Week 2 HW: DNA Read, Write, and Edit

HW2
Part 1: Benchling & In-silico Gel Art
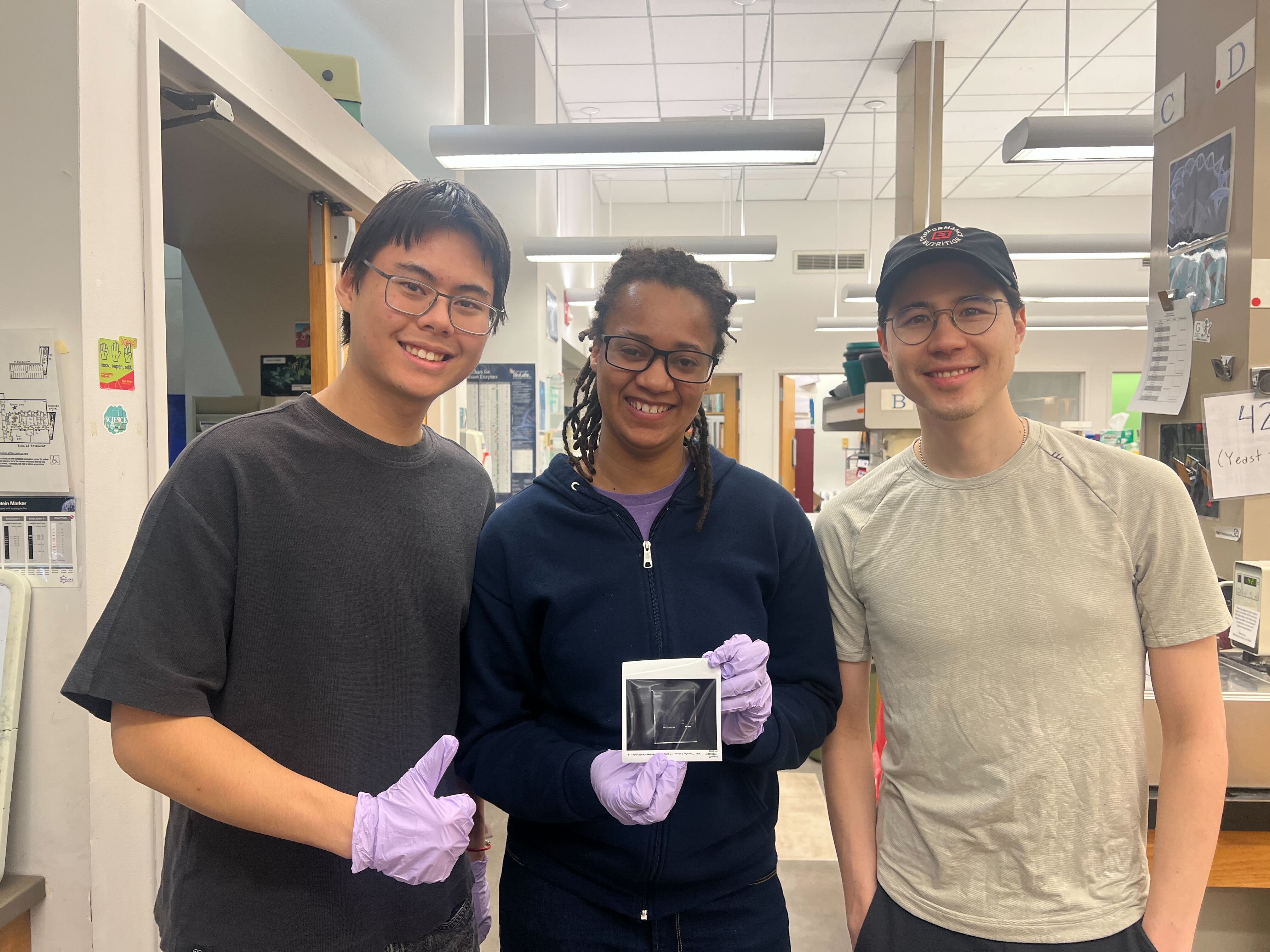
This week, we made gel electrophoresis art using Lambda phage DNA and ten restriction enzymes. Gel electrophoresis uses a positive charge to pull negatively charged DNA through a conductive gel. Longer strands move slower and shorter strands move faster meaning that different lengths of DNA fragments will appear as different bars in your gel. To use this in an artistic context we take our input Lambda DNA and cut it to different lengths using different restriction enzymes which allows us to have coarse control over where these bars end up and thus we can make art with it. I have decided to really commit to my favorite animal, turtles, this semester and try to have a turtle-inspired theme to all of my projects. In an ideal world this is what I wanted my gel art to look like.
