Week 6 HW: Genetic Circuits Part I

HW6
PCR and DNA Assembly
- What are some components in the Phusion High-Fidelity PCR Master Mix and what is their purpose?
The Phusion High-Fidelity PCR Master Mix contains a high-fidelity DNA polymerase for accurate DNA replication, dNTPs as the building blocks for new DNA strands, a buffer to maintain optimal reaction conditions, and magnesium ions which act as a cofactor for the polymerase. Together, these components enable efficient and precise DNA amplification.
- What are some factors that determine primer annealing temperature during PCR?
Primer annealing temperature is primarily determined by the melting temperature of the primers, which depends on their length and GC content. Higher GC content increases the temp due to stronger hydrogen bonding.
- There are two methods from this class that create linear fragments of DNA: PCR, and restriction enzyme digests. Compare and contrast these two methods, both in terms of protocol as well as when one may be preferable to use over the other.
PCR generates linear DNA fragments by amplifying a target sequence using primers and a polymerase through thermal cycling, allowing for flexible design and modification of DNA ends. In contrast, restriction enzyme digestion cuts DNA at specific recognition sites, producing predictable fragments but with less flexibility. PCR is better when creating new sequences or adding overlaps, while restriction digests are better when cutting existing DNA at known sites for cloning.
- How can you ensure that the DNA sequences that you have digested and PCR-ed will be appropriate for Gibson cloning?
To ensure compatibility with Gibson cloning, the DNA fragments must have overlapping homologous regions that allow them to anneal during assembly. They should be designed to keep the correct orientation and reading frame, and checked using sequence analysis tools. Gel electrophoresis can be used to confirm fragment size before assembly.
- How does the plasmid DNA enter the E. coli cells during transformation?
Plasmid DNA enters cells through heat shock or electroporation. Heat shock temporarily increases membrane permeability because of the rapid temperature change, and electroporation uses an electric field to create pores in the cell membrane so DNA can pass into the cell.
- Describe another assembly method in detail
Golden Gate Assembly uses Type IIS restriction enzymes that cut outside of their recognition sites to generate custom overhangs. These overhangs allow multiple DNA fragments to be assembled in a specific order in a single reaction. DNA ligase then joins the fragments together, resulting in a seamless construct without extra bases.
- Explain the other method in 5–7 sentences plus diagrams
