Albert Manrique — HTGAA Spring 2026
About me
Hello! I am an Electrical Engineer currently based out of Miami,FL looking to learn more about synbio!
Hello! I am an Electrical Engineer currently based out of Miami,FL looking to learn more about synbio!
Week 1 HW: Principles and Practices
Glowing Leather as a pilot for decentralized biomanufacturing of goods Bacterially produced leather is currently in the stage of being scaled up in centralized biomanufacturing plants around the world utilizing waste feedstocks from agricultural sources. Smaller scale experiments utilize kombucha SCOBYs to produce bacterial leather however they face variability in leather quality due to different growing conditions, feedstock, and SCOBY relationships in regards to each individual member’s reaction to the feedstock/growing conditions. The genetic drift that the SCOBY undergoes specifically the K. xylinus also makes it unreliable for consistent bacterial leather production. I am interested in exploring the development of an open source decentralized protocol to make growing bioleather easier, more climate resilient than growing cows, and more equitable than current centralized operations.
Week 2 HW: DNA Read, Write, & Edit
Week 2 Homework Part 1: Benchling & In-silico Gel Art MIT/Harvard students Required Committed Listeners Required See this week’s lab protocol “Gel Art: Restriction Digests and Gel Electrophoresis” for details. Overview: Make a free account at benchling.com Import the Lambda DNA Simulate Restriction Enzyme Digestion with the following enzymes: EcoRI HindIII BamHI KpnI EcoRV SacI SalI Create a pattern/image in the style of Paul Vanouse’s Latent Figure Protocol artworks You might find Ronan’s website a helpful tool for quickly iterating on designs! Response I was able to import all the restriction enzymes into benchling,
Homework Assignment: Python Script for Opentrons Artwork — DUE BY YOUR LAB TIME! Assignees for this section MIT/Harvard students Required Committed Listeners Required Your task this week is to Create a Python file to run on an Opentrons liquid handling robot.
I am interested in exploring the development of an open source decentralized protocol to make growing bioleather easier, more climate resilient than growing cows, and more equitable than current centralized operations.
Next, score (from 1-3 with, 1 as the best, or n/a) each of your governance actions against your rubric of policy goals. The following is one framework but feel free to make your own:
| Governance Actions / Policy Goals | PG1.1: Minimize regulatory & IP hurdles | PG1.2: Enable frugal deployment by end users | PG2.1: Implement unobtrusive & robust licensing | PG2.2: Utilize GRAS/BSL-1 organisms to minimize user error harm | PG3.1: Utilize existing waste streams | PG3.2: Avoid toxic processing byproducts | PG3.3: Carbon neutral manufacturing (blue-sky) |
|---|---|---|---|---|---|---|---|
| Govt Regulatory Agencies | |||||||
| Establish GMO importation & use standards | 2 | n/a | 2 | 2 | n/a | n/a | n/a |
| Monitor registered GMO users | 3 | n/a | 2 | n/a | n/a | n/a | n/a |
| Advocate for simplified BSL-1 pathways | 1 | 1 | 2 (i think?) | 1 | n/a | n/a | n/a |
| Distributed Ops Network + Strain Banks | |||||||
| Distribute certified sachets | 1 | 2 | 1 | 1 | n/a | n/a | n/a |
| Fluorescent reporter quality gate | 3 | n/a | 2 | 2 | n/a | n/a | n/a |
| Propagate regional strain banks | 1 | 1 | 1 | 1 | 2 | 2 | 1 |
| Global Academic Researchers | |||||||
| Develop low-resource-compatible strains | 1 | 1 | 1 | 1 | 2 | 2 | 1 |
| Require open-source publishing & licensing | 1 | 1 | 1 | 1 | 2 | 2 | 1 |
| Global test lab network | 2 | 1 | 2 | 2 | 1 | 3 | 1 |
| Cross-lab reproducibility protocols | 1 | 1 | 2 | 1 | 1 | 3 | 1 |
Our action plan would first prioritize organizing a global collective of interested scientists and engineers (perhaps at HTGAA!) to develop the deployment protocol that takes into account regulatory actions, equitable use, and licensing considerations. This would allow us to organize the foundational work needed for deployment. Our main deliverable would be a published and verified protocol that would allow us to distribute our findings for use in an open source license.
We would then prioritize interfacing with regional regulatory bodies in order to further refine protocols for use in target regions using a BSL-1 organism.
In order to find revenue we would seek to target luxury goods and create a beachhead market and further fund distributed deployment of manufacturing. More research is needed but our main tradeoff would be international regulatory work would have to wait until initial revenue or funding is secured.
AI usage Summary can be found here
Homework Questions from Professor Jacobson: [Lecture 2 slides]
Nature’s machinery for copying DNA is called polymerase. What is the error rate of polymerase? How does this compare to the length of the human genome. How does biology deal with that discrepancy?
The throughput error rate is 1:“10^6 and 10mS per base addition length of the human genome is 6.2 bil base pairs.
Biology deals with this by having the polymerase be able to error correct in addition to having other proteins correct remaining errors. On slide 14 it describes the MutS repair system.
How many different ways are there to code (DNA nucleotide code) for an average human protein? In practice what are some of the reasons that all of these different codes don’t work to code for the protein of interest?
Homework Questions from Dr. LeProust: [Lecture 2 slides]
What’s the most commonly used method for oligo synthesis currently?
Why is it difficult to make oligos longer than 200nt via direct synthesis?
Why can’t you make a 2000bp gene via direct oligo synthesis?
