Week 1 HW: Principles and Practices
HW1
Ultra-efficient DNA Synthesis Machine
My research is on designing a DNA synthesis machine that can reduce the cost and time to produce long strands of arbitrary DNA sequences. Right now we’re aiming for megabase strands of DNA but the goal of the project is to eventually get to the Gb range and establish a technology that can scale and improve similar to transistors in Moore’s Law. For context, I work mostly on the mechanical side and hope to learn more about the biochemistry and synthetic biology in general from this class.
Most of today’s DNA synthesis tools and services are good at making short DNA pieces, from hundreds to thousands of bps, but as strands get longer, they become much harder and costlier to produce. As an example, Integrated DNA Technologies sells ~125-3,000 bp fragments for uses like cloning single genes and antibody research, the cost is around $0.07-$0.20 per base pair.
Short and mid-length sequences are still really useful:
20–100 bp pieces are used for primers to amplify DNA or guide RNA design
1,000–3,000 bp can code for single proteins or small metabolic pathways
10,000–100,000 bp pieces could represent entire operons
Once hit the megabase range and beyond, you can do a whole lot more:
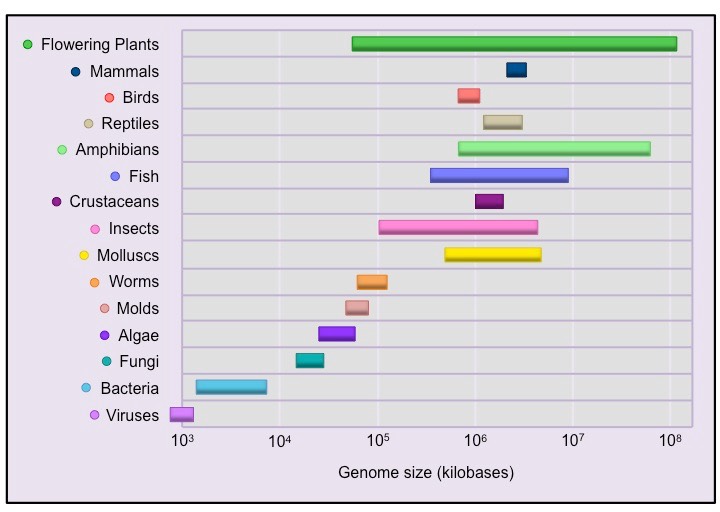
A typical bacterial genome like E. coli is ~1–13Mbp
Yeast has a genome around 12Mbp long
A gigabase approaches the size of large eukaryotic chromosomes
3.2Gbp is the size of the human genome
The goal is not just to achieve longer strands, but to make long, accurate DNA affordable and reliable, opening the door to lots more possibilities.
Governance/Policy
At the gigabase scale, DNA synthesis moves beyond individual genes and can be used to create entire genomes or chromosomes, which raises new safety, security, and ethical considerations. The main governance challenge for lowering the barrier to creating these large constructs is: how this technology and its outputs be developed and deployed responsibly as its capabilities scale.
Goal
Enable the constructive use of large-scale DNA synthesis while preventing misuse or unintended harm enabled by scale and accessibility.
Governance should aim to prevent malfeasance enabled by longer DNA constructs. This involves:
- Preventing the synthesis of harmful DNA sequences
- Reducing the risk posed by unintentional misuse
- Ensuring that increased automation does not eliminate safety checks and that safety checks adapt to the technology
One way to enable this is through system-level design. The hierarchy of controls encourages eliminating risk before having to rely on training or user intent. By incorporating safety checks, containment options, and traceability mechanisms directly into hardware, safe operation becomes the default outcome. Without governance and training, things can still go wrong, and although accessibility is powerful, there should be limits set to ensure compliance and training. In case something goes wrong, there should be an established line of accountability and transparency with oversight.
Governance Actions
To ensure that large-scale DNA synthesis develops in a safe and constructive way there should be a mix of technical, institutional, and regulatory governance actions.
Hardware-Level Constraints on Maximum Assembly Length
Purpose Currently, DNA synthesis limits are largely economic and biochemical but they could be enforced by hardware limits
Design Mechanical or software-enforced limits on assembly length, these would be enforced by governance and added by design choices made by manufacturers and research labs building synthesis platforms.
Assumptions One assumption is that limiting the strand length meaningfully reduces misuse risk. I don’t know where this limit would be set at but it’s possible that you could get around this or that the set length can still lead to misuse. It is also assumed that this wouldn’t hinder legitimate research.
Risks of Failure & “Success” As mentioned before, you could find ways to misuse shorter strands or combine shorter strands into longer ones. Or this length could keep legitimate research from occurring successfully.
