Week 9 HW: Cell Free Systems
HW9
General homework questions
- Explain the main advantages of cell-free protein synthesis over traditional in vivo methods, specifically in terms of flexibility and control over experimental variables. Name at least two cases where cell-free expression is more beneficial than cell production.
Cell-free gives more flexibility and control because it operates in a system where you can directly control concentrations of DNA, ions, and cofactors. This allows rapid prototyping and expression of toxic or non-natural proteins without killing your host cell. It is especially beneficial for producing toxic proteins and for quickly testing genetic circuits.
- Describe the main components of a cell-free expression system and explain the role of each component.
A cell-free expression system contains a cell extract with transcription and translation machinery, a DNA or mRNA template encoding the protein, and an energy system to fuel synthesis. Also amino acids, salts, and cofactors that support proper folding and reaction stability.
- Why is energy provision regeneration critical in cell-free systems? Describe a method you could use to ensure continuous ATP supply in your cell-free experiment.
Energy regeneration is critical because protein synthesis quickly consumes ATP and GTP. Without replenishment the reaction stops. Continuous ATP supply can be maintained by using systems like phosphoenolpyruvate with pyruvate kinase to regenerate ATP from ADP.
- Compare prokaryotic versus eukaryotic cell-free expression systems. Choose a protein to produce in each system and explain why.
Prokaryotic cell-free systems are fast and high-yield while eukaryotic systems support proper folding and modifications but are slower and more expensive. A simple protein like GFP is best produced in a prokaryotic system, but a complex protein like an antibody requires a eukaryotic system.
- How would you design a cell-free experiment to optimize the expression of a membrane protein? Discuss the challenges and how you would address them in your setup.
A cell-free system can include liposomes or nanodiscs to mimic a membrane environment and prevent aggregation. Challenges include hydrophobic misfolding and instability, which can be addressed with careful tuning of reaction conditions which improves proper insertion and folding.
- Imagine you observe a low yield of your target protein in a cell-free system. Describe three possible reasons for this and suggest a troubleshooting strategy for each.
Low protein yield might be caused by poor transcription, inefficient translation, or energy depletion. These can be addressed by using stronger promoters or more DNA, optimizing ribosome binding sites or codons, and improving ATP regeneration systems.
Homework question from Kate Adamala
Design an example of a useful synthetic minimal cell as follows:
- Pick a function and describe it.
What would your synthetic cell do? What is the input and what is the output?
For my final project, I want to make a lactate-detecting cell-free system. It would detect lactate concentration and produce a measurable signal. It acts as a biochemical sensor that converts a lactate input into a fluorescent output and is useful for monitoring physiological or environmental lactate levels
Could this function be realized by cell-free Tx/Tl alone, without encapsulation?
Yes, that’s how I plan to do it for my final project, but encapsulation would improve control over sensing and environmental interaction and might also let me make the reaction resettable. Without a membrane, diffusion and signal localization are harder to regulate.
Could this function be realized by a genetically modified natural cell?
A genetically modified natural cell could also perform lactate sensing using my genetic circuit. However, natural cells introduce complexity, slower response times, and biosafety concerns. My circuit could also produce a large amount of h2o2 which is harmful to natural cells but maybe not to synthetic ones.
- Describe the desired outcome of your synthetic cell operation.
The desired outcome is a system that produces a reliable, quantifiable signal proportional to lactate concentration. The response should be sensitive, fast, and specific to lactate. This allows accurate monitoring in real time.
- Design all components that would need to be part of your synthetic cell.
- What would the membrane be made of?
The membrane would be composed of a lipid bilayer combined with cholesterol for stability. This provides a semi-permeable barrier similar to natural cells. It also helps maintain internal reaction conditions.
- What would you encapsulate inside? Enzymes, small molecules.
Inside the cell, would be transcription/translation machinery, DNA encoding the sensor and reporter, amino acids, ATP regeneration components, and cofactors. My cell in particular would also include lactate oxidase and DFBH dye.
- Which organism your Tx/Tl system will come from? Is bacterial OK, or do you need a mammalian system for some reason? (hint: for example, if you want to use small molecule modulated promotors, like Tet-ON, you need mammalian)
I plan to use E. coli, which is sufficient for expressing simple reporter proteins and oxyR. A mammalian system is not necessary unless complex regulatory elements are required, my second goal was to track sodium levels which a mammalian system might be best for
- How will your synthetic cell communicate with the environment? (hint: are substrates permeable? or do you need to express the membrane channel?)