Golden Gate Assembly relies on Type IIS restriction enzymes, which cut DNA outside of their recognition sequences to produce specific overhangs. These overhangs are designed so that different DNA fragments can anneal in a predetermined order. During the reaction, restriction enzymes cut the DNA while ligase simultaneously joins compatible fragments, driving the system toward the correct final product. Because the recognition sites are removed during assembly, the resulting DNA sequence is seamless. This method is best for assembling multiple fragments in a single reaction and is popular for synthetic biology, but it requires careful design of overhang sequences to avoid incorrect assembly. It is a fast and scalable method for constructing complex DNA designs.
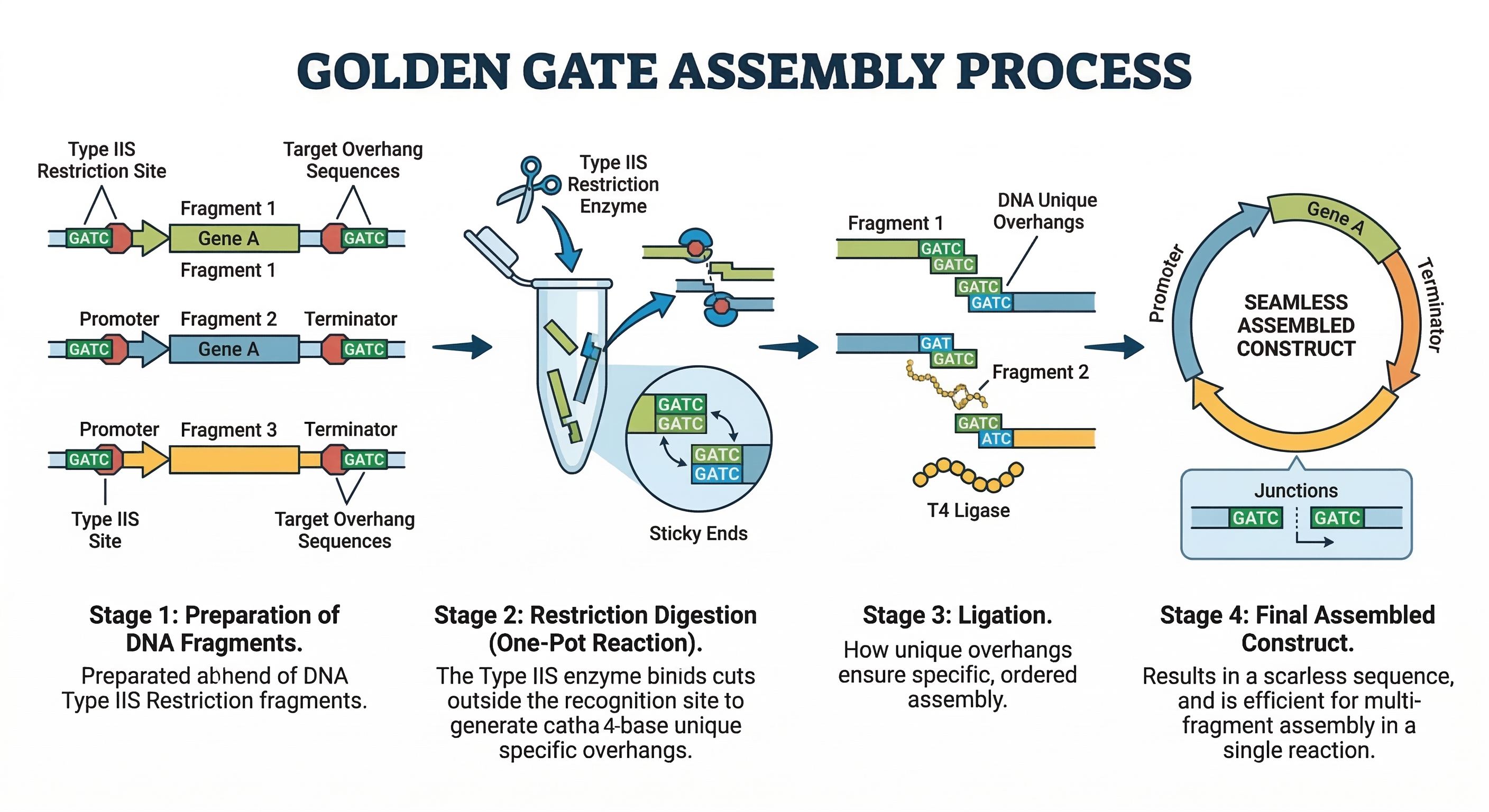
- Model this assembly method with Benchling or Asimov Kernel
For this part of the assignment, I used Benchling to create a Golden Gate Assembly of the pUC19 backbone with the GFP insert.
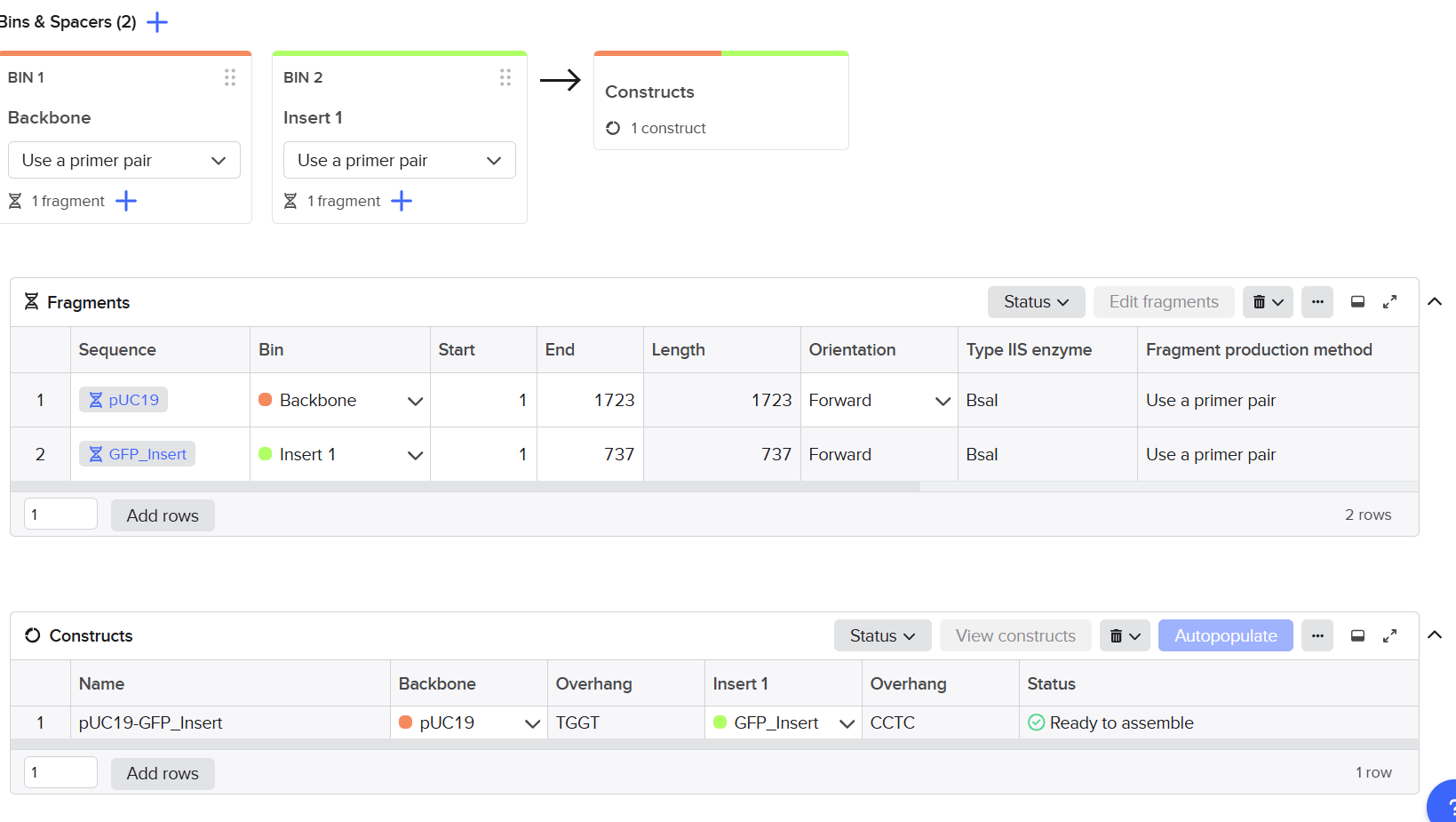
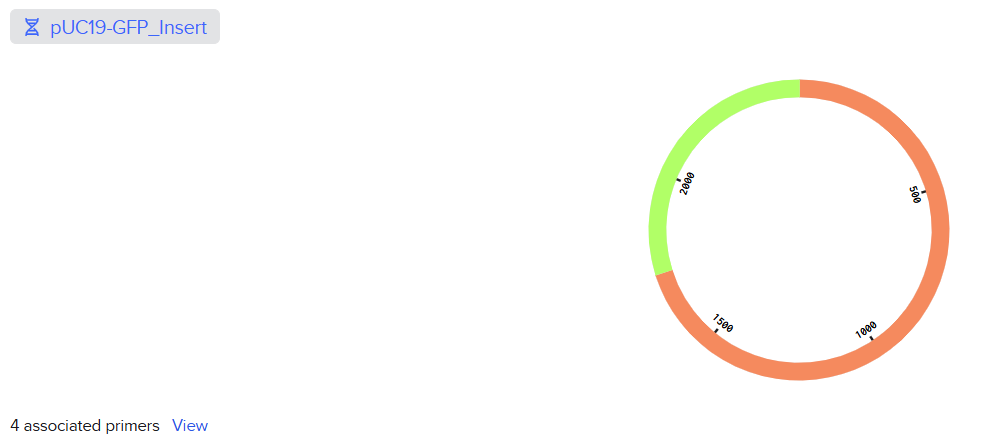
Asimov Kernel
For the first part of the kernel homework I looked into the repressilator and multiplexer circuits. The repressilator is a synthetic oscillatory circuit made of three genes that repress each other in a loop, causing time-delayed negative feedback that creates in periodic oscillations of protein expression. The multiplexer is a logic-based circuit that uses L-arabinose, aTc, and IPTG as inputs and produces a high GFP output only for specific combinations of these inputs. It was interesting to see their simulations and compare them to electronic circuits.