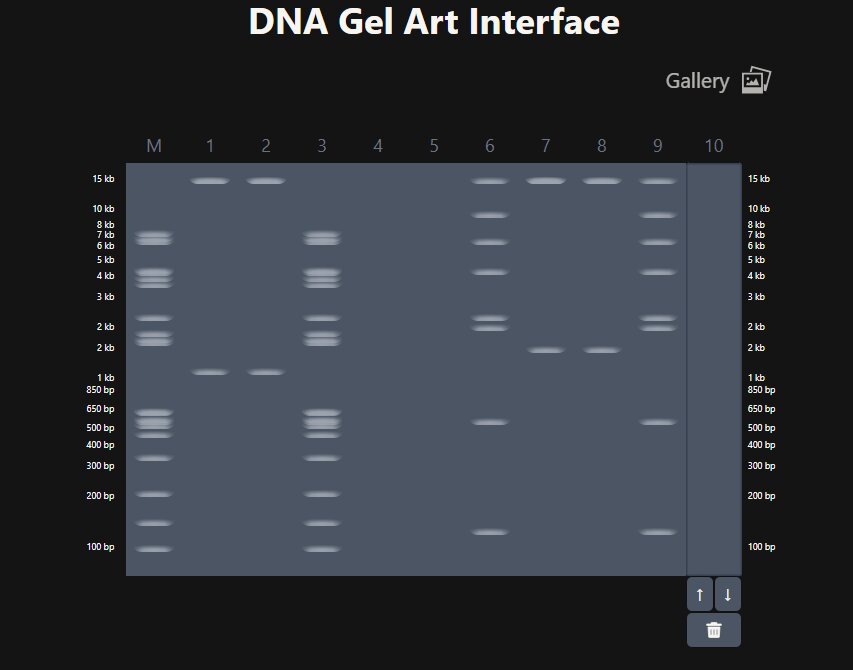

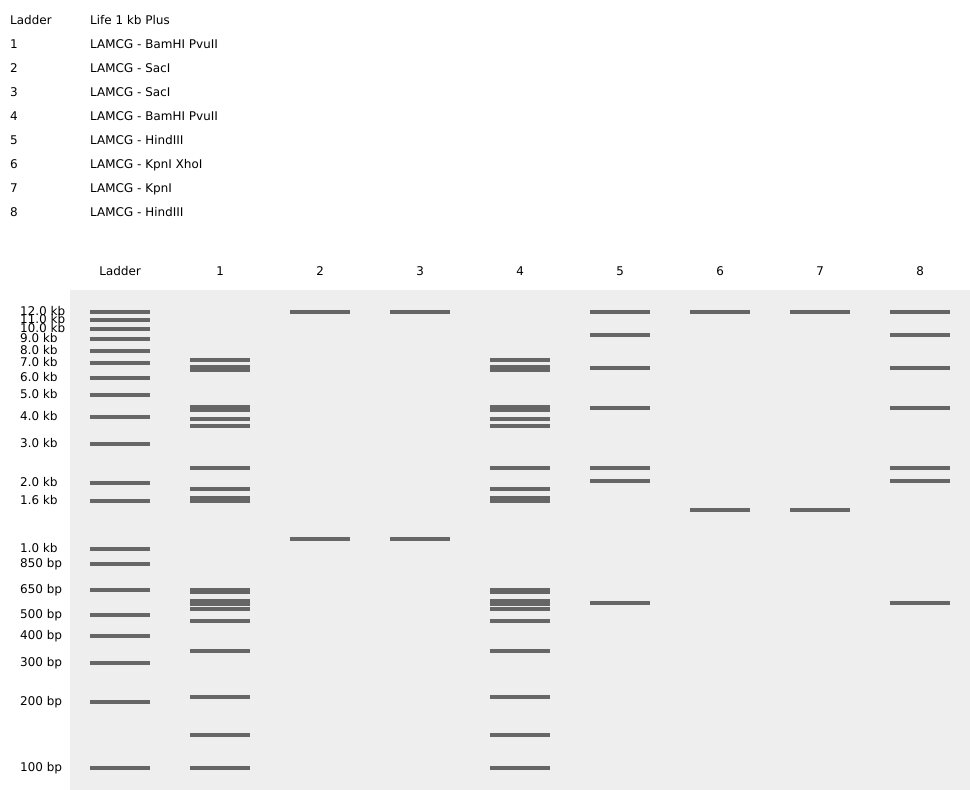

Part 2: Gel Art - Restriction Digests and Gel Electrophoresis
This was my first time going through the whole process of making a gel, but I’ll describe more about the experiment in the lab2 page on my website.