Homework Question from George Church: [Lecture 2 slides]
Choose ONE of the following three questions to answer; and please cite AI prompts or paper citations used, if any.
[Using Google & Prof. Church’s slide #4] What are the 10 essential amino acids in all animals and how does this affect your view of the “Lysine Contingency”?
Phenylalanine, Valine, Threonine, Tryptophan, Isoleucine, Methionine, Histidine, Arginine, Leucine, Lysine
Turns out in the Jurassic Park franchise they’re just eating Lysine from sources in the environment.
Would be more prudent to use an amino acid that’s not available in nature and controllable.
| MIT/Harvard students | Required |
| Committed Listeners | Required |
See this week’s lab protocol “Gel Art: Restriction Digests and Gel Electrophoresis” for details. Overview:
I was able to import all the restriction enzymes into benchling,
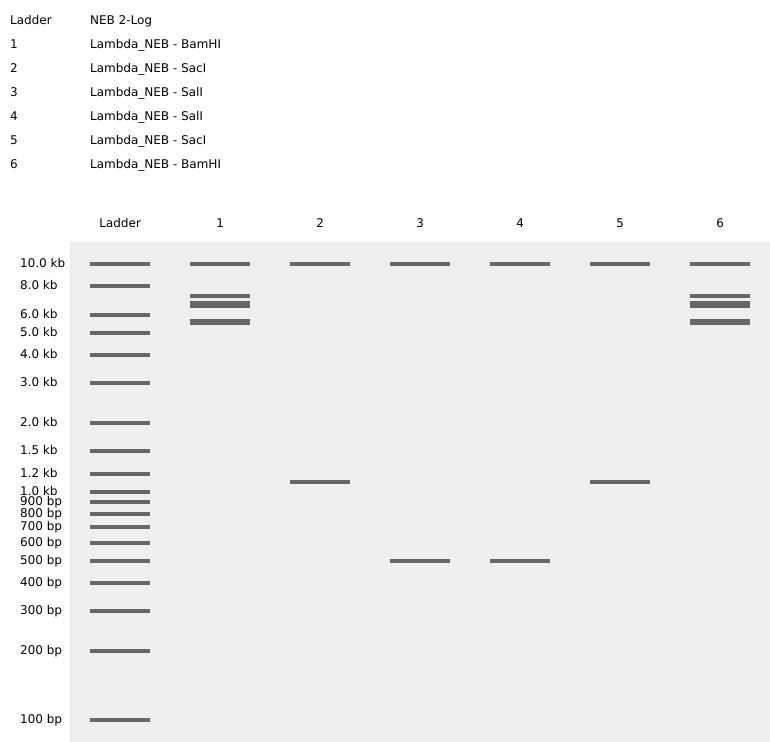
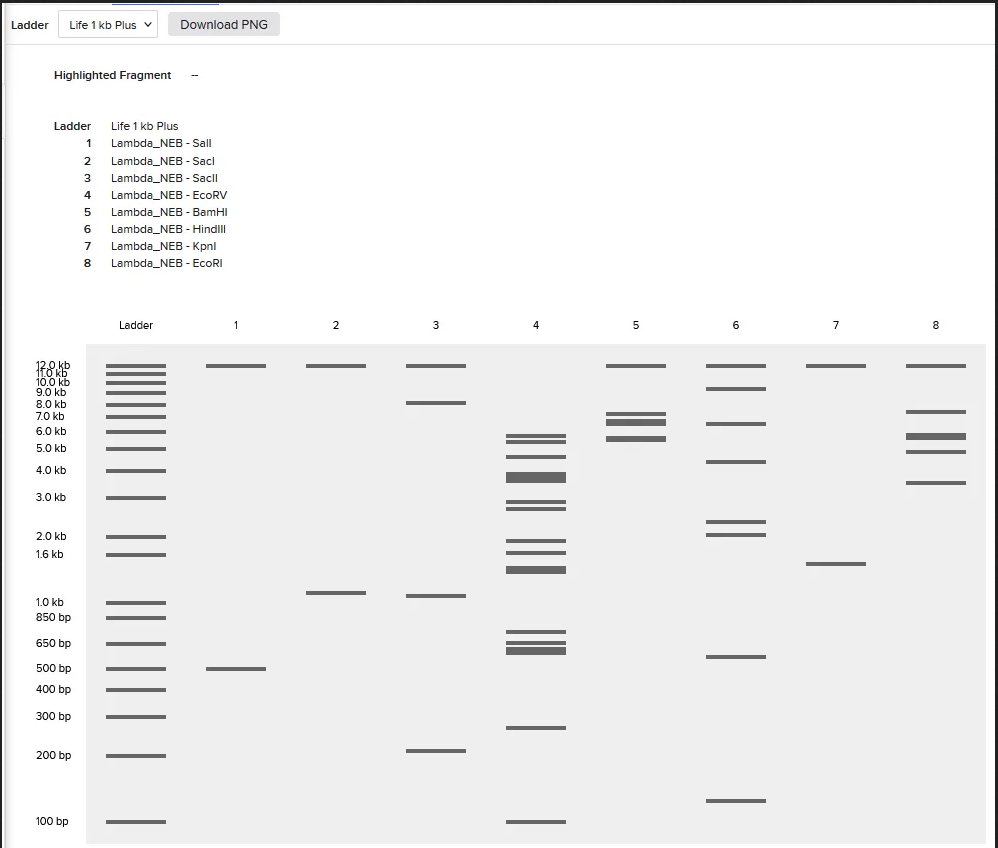

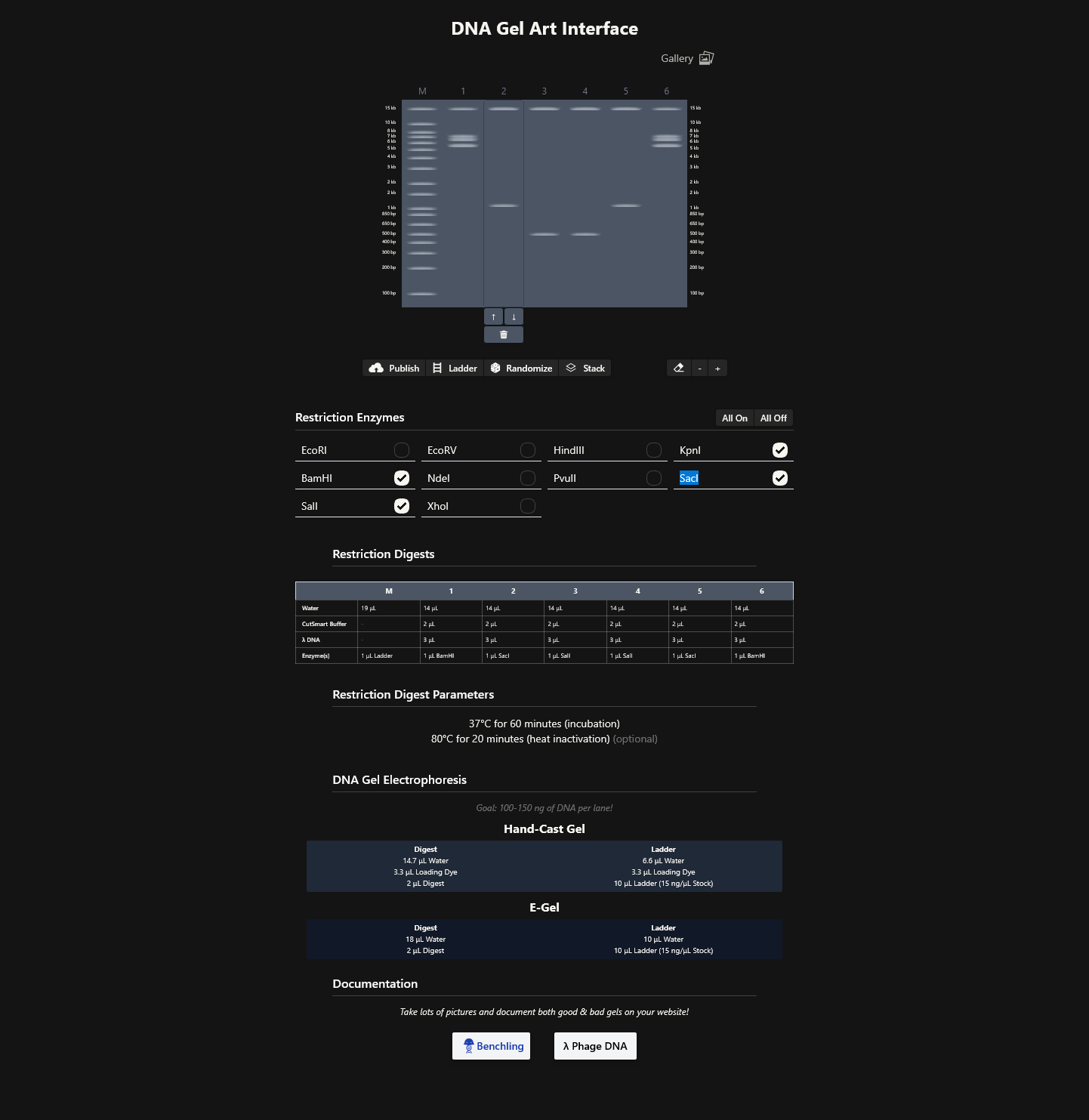



In recitation, we discussed that you will pick a protein for your homework that you find interesting. Which protein have you chosen and why? Using one of the tools described in recitation (NCBI, UniProt, google), obtain the protein sequence for the protein you chose.
Example from group homework:
>sp|P03609|LYS_BPMS2 Lysis protein OS=Escherichia phage MS2 OX=12022 PE=2 SV=1 METRFPQQSQQTPASTNRRRPFKHEDYPCRRQQRSSTLYVLIFLAIFLSKFTNQLLLSLL EAVIRTVTTLQQLLT
Response
