Week 5 HW: Protein Design Part II
PART A: Therapeutic Peptide Design for SOD1 (ALS)
1. Objective
The goal of this section was to design a synthetic peptide binder capable of stabilizing the mutant SOD1 (A4V) protein, a primary cause of Amyotrophic Lateral Sclerosis (ALS). We aimed to create a “molecular shield” to prevent toxic protein aggregation.
2. Methodology & Results
We used AI-driven models to generate and validate peptide candidates. Our best result achieved a significant binding confidence score.
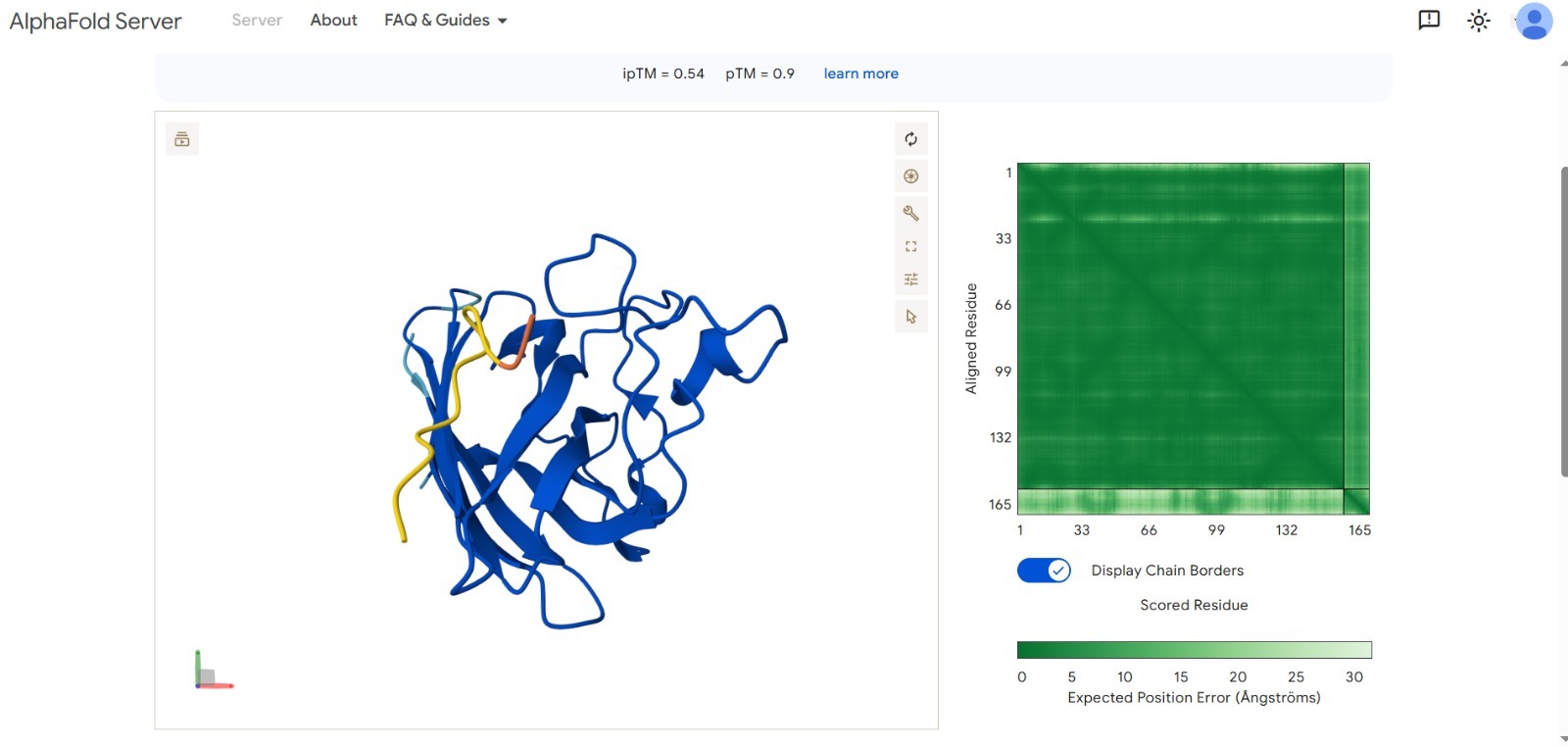
- Best Score: ipTM 0.54 (Optimized via targeted design).