Part 3: DNA Design Challenge
3.1: Choose your protein
For my design challenge homework, I have picked Green Fluorescent Protein (GFP) because I was inspired by the HTGAA website photos, it is great for visualizing gene expression, and I want a second chance at making a turtle this time I want to make a turtle-shaped fluorescence pattern.
Here’s the protein sequence from Uniprot: >sp|P42212|GFP_AEQVI Green fluorescent protein OS=Aequorea victoria OX=6100 GN=GFP PE=1 SV=1 MSKGEELFTGVVPILVELDGDVNGHKFSVSGEGEGDATYGKLTLKFICTTGKLPVPWPTL VTTFSYGVQCFSRYPDHMKQHDFFKSAMPEGYVQERTIFFKDDGNYKTRAEVKFEGDTLV NRIELKGIDFKEDGNILGHKLEYNYNSHNVYIMADKQKNGIKVNFKIRHNIEDGSVQLAD HYQQNTPIGDGPVLLPDNHYLSTQSALSKDPNEKRDHMVLLEFVTAAGITHGMDELYK
3.2: Reverse Translate: Protein (amino acid) sequence to DNA (nucleotide) sequence
I used NCBI to get this reverse translation: >reverse translation of sp|P42212|GFP_AEQVI Green fluorescent protein OS=Aequorea victoria OX=6100 GN=GFP PE=1 SV=1 to a 714 base sequence of most likely codons. atgagcaaaggcgaagaactgtttaccggcgtggtgccgattctggtggaactggatggc gatgtgaacggccataaatttagcgtgagcggcgaaggcgaaggcgatgcgacctatggc aaactgaccctgaaatttatttgcaccaccggcaaactgccggtgccgtggccgaccctg gtgaccacctttagctatggcgtgcagtgctttagccgctatccggatcatatgaaacag catgatttttttaaaagcgcgatgccggaaggctatgtgcaggaacgcaccatttttttt aaagatgatggcaactataaaacccgcgcggaagtgaaatttgaaggcgataccctggtg aaccgcattgaactgaaaggcattgattttaaagaagatggcaacattctgggccataaa ctggaatataactataacagccataacgtgtatattatggcggataaacagaaaaacggc attaaagtgaactttaaaattcgccataacattgaagatggcagcgtgcagctggcggat cattatcagcagaacaccccgattggcgatggcccggtgctgctgccggataaccattat ctgagcacccagagcgcgctgagcaaagatccgaacgaaaaacgcgatcatatggtgctg ctggaatttgtgaccgcggcgggcattacccatggcatggatgaactgtataaa
3.3. Codon optimization
I used a handy website called https://www.novoprolabs.com/tools/codon-optimization to optimize my codon for Escherichia coli (E.coli). I chose this organismbecause it is commonly used in synthetic biology, safe, robust, and easy for people new to the wet lab. Why did I have to optimize it at all? GFP comes from a jellyfish it occurs naturally in that species but not in everything, in order to ensure that it will work well with E.coli I have to pick the codons it prefers (codon bias). This is possible because multiple codons can code for the same amino acid. Codon bias can be a problem if the host organism has low amounts of the matching tRNA. So codon optimization replaces rare codons with preferred codons and can remove unwanted restriction sites. Here’s the optimized sequence: ATGTCTAAAGGCGAAGAACTGTTCACCGGTGTGGTTCCGATCCTGGTCGAGCTGGACGGCGACGTAAACGGCCACAAGT TCTCTGTATCTGGTGAAGGCGAGGGTGATGCAACCTACGGTAAACTGACTCTGAAGTTCATTTGCACTACTGGTAAACT GCCGGTTCCGTGGCCGACTCTGGTCACTACTTTCAGCTACGGTGTACAATGTTTTTCCCGTTACCCGGATCACATGAAG CAGCATGACTTCTTCAAATCTGCTATGCCGGAAGGCTACGTTCAGGAACGCACCATCTTCTTCAAAGACGACGGTAACT ACAAAACTCGCGCTGAGGTTAAGTTTGAAGGCGACACCCTGGTTAATCGTATCGAACTGAAAGGCATTGACTTCAAAGA AGATGGTAACATCCTGGGTCACAAACTGGAATACAACTACAACAGCCATAACGTTTACATCATGGCAGACAAACAGAAA AACGGCATCAAGGTGAACTTCAAAATTCGTCACAATATCGAAGATGGTTCCGTGCAGCTGGCCGATCACTACCAGCAGA ACACTCCGATCGGTGACGGTCCGGTGCTGCTGCCGGACAATCACTATCTGAGCACTCAAAGCGCCCTGAGCAAAGACCC GAACGAAAAACGTGATCACATGGTGCTGCTGGAATTCGTTACCGCGGCAGGCATCACTCACGGCATGGATGAACTGTAT AAA
And stats about what’s changed: CAI before optimization: 0.80 CAI after optimization: 0.83 GC content before optimization: 48.60% GC content after optimization: 49.30%
