Week 10 — Advanced Imaging & Measurement Technology
Homework
Homework is partly based on data that will be generated in the Waters Immerse Lab in Cambridge, MA. Students will characterize green fluorescent protein (eGFP, a recombinant protein standard) structure (primary, secondary/tertiary) in the lab using liquid chromatography and mass spectrometry, as well as Keyhole Limpet Hemocyanin (KLH) oligomeric states using charge detection mass spectrometry (CDMS). Data generated in the lab needed to do the homework is included both within this document and in the Appendix of the laboratory protocol.
Homework: Final Project
For your final project:
Please identify at least one (ideally many) aspect(s) of your project that you will measure. It could be the mass or sequence of a protein, the presence, absence, or quantity of a biomarker, etc.
- The integration of the VRC01 sequence in the hematopoietic cells.
Please describe all of the elements you would like to measure, and furthermore describe how you will perform these measurements.
- VRC01 sequence (bnAbs that target Env) in the hematopoietic cells
- A GFP reporter gene
What are the technologies you will use (e.g., gel electrophoresis, DNA sequencing, mass spectrometry, etc.)? Describe in detail.
- Flow cytometry we can get to know how many cells have been genetically modified.
- PCR can be done too, targetting the construct we’ve inserted with primers designed just for that.
Homework: Waters Part I — Molecular Weight
We will analyze an eGFP standard on a Waters Xevo G3 QTof MS system to determine the molecular weight of intact eGFP and observe its charge state distribution in the native and denatured (unfolded) states. The conditions for LC-MS analysis of intact protein cause it to unfold and be detected in its denatured form (due to the solvents and pH used for analysis).
Based on the predicted amino acid sequence of eGFP (see below) and any known modifications, what is the calculated molecular weight? You can use an online calculator like the one at https://web.expasy.org/compute_pi/
eGFP Sequence: MVSKGEELFTG VVPILVELDG DVNGHKFSVS GEGEGDATYG KLTLKFICTT GKLPVPWPTL VTTLTYGVQC FSRYPDHMKQ HDFFKSAMPE GYVQERTIFF KDDGNYKTRA EVKFEGDTLV NRIELKGIDF KEDGNILGHK LEYNYNSHNV YIMADKQKNG IKVNFKIRHN IEDGSVQLAD HYQQNTPIGD GPVLLPDNHY LSTQSALSKD PNEKRDHMVL LEFVTAAGIT LGMDELYKLE HHHHHH Note: This contains a His-purification tag (HHHHHH) and a linker (the LE before it).
The molecular weight is 28006.60
Calculate the molecular weight of the eGFP using the adjacent charge state approach described in the recitation. Select two charge states from the intact LC-MS data (Figure 1) and:
1. Determine for each adjacent pair of peaks using (n,n + 1):
Chosen values are: m/zn = 903.7148 m/zn+1 = 875.4421
= 903.7148
Substitution of the values z = 875.4421 / (903.7148 - 875.4421)
z = 875.4421 / (28.2727)
z = 30.96, but we’ll round it up to 31
2. Determine the MW of the protein using the relationship between m/zn, MW and z
Now, MW = zn (m / zn - H) is the formula we’re gonna use
MW = 31 (903.7148 − 1)
MW = 31 (902.7148)
MW = 27,984.15Da
MW = 27.98KDa
Quite close to 28
3. Calculate the accuracy of the measurement using the deconvoluted MW from 2.2 and the predicted weight of the protein from 2.1 using:
Accuracy = (MW experiment - MW theory) / MW theory
Substituting these would be
Accuracy = |(27,984.15 - 28,000)| / 28,000
Accuracy = 15.85 / 28,000
Accuracy = 0.000566, as in, error of 0.000566%
Homework: Waters Part II — Secondary/Tertiary structure
Optional, so I’ll skip on it for now. I do have plans on coming back to it but thesis and meetings in the laboratory + classes have kept me busy!
Homework: Waters Part III — Peptide Mapping - primary structure
We will digest the eGFP protein standard into peptides using trypsin (an enzyme that selectively cleaves the peptide bond after Lysine (K) and Arginine (R) residues. The resulting peptides will be analyzed on the Waters BioAccord LC-MS to measure their molecular weights and fragmented to confirm the amino acid sequence within each peptide – generating a “peptide map”. This process is used to confirm the primary structure of the protein.
There are a variety of tools available online to calculate protein molecular weight and predict a list of peptides generated from a tryptic digest. We will be using tools within the online resource Expasy (the bioinformatics resource portal of the Swiss Institute of Bioinformatics (SIB)) to predict a list of tryptic peptides from eGFP.
- How many Lysines (K) and Arginines (R) are in eGFP? Please circle or highlight them in the eGFP sequence given in Waters Part I question 1 above. (Note: adding the sequence to Benchling as an amino acid file and clicking biochemical properties tab will show you a count for each amino acid).
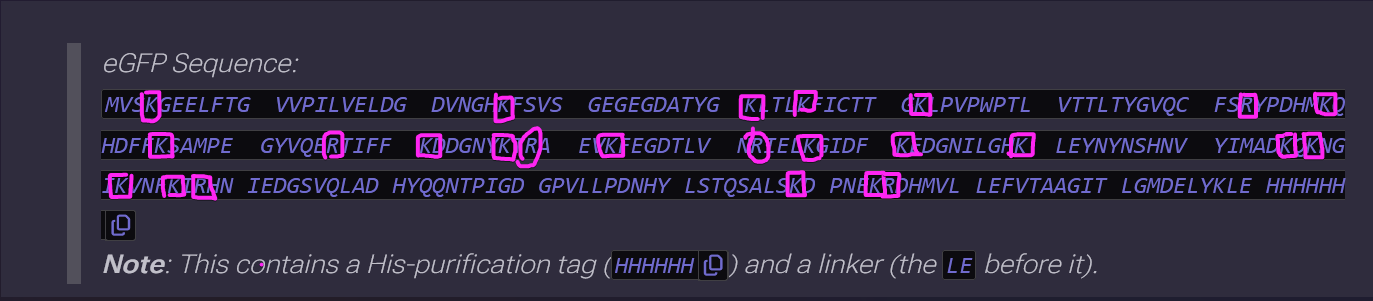

There’s 21 Lysines (K) and 5 Arginines (R)
- How many peptides will be generated from tryptic digestion of eGFP?
27, and it’s not 26 because, let’s say there’s only 1 lysine 0 arginines; we’d have 2 peptides in the end.
1. Navigate to https://web.expasy.org/peptide_mass/
2. Copy/paste the sequence above into the input box in the PeptideMass tool to generate expected list of peptides.
3. Use Figure 4 below as a guide for the relevant parameters to predict peptides from eGFP.
4. Click “Perform the Cleavage” button in the PeptideMass tool and report the number of peptides generated when using trypsin to perform the digest.
Oh, 19 peptides
- Based on the LC-MS data for the Peptide Map data generated in lab (please use Figure 5a as a reference) how many chromatographic peaks do you see in the eGFP peptide map between 0.5 and 6 minutes? You may count all peaks that are >10% relative abundance.
23 peaks
I’ll resume the next parts the next day