Sequence Screening and Logging
Purpose Many DNA synthesis companies perform sequence screening to flag known pathogenic or regulated sequences. If synthesis moves toward in-house and machine-based systems, this screening could still take place or even be extended to logging
Design Automated sequence screening could occur within each synthesis machine with flags dispersed or hardware/software locks enabled if needed. It might also be wise to log synthesized sequences above a certain size threshold. This can be implemented by academic institutions, commercial developers, and funding agencies. When working in MIT nano, your process must be approved by a committee before you begin and EHS reviews new and acceptable chemicals, this would work the same way.
Assumptions This assumes that known harmful sequences can be meaningfully identified. And that users will accept limited logging in exchange for access.
Risks of Failure & “Success” It is possible that screening can miss novel or emergent risks. Or that logging could raise intellectual-property concerns. If “too successful,” logging could discourage exploratory research using these systems.
Tiered Access
Purpose I think one of the best ways to govern this technology is with a tiered access model where synthesis capabilities scale with demonstrated ability, infrastructure, and oversight.
Design You could start with basic access for short and mid-length synthesis, with fairly open access to this. Then more advanced capabilities could be unlocked by agreeing to institutional reviews, trainings, and safety approval, along with oversight by universities or national research bodies. It might even be better to have a centralized location with the extra advanced machines with specialized oversight on them though still granting access.
Assumptions This assumes that governing institutions can fairly evaluate readiness and risk and that training and review improve safety outcomes. It also assumes that access tiers won’t become arbitrary gatekeeping, and won’t come down to the same financial barriers in place now.
Risks of Failure & “Success” This could disadvantage smaller or less well-funded labs, and “success” might slow innovation if approval processes lag behind technology.
| Does the option: | Hardware/Software Constraints | Screening/Logging | Tiered Access |
|---|---|---|---|
| Enhance Biosecurity | |||
| • By preventing incidents | 1 | 2 | 2 |
| • By helping respond | 3 | 1 | 2 |
| Foster Lab Safety | |||
| • By preventing incidents | 2 | 2 | 2 |
| • By helping respond | 3 | 1 | 2 |
| Protect the environment | |||
| • By preventing incidents | 1 | 3 | 2 |
| • By helping respond | 3 | 2 | 2 |
| Other considerations | |||
| • Minimizing costs and burdens to stakeholders | 1 | 3 | 2 |
| • Feasibility? | 1 | 2 | 2 |
| • Not impede research | 2 | 3 | 2 |
| • Promote constructive applications | 2 | 2 | 1 |
Recommended Governance Approach and Trade-offs
Based on the scoring, I would prioritize a combined approach using sequence screening and logging (Option 2) and tiered access (Option 3). Together, these options provide the strongest balance between preventing misuse, enabling response if something goes wrong, and still enabling legitimate research.
Option 2 performs best in terms of biosecurity and response. As DNA synthesis moves toward in-house, automated, and large-scale systems, maintaining some form of sequence screening becomes increasingly important. Screening and logging help ensure that synthesis at larger scales includes visibility and accountability. It also enables backtracking to find problems or errors, which is important when failures or misuse may not be immediately obvious.
Option 3 complements this by recognizing that not all synthesis capabilities carry the same level of risk. A tiered access model allows safe DNA synthesis to remain relatively open, while reserving the most powerful capabilities for users and institutions with appropriate infrastructure, training, and oversight.
Week 2 Lecture Prep
Homework Questions from Professor Jacobson:
- 1: DNA polymerase has an error rate of 1:10^6. With the 3.2 Gb human genome, that’s ~3,200 errors per replication. Biology fixes this discrepancy using polymerase proofreading and the MutS repair system, which functions likea multi-stage error-correction protocol to drop the effective mutation rate.
- 2: If an average human protein has around 400 amino acids, then redundancy allows for 400!/(20!)20 which is on the order of ~10501 DNA sequences to code for an average protein. But many versions fail because RNA secondary structures physically block ribosomes, or sequences trigger RNA cleavage and codon bias issues.
Homework Questions from Dr. LeProust:
- 1: Phosphoramidite synthesis
- 2: Synthesis hits a wall due to an exponential yield drop. Following the (1 - error rate)^N, curve, the probability of a “perfect” strand decreases with every base added.
- 3: At 2kb, the yield of perfect strands is basically zero. To reach the Gb range, we have to assemble smaller, verified oligos using PCA or Gibson Assembly rather than making them in one shot.
Homework Question from George Church:
- 1: Phenylalanine, Valine, Threonine, Tryptophan, Isoleucine, Methionine, Histidine, Arginine, Leucine, and Lysine
- 2: As a fail-safe, the Jurassic Park lysine contingency is flawed because all animals are already naturally unable to synthesize lysine; they get it from food. An escaped organism would do the same and just find lysine-rich food.
Sources
- https://www.reddit.com/r/JurassicPark/comments/12ib7to/the_lysine_contingency_makes_no_sense_right/
- image credit: Bioninja (https://old-ib.bioninja.com.au/standard-level/topic-3-genetics/32-chromosomes/genome-size.html)
AI Prompts
- “how long is an average human protein?”
- “give me some examples of what certain lengths of DNA can achieve?”