3.4. You have a sequence! Now what?
Cell-Dependent Expression:
To get the GFP DNA to express in a cell, I would insert the codon-optimized GFP gene into a plasmid and transformed into E. coli. The plasmid includes a promoter that allows the cell to recognize and transcribe the gene. Once inside the bacteria, RNA polymerase transcribes the GFP DNA into mRNA, and ribosomes translate the mRNA into the GFP protein. As the protein folds into its final structure, it begins to fluoresce. In this system, the living cell provides all the machinery needed for transcription and translation.
Cell-Independent Expression:
To get GFP to be produced without living cells, I’d need to mix a cell extract containing ribosomes, enzymes, and tRNAs with the GFP DNA template in a test tube. The extract carries out transcription and translation directly in solution. This allows faster protein production and more control over reaction conditions since there is no need to grow or maintain cells. It is useful for rapid testing of gene designs before moving into full bacterial expression.
Part 4: Prepare a Twist DNA Synthesis Order
I chose a similar protein to the example in class but replaced the sfGFP coding sequence with my regular GFP coding sequence. I wonder what the differences are and if they are different enough to create a pattern with. ANyways here’s my benchling linear map.
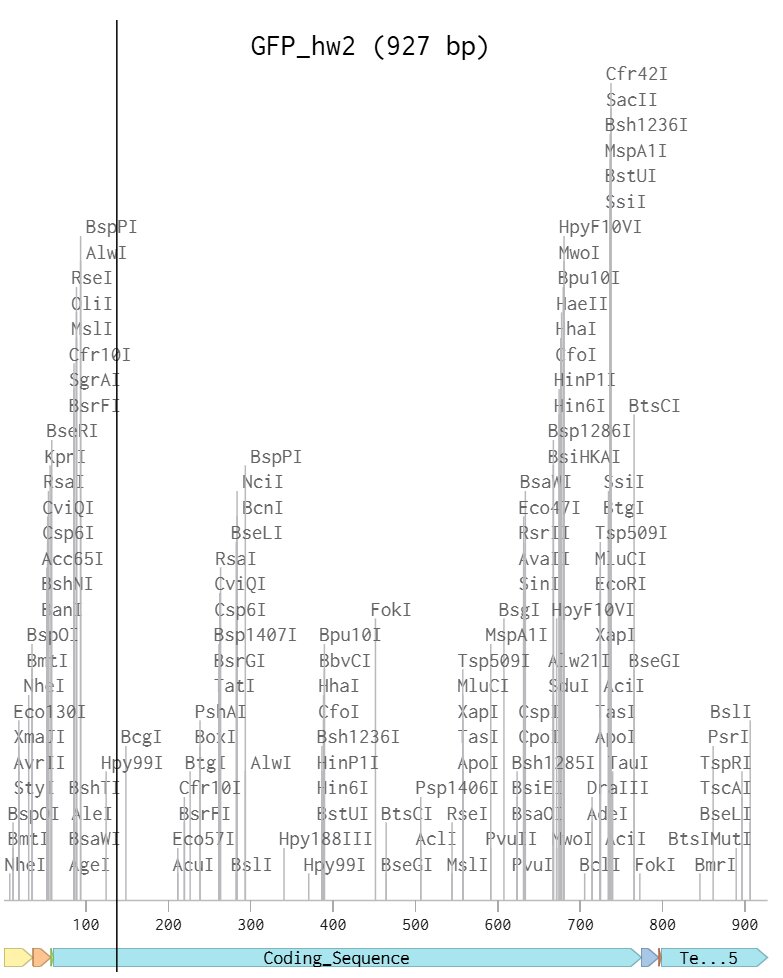

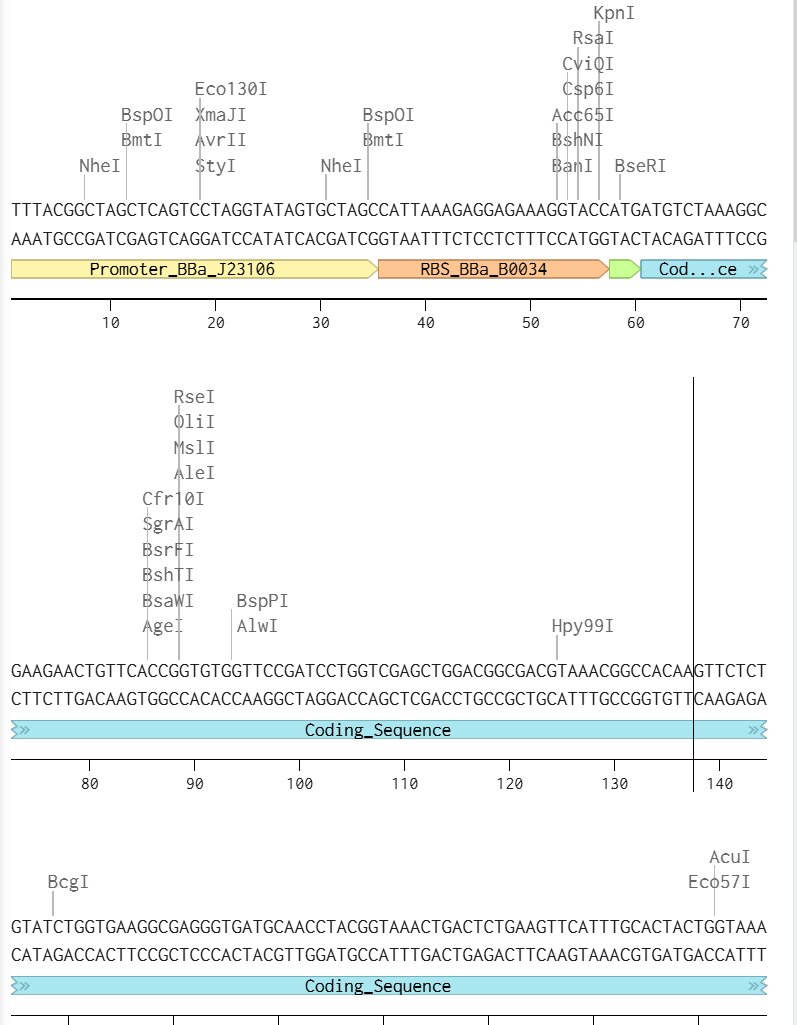
Here’s my final plasmid:
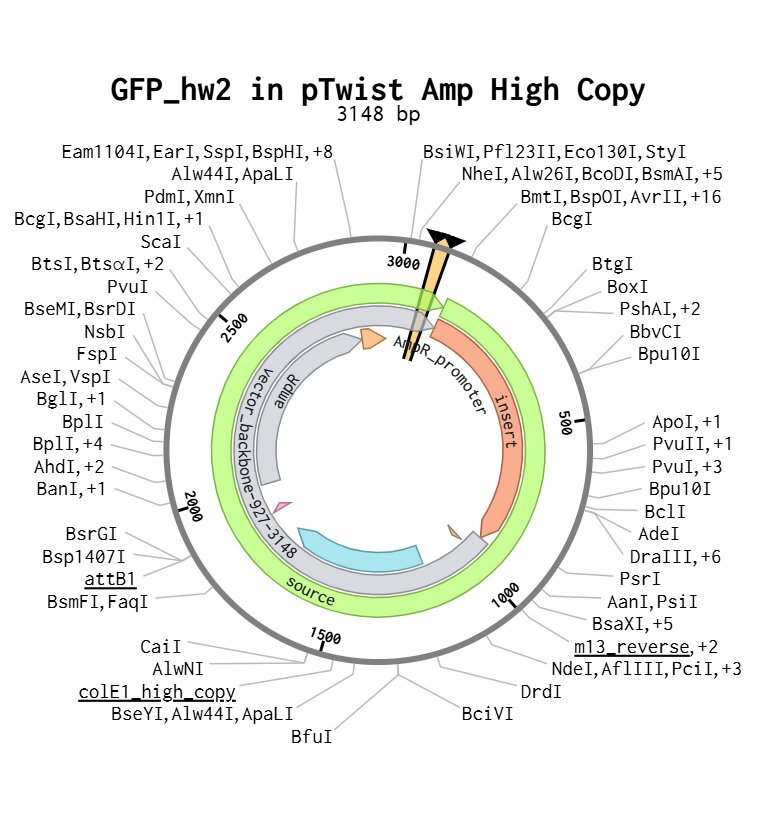

Part 5: DNA Read/Write/Edit
5.1 DNA Read
If I could choose DNA to sequence, I would choose synthetic DNA used for digital data storage. DNA data storage is interesting because it treats DNA as away of encoding information, like a hard drive, but at a molecular scale. Sequencing the DNA would allow us to read back the stored data and