Lactate would diffuse across the membrane or through membrane channels. If lactate permeability is low, the transporter protein lactate permease could be included.
- Experimental details
List all lipids and genes. Lipids: POPC and cholesterol. Genes: lactate-responsive regulator, a promoter controlled by OxyR, dye as a reporter, possibly lactate permease for transport, and lactate oxidase
How will you measure the function of your system?
By monitoring fluorescence intensity over time using a plate reader or microscope. Increased fluorescence indicates a higher lactate concentration.
Homework question from Peter Nguyen
- Write a one-sentence summary pitch sentence describing your concept.
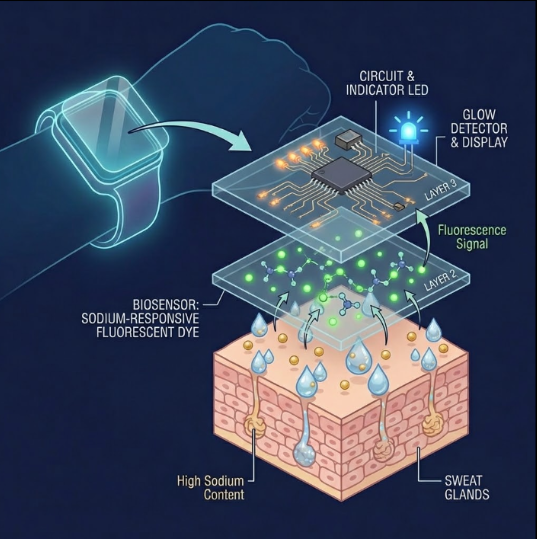
I want to make a freeze-dried cell-free system embedded in workout clothes that detects lactate buildup in sweat a change color to indicate fatigue and performance.
- How will the idea work, in more detail? Write 3-4 sentences or more.
Cell-free reactions containing a lactate-responsive genetic circuit are freeze-dried into removable patches that are attached to various locations on the fabric. When the wearer sweats, moisture rehydrates the system and lactate diffuses into the reaction, activating expression of a fluorescent reporter dye. The intensity of the signal correlates with lactate concentration, providing real-time feedback on muscle fatigue.
- What societal challenge or market need will this address?
Not sure if anyone needs this, I just think it would be cool. Originally, I wanted to do something that informs the wearer of dehydration.
- How do you envision addressing the limitation of cell-free reactions (e.g., activation with water, stability, one-time use)?
I hope that sweat provides enough water to activate the freeze-dried systems, and using modular replaceable patches would allow for multiple uses and control stability.
Homework question from Ally Huang
- Provide background information that describes the space biology question or challenge you propose to address. Explain why this topic is significant for humanity, relevant for space exploration, and scientifically interesting.
Microgravity leads to rapid muscle atrophy due to reduced mechanical loading, posing a major risk for long-duration space missions. Current countermeasures, like exercise, are time-consuming and not fully effective. Understanding and monitoring molecular pathways that regulate muscle growth and degradation is essential for developing better interventions. This is significant for maintaining astronaut health, enabling deep space exploration, and advancing treatments for muscle-wasting diseases on Earth. Studying these pathways in space also provides insight into fundamental mechanisms of mechanobiology.
- Name the molecular or genetic target that you propose to study. Examples of molecular targets include individual genes and proteins, DNA and RNA sequences, or broader approaches.
Myostatin (MSTN) and muscle growth regulators, including IGF-1 pathway gene expression markers.
- Describe how your molecular or genetic target relates to the space biology question or challenge your proposal addresses.
Myostatin negatively regulates muscle growth, while IGF-1 promotes muscle synthesis and repair. In microgravity, increased myostatin activity and reduced anabolic signaling contribute to muscle loss. By monitoring these molecular targets, we can track the balance between muscle degradation and growth. This helps identify when and how muscle atrophy occurs in space. Understanding these changes enables development of targeted countermeasures to maintain muscle mass.
- Clearly state your hypothesis or research goal and explain the reasoning behind it.
Microgravity conditions upregulate myostatin expression and downregulate IGF-1 signaling, leading to accelerated muscle atrophy. A cell-free system can be used to detect changes in expression of these markers from biological samples in space. The goal is to develop a lightweight, rapid diagnostic platform that monitors muscle health in real time using the stability and simplicity of freeze-dried cell-free systems to make it suitable for spaceflight. If successful, it could guide personalized countermeasures such as exercise timing or therapeutic interventions. This would improve astronaut health and mission performance during long-duration space travel.