I used UniProt to source Cellulose synthase from Komagataeibacter xylinus as I wanted to investigate growing bacterial cellulose as leather substitute for my final project.
Source: UniProt P19449 — BCSA1_KOMXY
The Central Dogma discussed in class and recitation describes the process in which DNA sequence becomes transcribed and translated into protein. The Central Dogma gives us the framework to work backwards from a given protein sequence and infer the DNA sequence that the protein is derived from. Using one of the tools discussed in class, NCBI or online tools (google “reverse translation tools”), determine the nucleotide sequence that corresponds to the protein sequence you chose above.
Example: Get to the original sequence of phage MS2 L-protein from its genome — phage MS2 genome - Nucleotide - NCBI
Lysis protein DNA sequence:
atggaaacccgattccctcagcaatcgcagcaaactccggcatctactaatagacgccggccattcaaacatgaggattacccatgtcgaagacaacaaagaagttcaactctttatgtattgatcttcctcgcgatctttctctcgaaatttaccaatcaattgcttctgtcgctactggaagcggtgatccgcacagtgacgactttacagcaattgcttacttaa
Response
I used the Sequence Manipulation Suite to do the reverse translation :)
Reverse translation of
sp|P19449|BCSA1_KOMXYCellulose synthase catalytic subunit [UDP-forming] OS=Komagataeibacter xylinus — 2262 base sequence of most likely codons:
Once a nucleotide sequence of your protein is determined, you need to codon optimize your sequence. You may, once again, utilize google for a “codon optimization tool”. In your own words, describe why you need to optimize codon usage. Which organism have you chosen to optimize the codon sequence for and why?
