Group Final Project
Hypothesis: Substitution of a bacteriophage’s replisome with an orthogonal T7 replisome for continuous hypermutation directed towards stability
The idea of a proposal comes from an article by Diercks et al., 2024, in which they use a very faulty replisome that induces hypermutation, in which they direct towards a very high antibiotic resistance.
Bacteriophage engineering
Challenge chosen: Stability
Group members
Alan Bravo https://pages.htgaa.org/2026a-alan-bravo/ Samudera Mukhalid https://pages.htgaa.org/2026a-samudera-mukhalid/
Choose one or two main goals from the list that you think you can address computationally (e.g., “We’ll try to stabilize the lysis protein,” or “We’ll attempt to disrupt its interaction with E. coli DnaJ.”).
Goal #1: We’ll try to make the bacteriophage more resistant to varying environmental conditions such as temperature and pH. Goal #2: We’ll try to reach this stability without compromising the bacteriophage’s infectivity
Write a 1-page proposal (bullet points or short paragraphs) describing:
Which tools/approaches from recitation you propose using (e.g., “Use Protein Language Models to do in silico mutagenesis, then AlphaFold-Multimer to check complexes.”).
Why do you think those tools might help solve your chosen sub-problem?
Name one or two potential pitfalls (e.g., “We lack enough training data on phage–bacteria interactions.”).
Include a schematic of your pipeline.
Proposal
My proposal consists on a bacteriophage based on the T7 replisome, since T7 encodes much of its own replication machinery, a lower-fidelity replisome, we can generate phages that are both faulty, but also excel at our characteristic of interest: stability. By exposing that population to a defined stress linked to stability, and then isolate the phages that remain infectious using plaque-based assays, we can recover viable survivors. After sequencing several independently recovered survivor phages, I would compare their encoded proteins with Clustal Omega, from that, we could build a consensus-style view of which residues remain strongly conserved and which sites may repeatedly change under selection, and so, we can make a construct of, hopefully, a very stable bacteriophage
Two potential pitfalls could be
- Results for clustal-omega could be too divergent, making it difficult to decide on conserved residues
- There could be no stable bacteriophages due to the faultiness of the replisome
Results for clustal-omega
(We’re doing a theoretical alignment, given that we cannot perform the proposed experiment to carry on real alignments)


Schematic
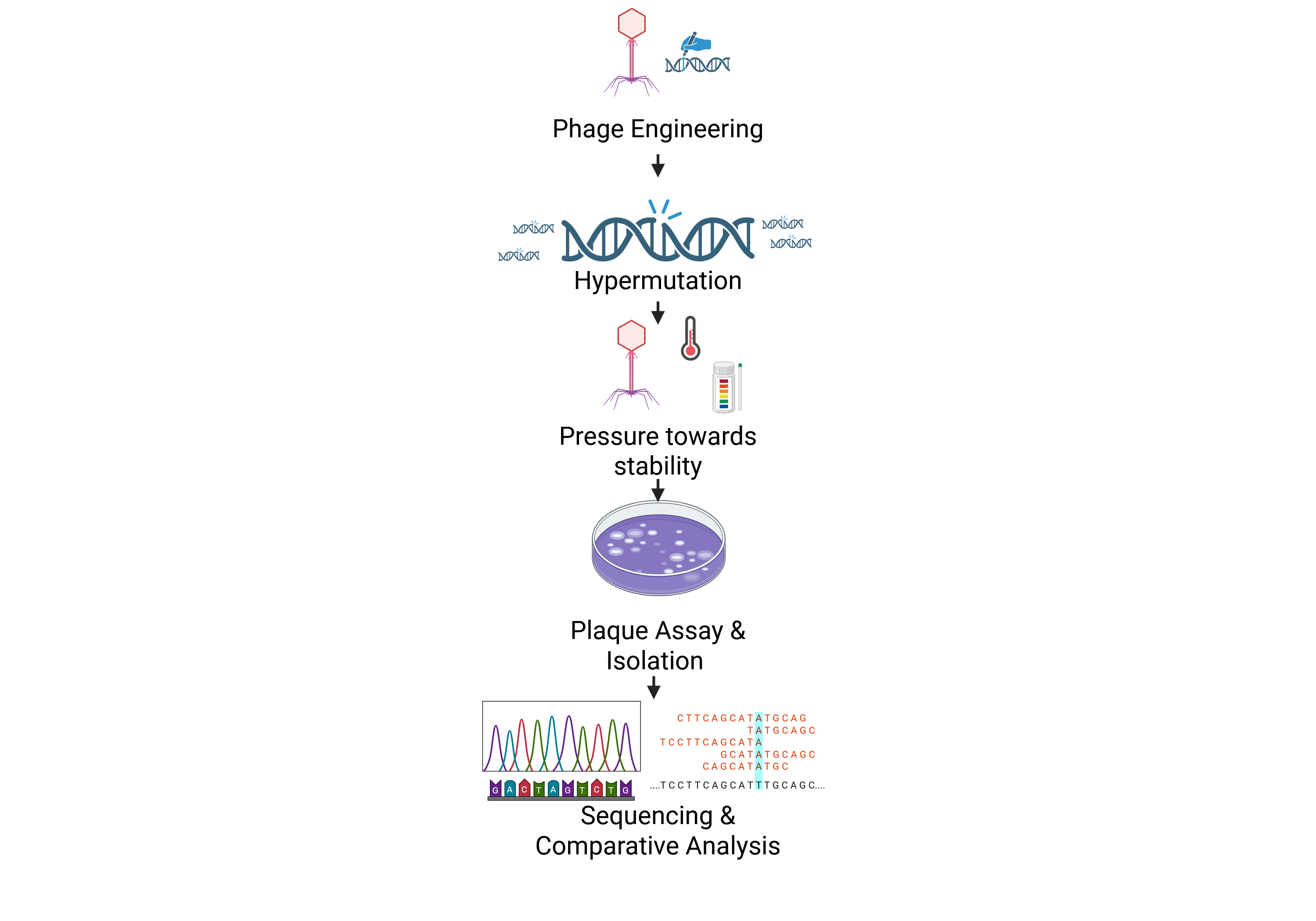

Bibliographic references
Diercks, C. S., Sondermann, P. J., Rong, C., Dik, D. A., Gillis, T. G., Ban, Y., & Schultz, P. G. (2024). An Orthogonal T7 Replisome for Continuous Hypermutation and Accelerated Evolution in E. coli. bioRxiv : the preprint server for biology, 2024.07.25.605042. https://doi.org/10.1101/2024.07.25.605042