- Structural Validation: The AlphaFold visualization shows the peptide (in yellow/orange) effectively docking onto the SOD1 surface.

3. Physicochemical & Safety Profile
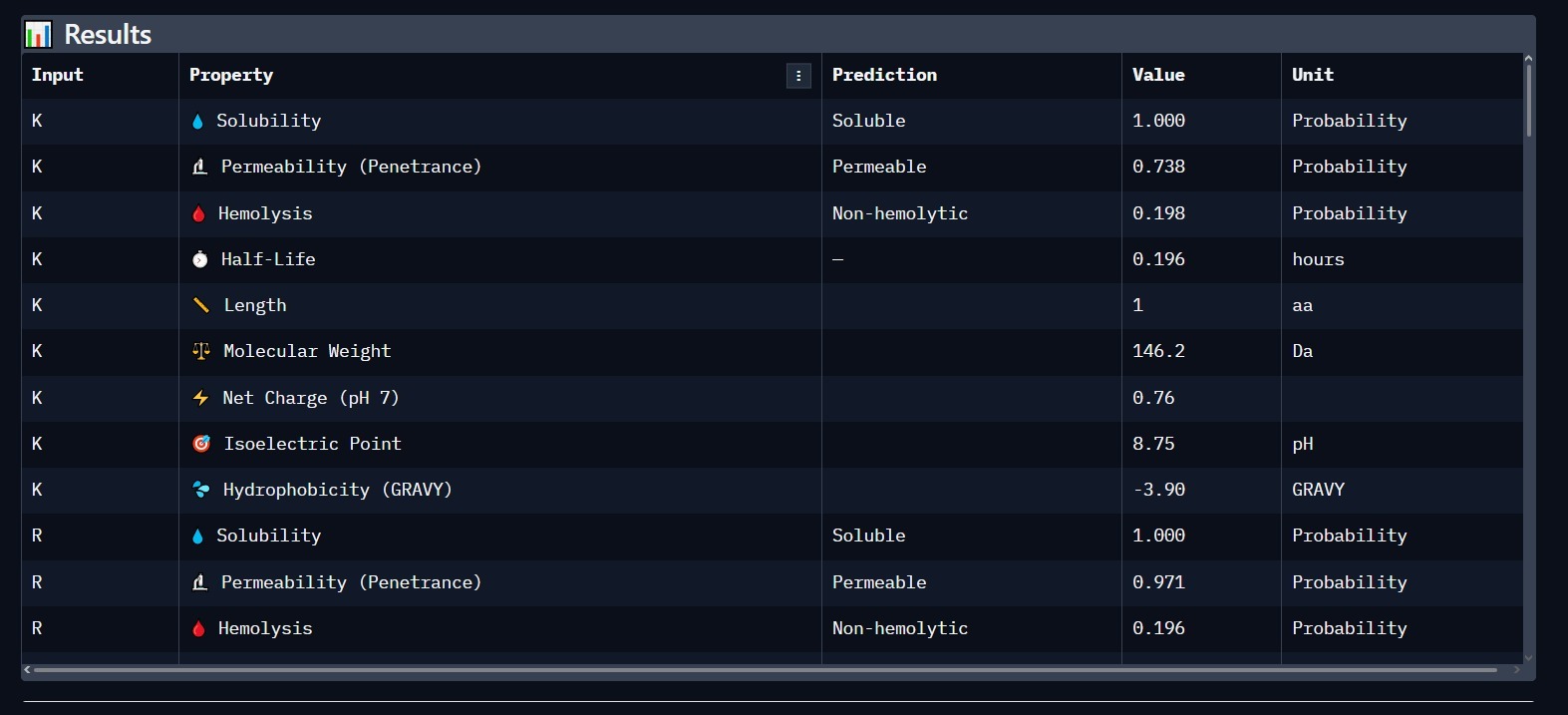
Using PeptiVerse, we analyzed the lead candidate for clinical viability. The results confirm that the peptide is highly soluble and safe for biological systems.