I also had AI help create a cheat sheet: Cheat Sheet pBad – activated by arabinose pTet – repressed by TetR, activated by aTc pLacI / pTac – repressed by LacI, activated by IPTG RBS – controls translation strength TetR – represses pTet LacI – represses Lac promoters GFP – fluorescent output L3S2P24 Terminator – stops transcription aTc – inhibits TetR IPTG – inhibits LacI Arabinose – activates pBad structure - Promoter → RBS → CDS → Terminator → Promoter → RBS → CDS → Terminator
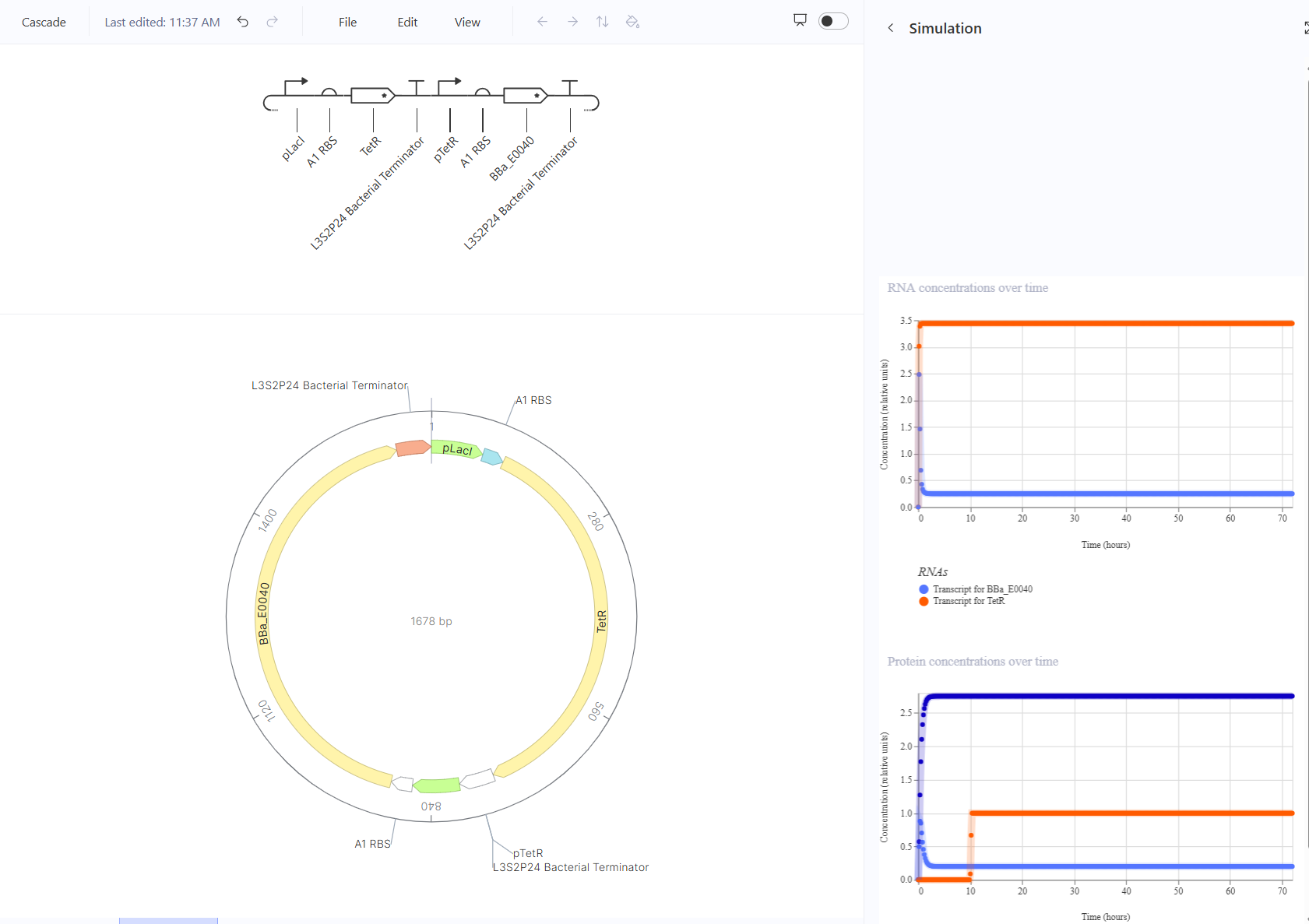
For the second part, I recreated the repressilator and simulated it.