measure how accurately the system preserves information over time, this seems like an idea straight out of science fiction though it is possible now. To sequence this DNA, I would use Illumina sequencing, a second-generation sequencing technology. It works well for short, synthetic DNA fragments and provides high accuracy at relatively low cost. The input would be the synthetic DNA fragments that encode the information. These fragments wouldneed to be prepared by adding adapter sequences to their ends, amplifying them by PCR, and loading them onto a flow cell. During sequencing, fluorescently labeled nucleotides are incorporated one base at a time, and a camera detects the color signal to determine which base was added. This process converts fluorescence into a digital readout of A, T, C, and G. The output is a large dataset of DNA reads that can be reconstructed into the original digital file.
5.2 DNA Write
If I could synthesize DNA, I would create my turtle-themed GFP expression construct but with multiple colors of fluorescent proteins. I would synthesize codon-optimized Fluorescent Protein genes under the control of a bacterial promoter so that it could be expressed in E. coli. The goal would be to design a system that produces a turtle-shell-like hexagonal fluorescence pattern. This connects synthetic biology with spatial design and pattern formation maybe I could even get a system for animating it and getting the turtle to move. The core sequence would be the coding region, inserted into a plasmid backbone.
To synthesize this DNA, I would use chemical DNA synthesis and fragment assembly. Short DNA oligos are chemically synthesized, assembled into the full gene using overlapping regions, cloned into a plasmid, and sequence-verified. This method is good for constructs around 1 kb, such as GFP. Limitations include cost increasing with length, possible synthesis errors, and longer turnaround times for larger constructs.
