knock-in heavy and light chains genes https://pmc.ncbi.nlm.nih.gov/articles/PMC10140475/
Extraction and purification of CD34+ https://pubmed.ncbi.nlm.nih.gov/33039133/
Homologous recombination in human HSPCs using CRISPR/Cas9 plus rAAV6 delivery https://pmc.ncbi.nlm.nih.gov/articles/PMC5826598/#S31
After genetic engineering, HSPC culturing https://pmc.ncbi.nlm.nih.gov/articles/PMC11877881/
Downstream differentiation of CD34+ to B-cell differentiation https://www.nature.com/articles/s41598-025-30732-9
Molecular assays for validation https://pubmed.ncbi.nlm.nih.gov/40773349/

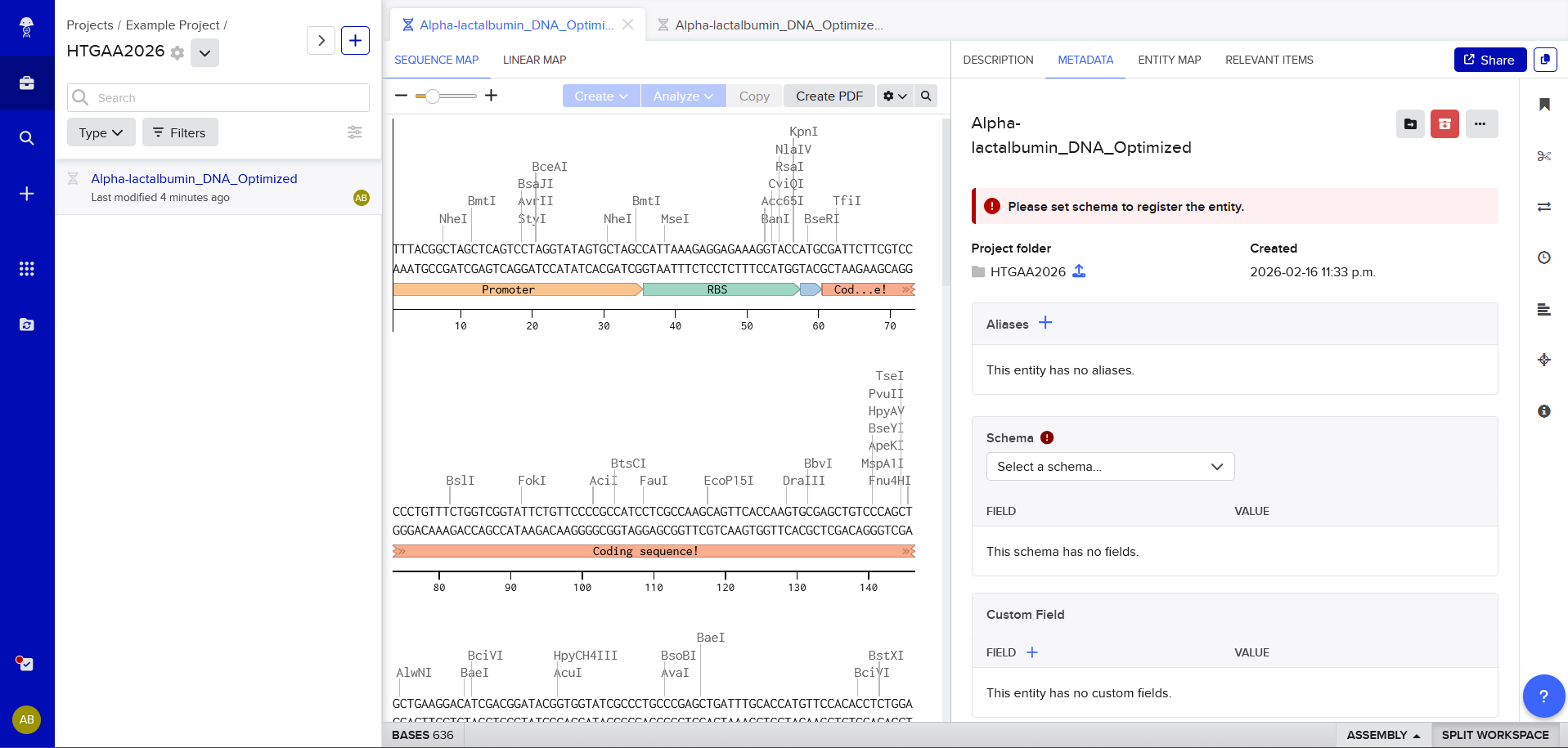
Review Part 3: DNA Design Challenge of the week 2 homework. Design at least 1 insert sequence and place it into the Benchling/Kernel/Other folder you shared in the Google Form above. Document the backbone vector it will be synthesized in on your website.
https://benchling.com/s/seq-JUNnre8HZRwtYLbAV7ER?m=slm-NLI7GfaPtF5eARNkdXdJ
Bibliographic references:
Lagashetti, A. C., Dufossé, L., Singh, S. K., & Singh, P. N. (2019). Fungal Pigments and Their Prospects in Different Industries. Microorganisms, 7(12), 604. https://doi.org/10.3390/microorganisms7120604
Venil, C. K., Velmurugan, P., Dufossé, L., Devi, P. R., & Ravi, A. V. (2020). Fungal Pigments: Potential Coloring Compounds for Wide Ranging Applications in Textile Dyeing. Journal of fungi (Basel, Switzerland), 6(2), 68. https://doi.org/10.3390/jof6020068
Yuan J., Ching C. B. (2016). Mitochondrial acetyl-CoA utilization pathway for terpenoid productions. Metab. Eng. 38, 303–309. 10.1016/j.ymben.2016.07.008