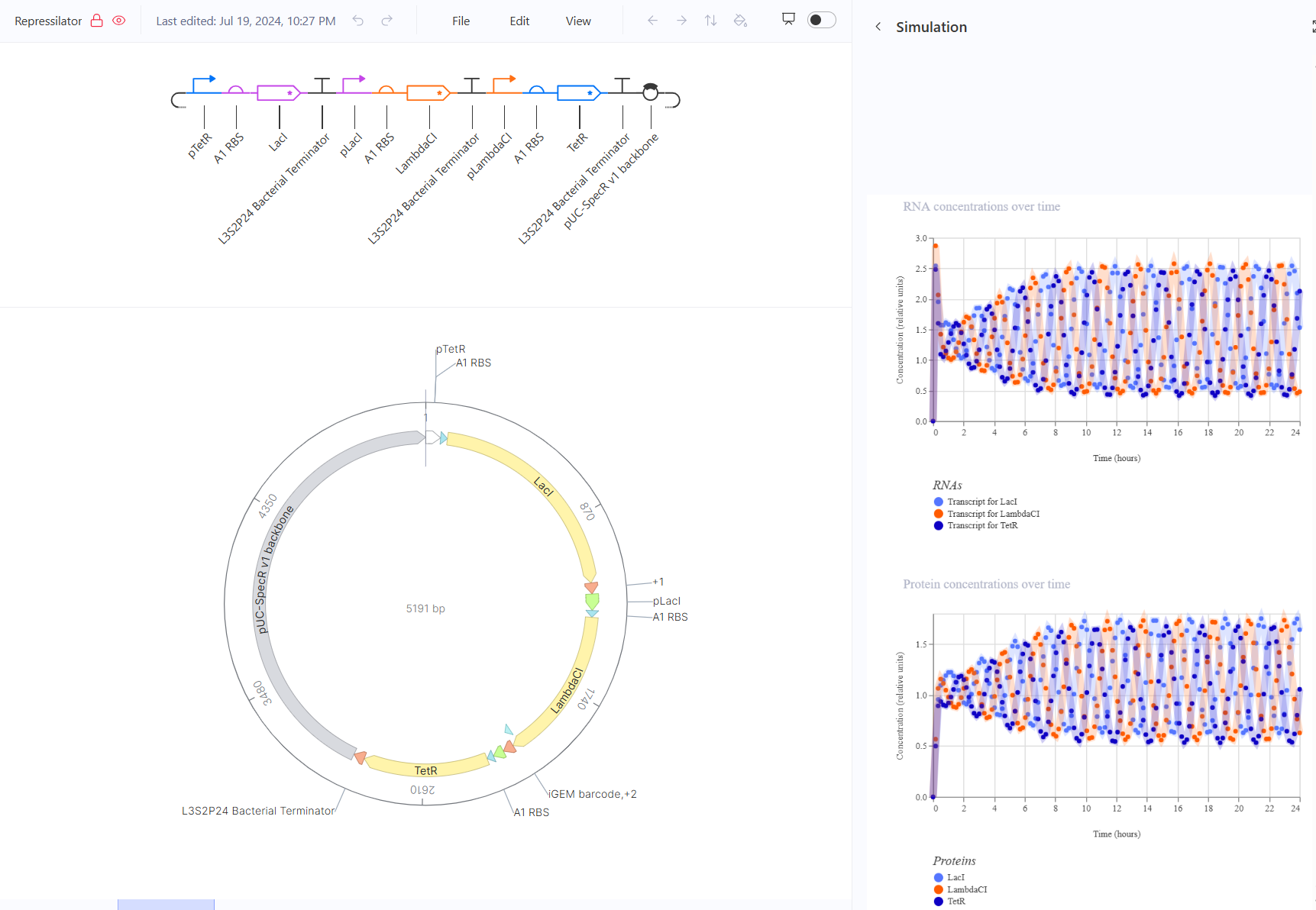
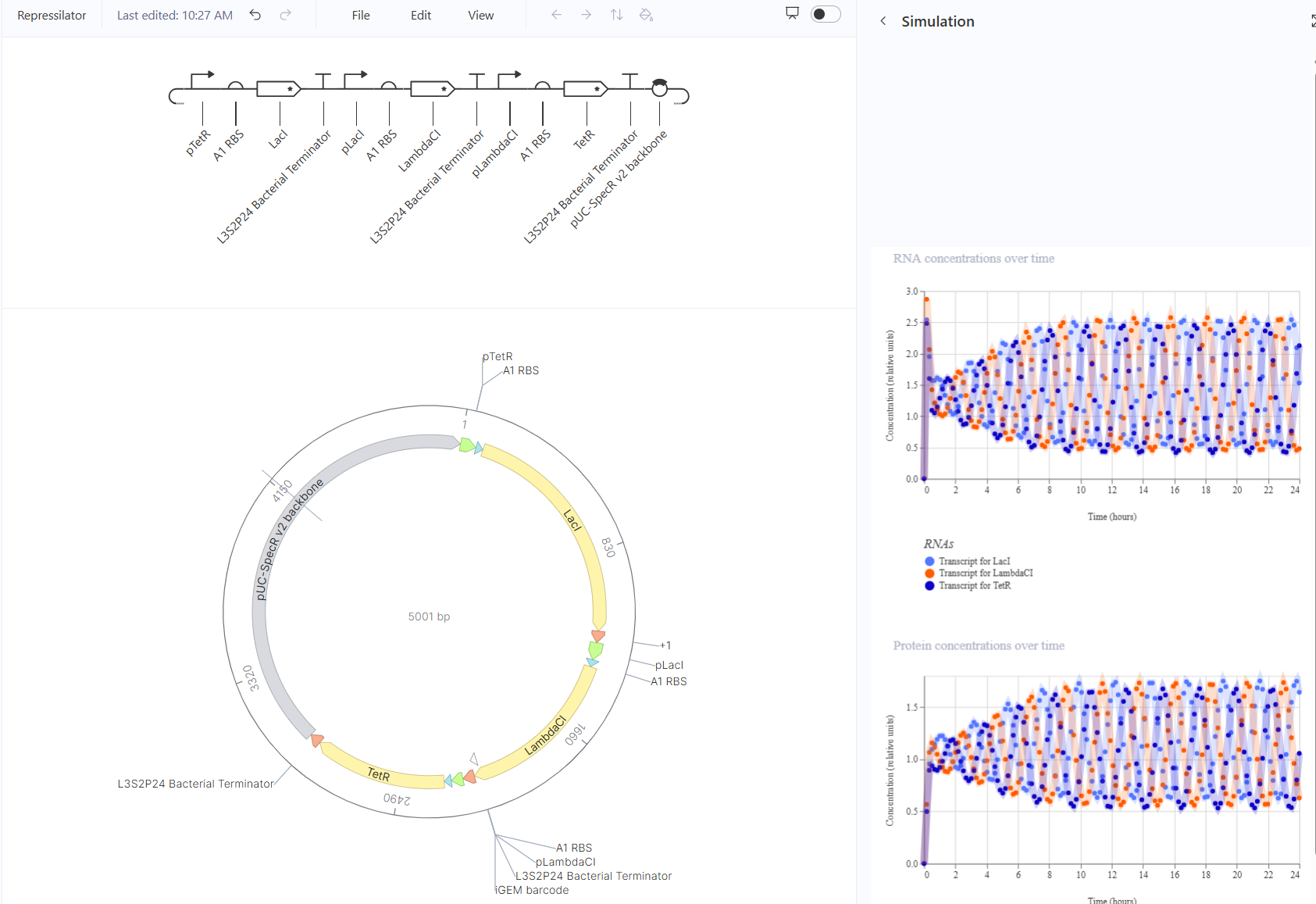
For the third part, I tried to design 3 circuits, a toggle switch, a pulse, and a tiered cascade.
The toggle switch should turn off when aTc is added and on with IPTG. This should happen by turning the transcript for LacI on and the transcript for TetR off, causing GFP production to stop. This one worked in simulation, though I also expected it to respond to IPTG.