Example from Codon Optimization Tool | Twist Bioscience while avoiding Type IIs enzyme recognition sites BsaI, BsmBI, and BbsI
Lysis protein DNA sequence with Codon-Optimization:
ATGGAAACCCGCTTTCCGCAGCAGAGCCAGCAGACCCCGGCGAGCACCAACCGCCGCCGCCCGTTCAAACATGAAGATTATCCGTGCCGTCGTCAGCAGCGCAGCAGCACCCTGTATGTGCTGATTTTTCTGGCGATTTTTCTGAGCAAATTCACCAACCAGCTGCTGCTGAGCCTGCTGGAAGCGGTGATTCGCACAGTGACGACCCTGCAGCAGCTGCTGACCTAA
Response
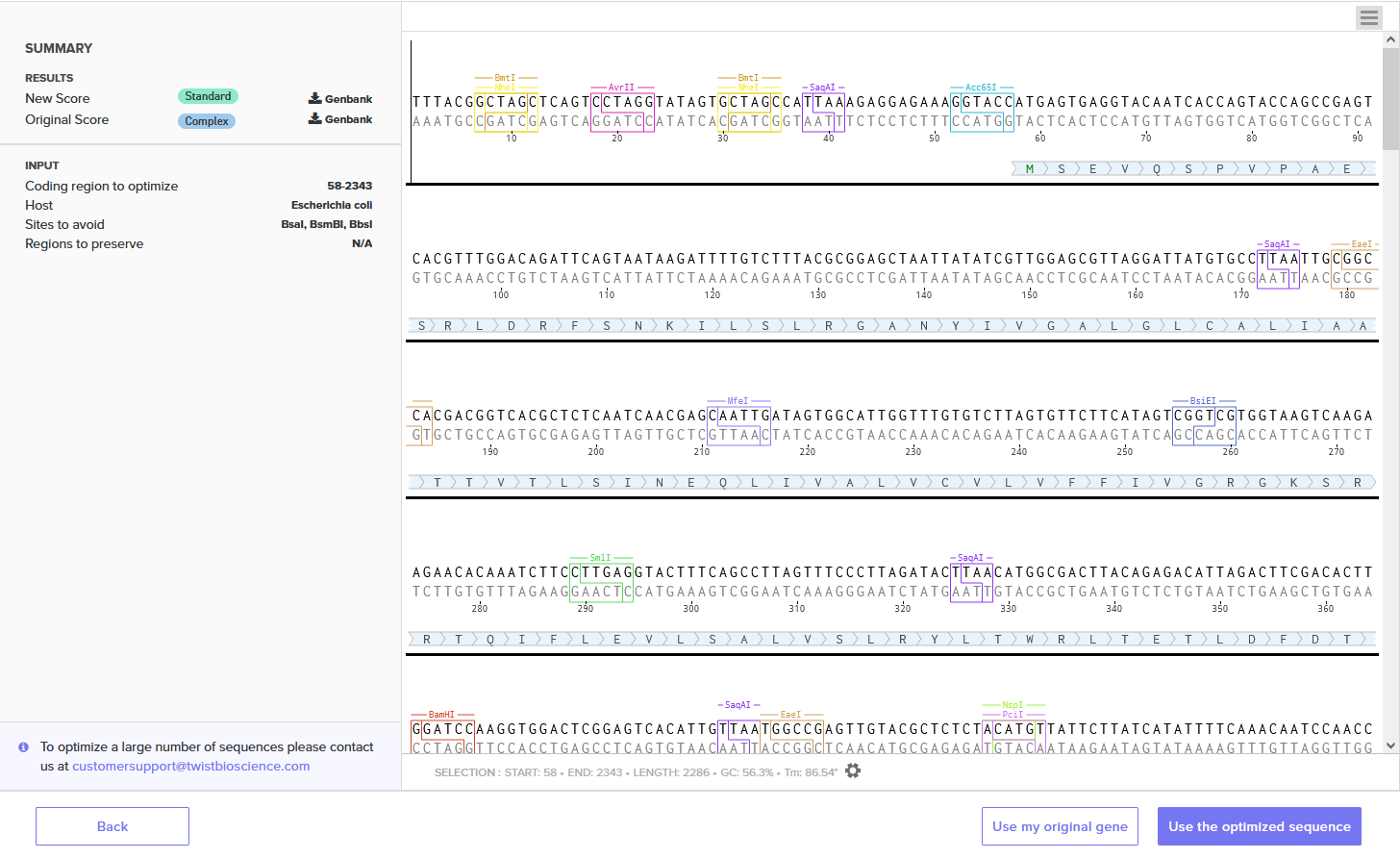
Different organisms preferentially use certain codons over others due to varying tRNA availability. By optimizing the BcsA sequence for E. coli, we ensure the ribosomes can read the sequence efficiently and produce high yields of protein. I used Twist Bioscience’s Codon Optimization Tool, optimized for Escherichia coli, avoiding restriction sites BsaI, BsmBI, and BbsI.
Improved DNA[1]: GC=55.06%, CAI=0.93
What technologies could be used to produce this protein from your DNA? Describe in your words the DNA sequence can be transcribed and translated into your protein. You may describe either cell-dependent or cell-free methods, or both.