- Solubility: 1.000 (Highly Soluble).
- Hemolysis: Non-hemolytic (Safe for blood contact).

PART C: Phage Lysis Protein Engineering
1. The Challenge: Overcoming E. coli Resistance
Bacteriophages use the L-Protein to lyse (kill) bacteria. E. coli develops resistance by mutating the DnaJ chaperone, which prevents the L-protein from folding correctly. Our objective was to engineer L-protein mutants that fold independently of DnaJ.
2. Structural Analysis (Wild-Type)
Using AlphaFold2 Multimer, we simulated the interaction between DnaJ and the L-Protein to identify structural vulnerabilities.
Sequence Coverage: Analysis shows high coverage for DnaJ but very low evolutionary conservation for the L-protein region (positions 380+), indicating it is a unique viral protein.
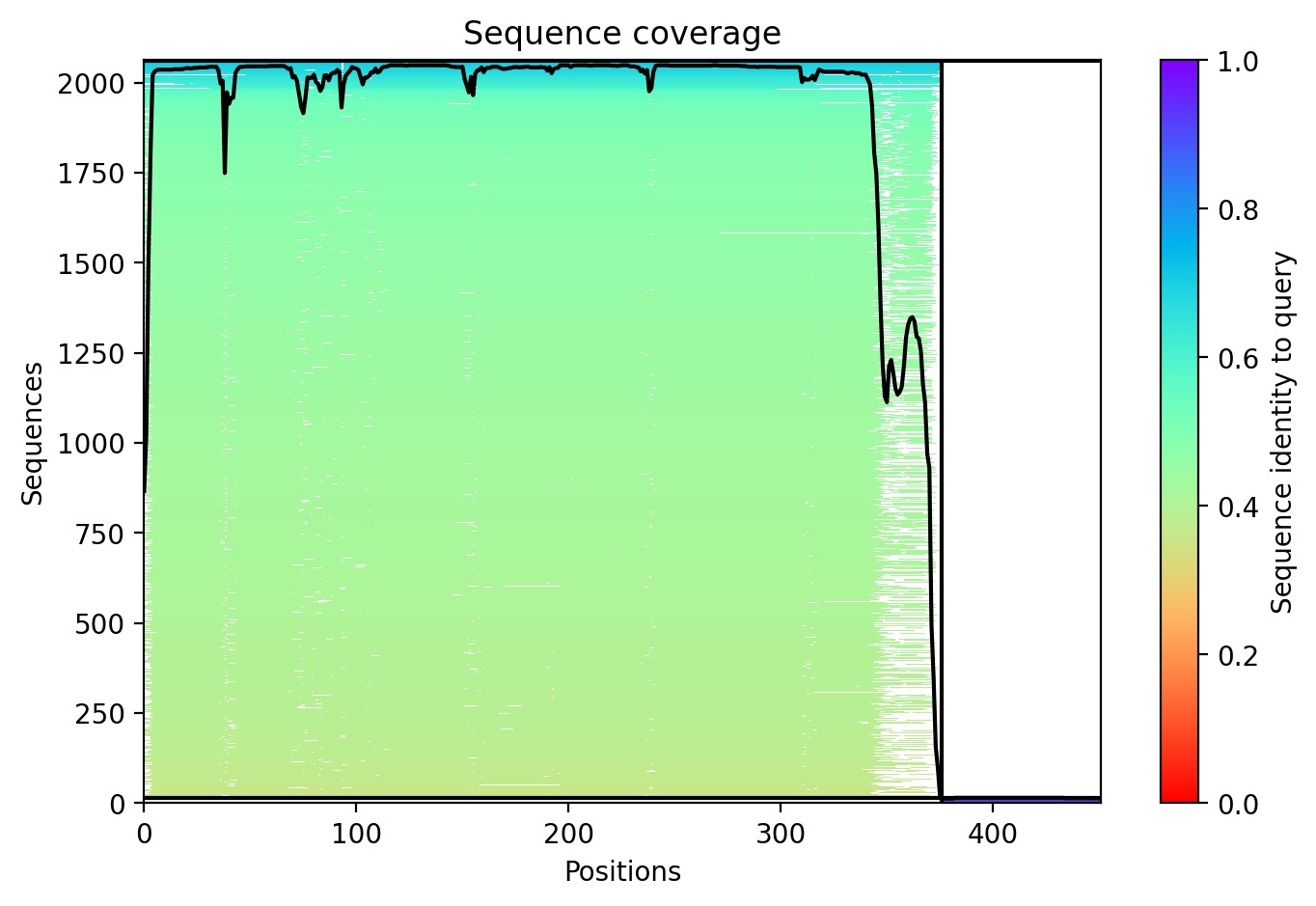

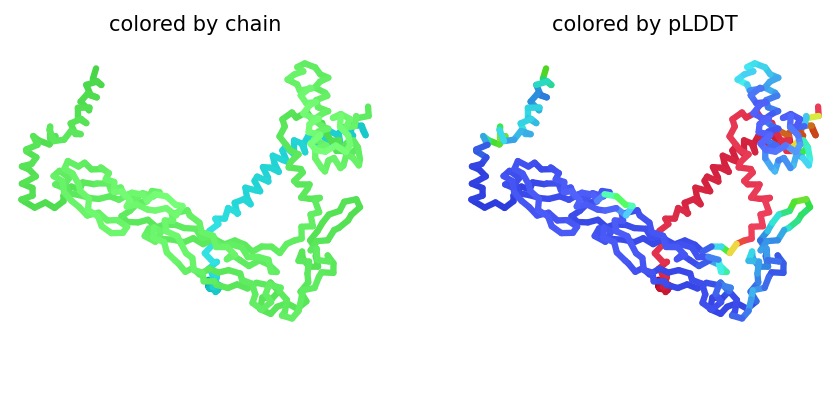
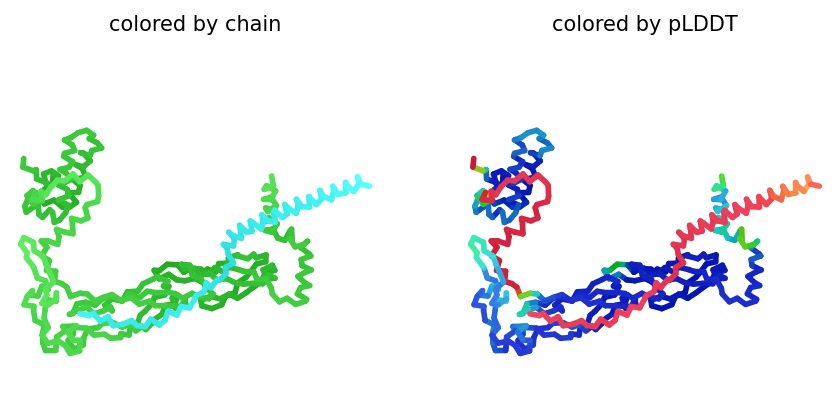
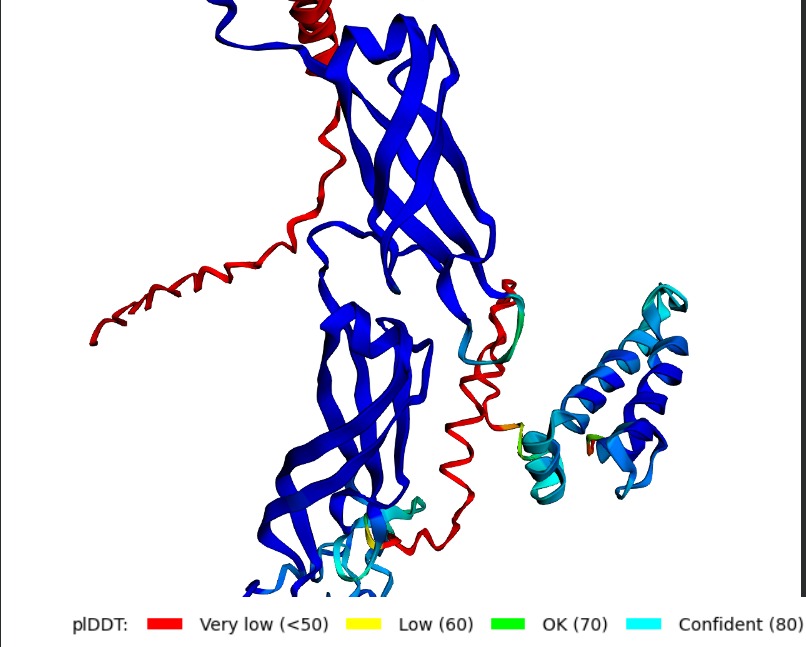
Structural Stability (pLDDT): The L-protein (Chain B) exhibits very low pLDDT scores (red color), confirming it is structurally unstable without the DnaJ chaperone.