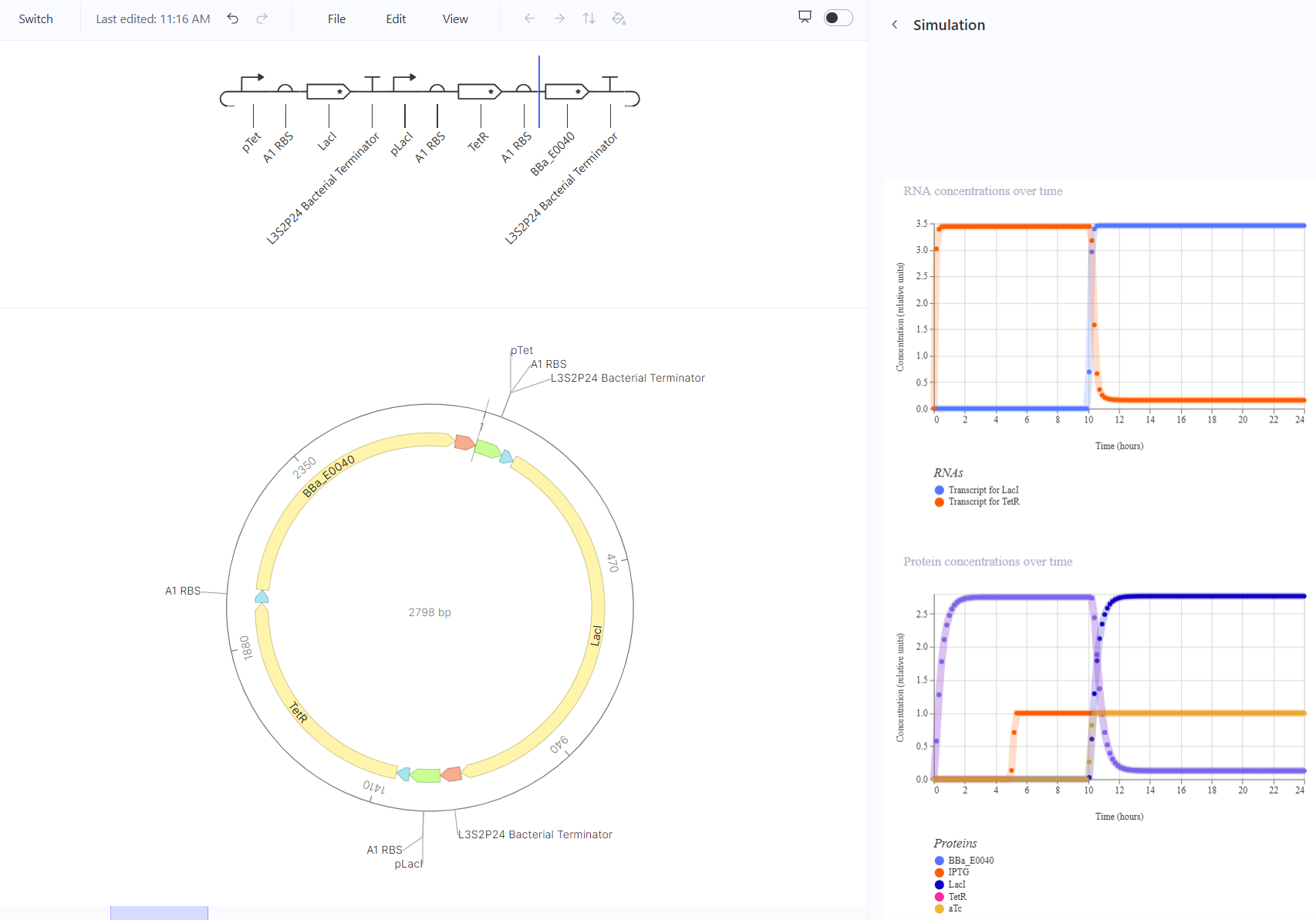
The pulse should show a sharp increase then decrease when arabinose is added. This should happen by turning the transcript for TetR on and the transcript for GFP will go from zero to high production, then decrease back to zero. This one kind of worked in simulation, but I expected GFP to start and end at zero and instead I still got a pulse but it was only a relatively low to high pulse which might affect the visibility of glowing but won’t cause a flicker.