- Outline your experimental plan - identify the sample(s) you will test in your experiment, including any necessary controls, the type of data or measurements that will be collected, etc.
Samples containing RNA or DNA proxies for MSTN and IGF-1 expression will be analyzed using BioBits® cell-free reactions designed to produce fluorescence in response to target sequences. miniPCR® will amplify target sequences if needed, and fluorescence will be measured using the P51 viewer. Controls will include samples without target sequences and with known concentrations. Data will be collected as fluorescence intensity over time and compared across simulated microgravity conditions. This will quantify changes in muscle-related gene expression.
First DNA Twist Order
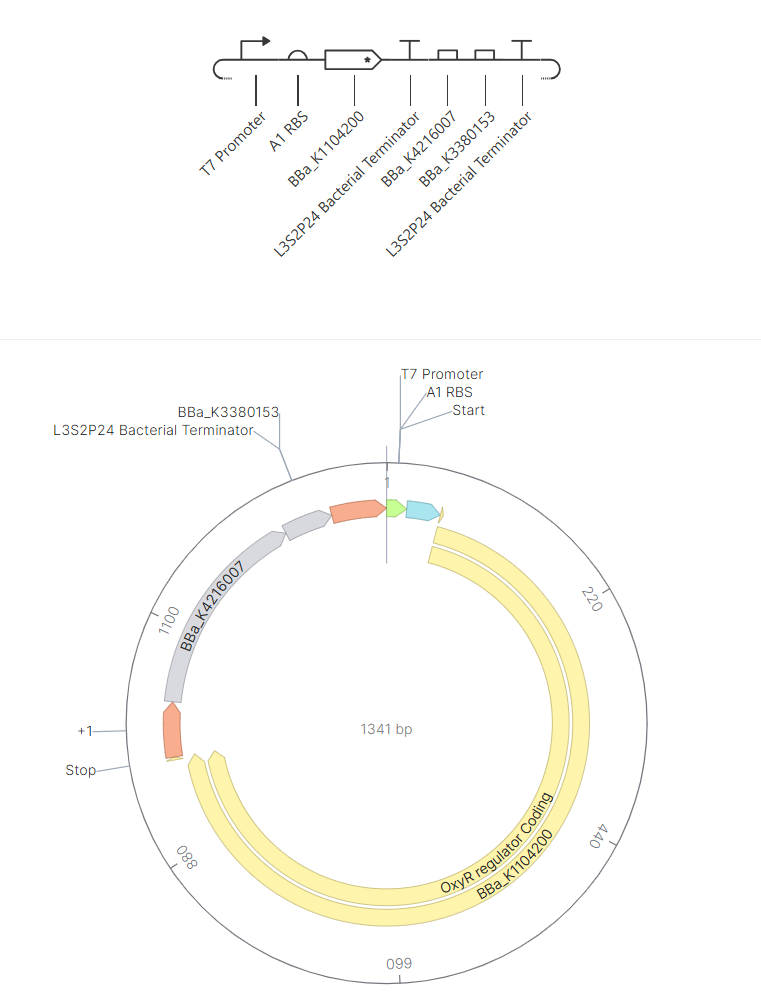
For my final project, I want to make a “hydration” checking wearable device. Originally, I wanted to sense increased sodium levels insweat but that proved to be difficult so instead I’m approximating increased hydration risk by just detecting lactate. I want this to be a cell free system to make it more compatible with a safe wearable device.
Lactate (input signal from sweat) -> Lactate Oxidase (breaks lactate down into) -> Pyruvate (byproduct not used) + H2O2 (which then activates OxyR by oxidizing it) -> OxyR (has two states starts reduced then is oxidized to become active and bind to DNA to activate transcription of PoxyS) -> PoxyS Promoter (controlled by OxyR, switches on reporter gene) -> RNA Aptamer (produced when PoxyS promoter is enabled binds to dye) -> DFHBI Dye (non-fluorescent when bound to RNA Aptamer)
All together it follows this chain: Lactate provides the biological input (increased sweat means increased need for hydration) Lactate Oxidase and H2O2 convert it into a detectable chemical signal OxyR and PoxyS act as switch RNA aptamer and dye generate a fast fluorescent output (that way the glow happens quicker than with protein translation and safer for a wearable because there are no cells)