Response
To produce cellulose synthase (BcsA) from Komagataeibacter xylinus in a lab setting, we can use a cell-dependent method with E. coli as the host organism. The codon-optimized BcsA gene is inserted into a plasmid, which is then introduced into E. coli via transformation. Once inside the cell, the E. coli transcription machinery reads the DNA and produces a messenger RNA (mRNA) copy of the BcsA gene. The ribosome then translates that mRNA into the BcsA protein by reading each 3-base codon and adding the corresponding amino acid. By cultivating the E. coli at scale, we can produce large quantities of cellulose synthase — and because we codon optimized the sequence for E. coli, this process happens efficiently. This approach also lets us modify the BcsA sequence to potentially engineer cellulose with different material properties for use in bacterial cellulose leather applications.
Response
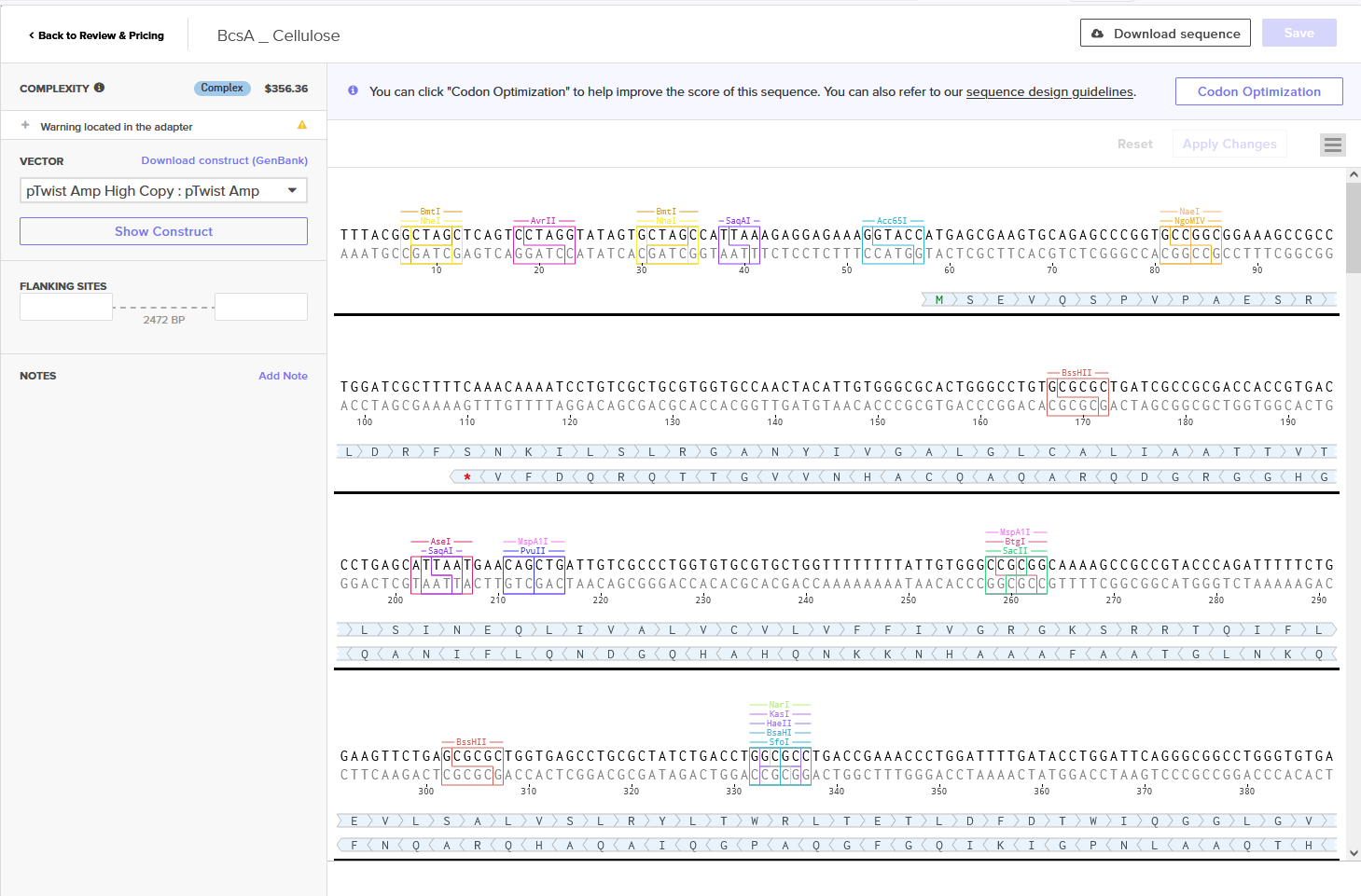
Annotated Benchling insert fragment and Twist order below:
Twist Optimized Cellulose Synthase in Plasmid
Twist Optimized Order
Twist Optimized
Debugging DNA
Download BcsA Cellulose 2nd Try (.gb)
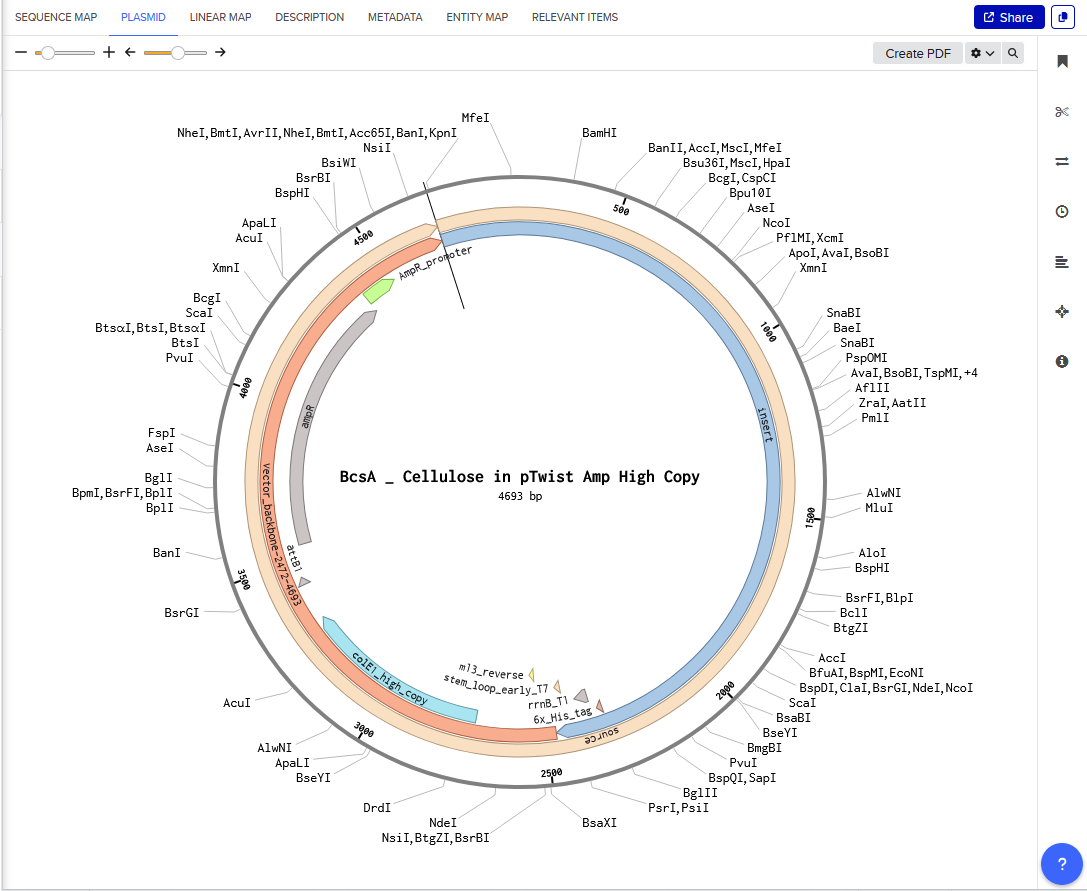

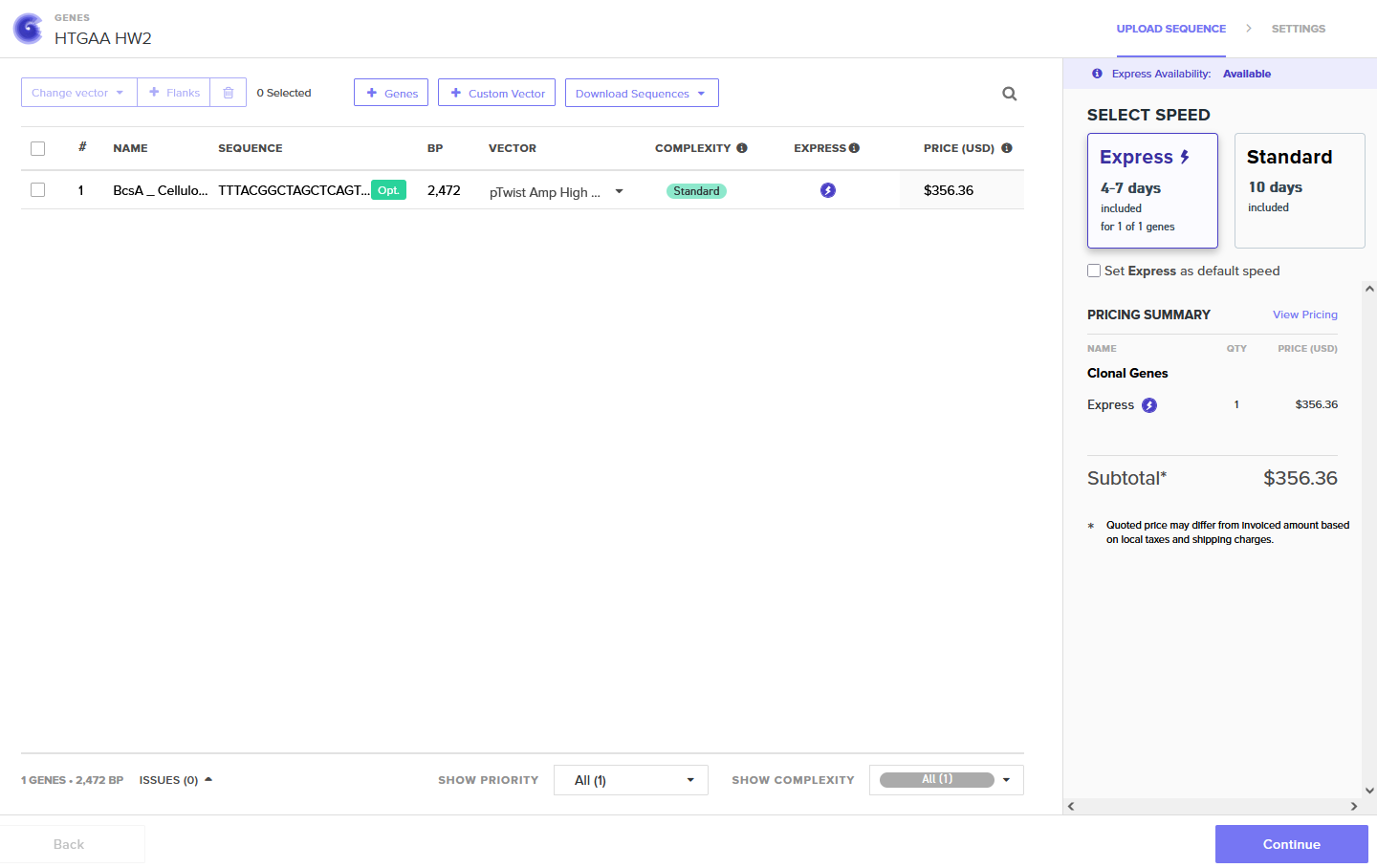



Download BcsA Cellulose Sequence (.pdf)
| MIT/Harvard students | Required |
| Committed Listeners | Required |
(i) What DNA would you want to sequence and why?