5.3 DNA Edit
An interesting DNA edit project would be engineering biological motors or force-generating systems that could act as microscopic actuators. It would be interesting to use the process that proteins such as Kinesin-1 use to convert chemical energy from ATP into mechanical motion along microtubules. By editing the genes that encode these motor proteins, we could potentially tune their speed, force output, or binding properties, creating nanoscale linear motors with maybe even the potential to scale up. Editing their DNA could allow us to design programmable biological actuators for soft robotics, microfluidics, or responsive materials.
To perform this editing, I would use CRISPR-Cas9. I’d design a guide RNA that targets the motor protein gene. Cas9, directed by the guide RNA, cuts the DNA at a specific location. If we want to introduce a modification, such as a mutation that alters motor speed or adds a binding domain, we would also provide a donor DNA template for repair. The inputs include the guide RNA sequence, Cas9, the donor template if inserting changes, and host cells. After editing, the modified cells would express the altered motor protein, and its mechanical properties could be measured experimentally. Limitations include variable editing efficiency and the possibility of off-target edits, but the approach might allow for precise modification of biological force-generating systems.
Sources
- https://www.uniprot.org/uniprotkb/P42212/entry#sequences
- https://www.novoprolabs.com/tools/codon-optimization
- https://www.bioinformatics.org/sms2/rev_trans.html
AI Prompts
- “How do you transcribe DNA and translate a protein with a cell-dependent method?”
- “How do you transcribe DNA and translate a protein with a cell-independent method?”