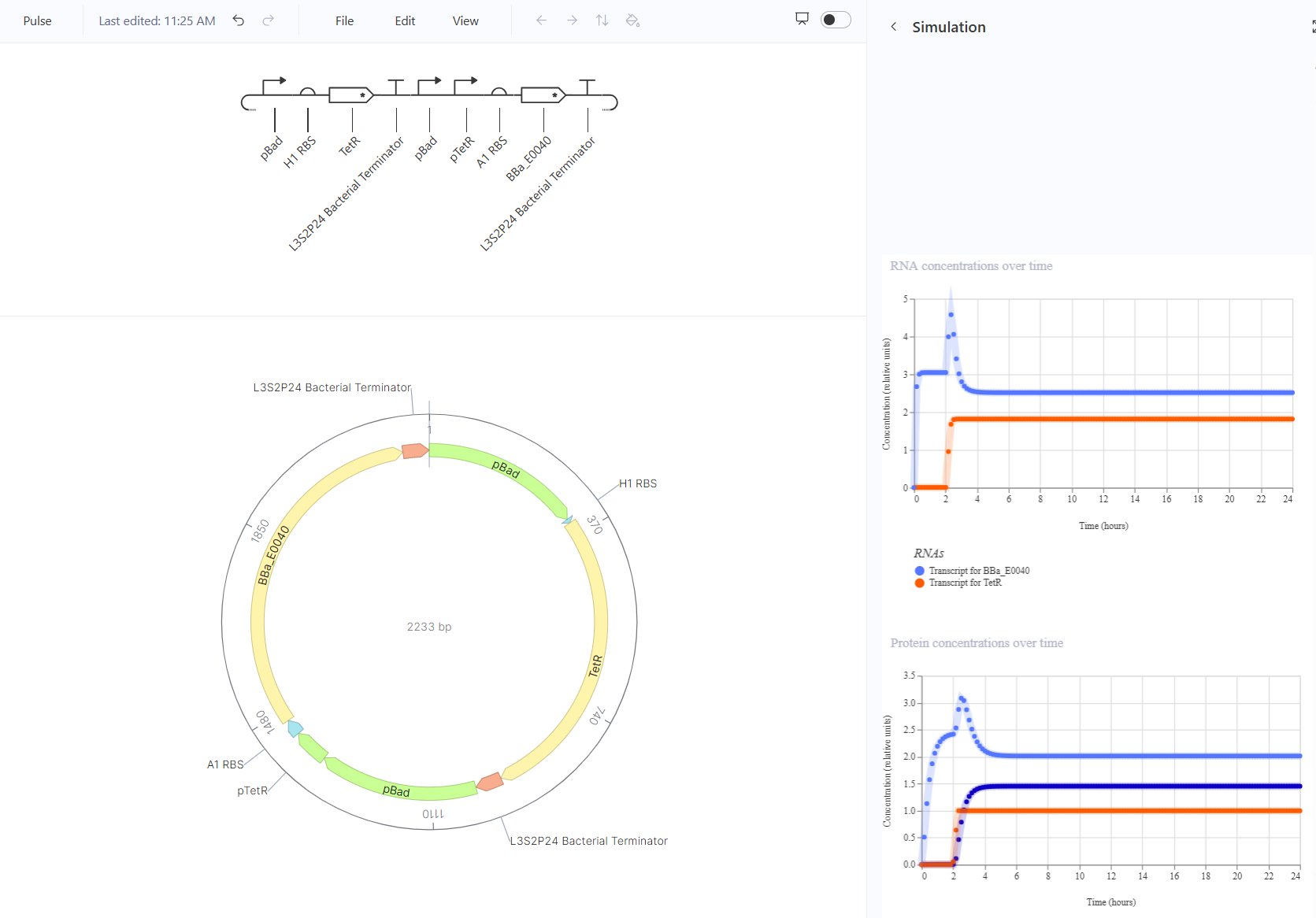
The cascade should look like stairs, starting from one rate of GFP production then stepping down to a lower rate of GFP production when arabinose is detected. This should happen by turning the transcript for TetR on and the transcript for GFP will go from low to high production, then decrease back to zero. This one did not work at all, there was no response to the arabinose. I believe that there was something wrong with the sensing part of the circuit, but I couldn’t troubleshoot it. Consulting ChatGPT recommended some changes to the TetR part to ensure detection of arabinose, but I wasn’t sure how to do that.