I want to read the sequence off of Komagataeibacter xylinus (the bacteria responsible for cellulose production in kombucha) to pull the cellulose synthase gene for further analysis.
I can also use this to validate my samples from Twist and QA my current batch of E. coli or other model bacteria I’m using.
(ii) What technology or technologies would you use to perform sequencing on your DNA and why?
I would use Oxford Nanopore sequencing to read my DNA.
Is your method first-, second- or third-generation or other? How so? Nanopore sequencing is a third generation technology that lets us read our DNA strands in one shot, without fragmentation or short-read reassembly.
What is your input? How do you prepare your input? List the essential steps.
What are the essential steps of your chosen sequencing technology — how does it decode the bases (base calling)? A protein nanopore sits in a membrane with an electrical current running through it. As a single DNA strand is ratcheted through the pore one base at a time, each base disrupts the current differently — A, T, G, C each produce a characteristic electrical signal. Software decodes this signal into a sequence.
What is the output of your chosen sequencing technology? Nanopore produces long reads up to 100kb+ at high accuracy, which means we can read our entire 2,472 bp plasmid insert in a single pass to verify the cellulose synthase gene transfer worked correctly.
(i) What DNA would you want to synthesize and why?
I want to write/manufacture the BcsA cellulose synthase expression cassette from Komagataeibacter xylinus, optimized for use in E. coli or yeast. I want to transfer the cellulose-producing capability into a faster but still cheap host organism to iterate on bacterial cellulose leather production.
(ii) What technology or technologies would you use to perform this DNA synthesis and why?
I would perform my synthesis using phosphoramidite chemical synthesis via Twist Bioscience, cloned into the pTwist Amp High Copy vector.
What are the essential steps of your chosen synthesis method?
What are the limitations of your synthesis method in terms of speed, accuracy, scalability?
(i) What DNA would you want to edit and why?
I want to edit the BcsA gene to change how the amino acids control the polymerization of cellulose chains and how they form fibers. By making targeted mutations we could potentially change:
(ii) What technology or technologies would you use to perform these DNA edits and why?
CRISPR-Cas9
How does your technology of choice edit DNA?
What preparation do you need to do and what are the inputs?
What are the limitations of your editing method in terms of efficiency or precision?
Assignees for this section
| MIT/Harvard students | Required |
| Committed Listeners | Required |
Your task this week is to Create a Python file to run on an Opentrons liquid handling robot.
https://opentrons-art.rcdonovan.com/?id=32iqzh4w9m44o5k


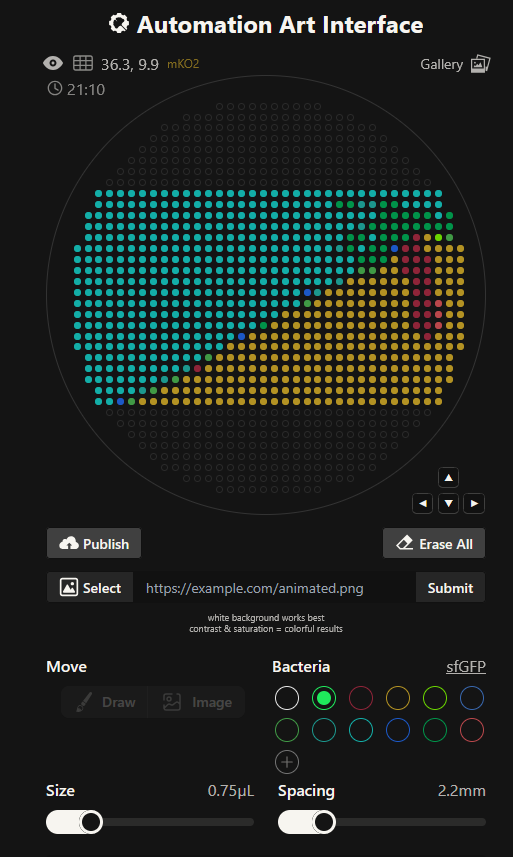

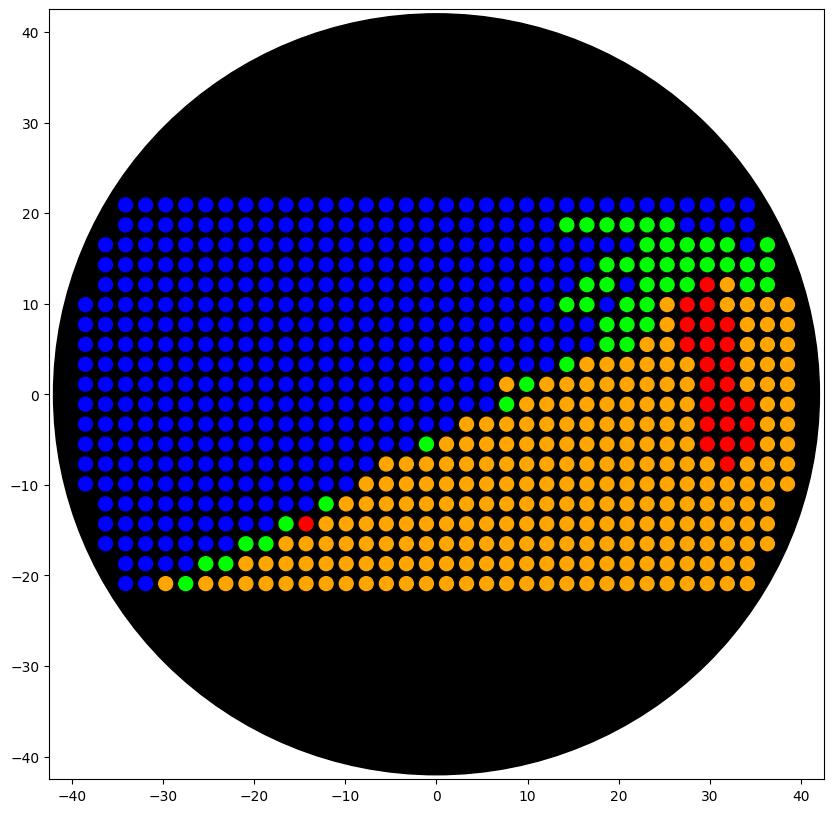

Using the coordinates from the GUI, follow the instructions in the HTGAA26 Opentrons Colab to write your own Python script which draws your design using the Opentrons.
Ask for help early!
If you are having any trouble with scripting, contact your TAs as soon as possible for help.
Do not wait until your scheduled robot time slot or you may not be able to complete this assignment!
If the Python component is proving too problematic even with AI and human assistance, download the full Python script from the GUI website and submit that:


Use the download icon pointed to by the red arrow in this diagram.
If you use AI to help complete this homework or lab, document how you used AI and which models made contributions.
Sign up for a robot time slot if you are at MIT/Harvard/Wellesley or at a Node offering Opentrons automation. The Python script you created will be run on the robot to produce your work of art!
Submit your Python file via this form.
I utilized gemini pro 3.5 and I was able to have it create the nested for loops needed to dispense on a per color basis.
Colab available here: Open My Opentrons Colab Script
Assignees for this section
| MIT/Harvard students | Required |
| Committed Listeners | Required |
One of the great parts about having an automated robot is being able to precisely mix, deposit, and run reactions without much intervention, and design and deploy experiments remotely.
For this week, we’d like for you to do the following:
https://pmc.ncbi.nlm.nih.gov/articles/PMC11933169/ Enhanced production of bacterial cellulose with a mesh dispenser vessel-based bioreactor.
From my reading of the paper the team created structures for the Komagataeibacter out of mesh and automated nutrient dispensing, the quality and speed of Komagataeibacter’s cellulose production were significantly improved over the control of pellicle formation to produce the bacterial leather.
Loh J, Arnardottir T, Gilmour K, Zhang M, Dade-Robertson M. Enhanced production of bacterial cellulose with a mesh dispenser vessel-based bioreactor. Cellulose (Lond). 2025;32(4):2209-2226. doi: 10.1007/s10570-024-06367-w. Epub 2025 Jan 29. PMID: 40144312; PMCID: PMC11933169.
While your description/project idea doesn’t need to be set in stone, we would like to see core details of what you would automate. This is due at the start of lecture and does not need to be tested on the Opentrons yet.
Example 1: You are creating a custom fabric, and want to deposit art onto specific parts that need to be intertwined in odd ways. You can design a 3D printed holder to attach this fabric to it, and be able to deposit bio art on top. Check out the Opentrons 3D Printing Directory.
Example 2: You are using the cloud laboratory to screen an array of biosensor constructs that you design, synthesize, and express using cell-free protein synthesis.
For my final project I can see the possibility of the Opentrons helping by inserting the plasmids into my target organism, automating PCRs for sequencing, and being able to test different combinations of variables very quickly such as growth media or quantity of plasmids needed to successfully express the cellulose synthase within my target organism.
For broader automation goals I would like to also implement automated nutrient delivery systems and leather harvesting systems in order to reduce human oversight of microbial leather production. I am thinking of implementing fluid transfer pumps, temperature controls, and computer vision models to monitor the growth of bacterial leather in my target organism. This would be done with custom made hardware and implementing transfer learning for the vision policies.
Assignees for this section
| MIT/Harvard students | Required |
| Committed Listeners | Required |
As explained in this week’s recitation, add 1-3 slides with 3 ideas you have for an Individual Final Project in the appropriate slide deck for MIT/Harvard/Wellesley students or for Commited Listeners. Be sure to put your name on your slide(s); for CLs, also put your city and country on your slide(s) and be sure you’re putting your slide(s) in your Node’s section of the deck.
Automated microbial leather production
Utilizing sensors to monitor target organism (Komagataeibacter) health and nutrient conditions in growth tank
Leveraging fluidics automations for a custom nutrient delivery fluidics system
Computer vision learning to develop contamination detection models
Expressing cellulose growth in model organisms
Adapting fluorescent protein expression in Komagataeibacter