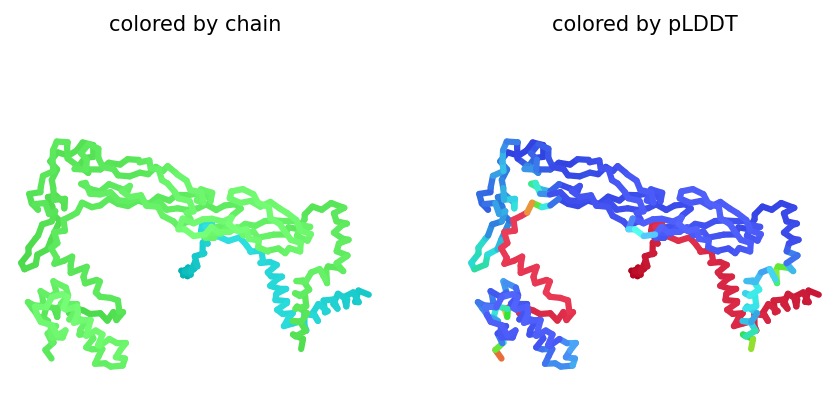



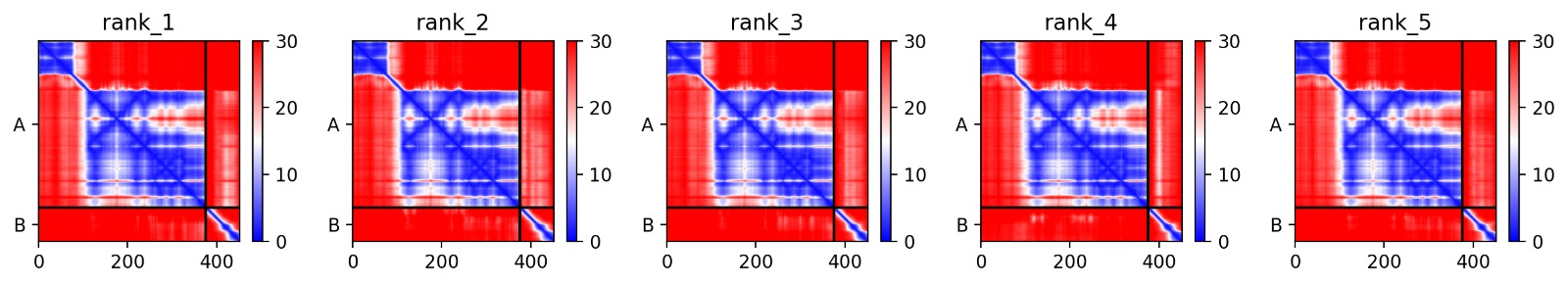
Interaction Matrix (PAE): The PAE plots show the specific residues where the L-protein contacts DnaJ. This data was used to guide our mutation strategy.

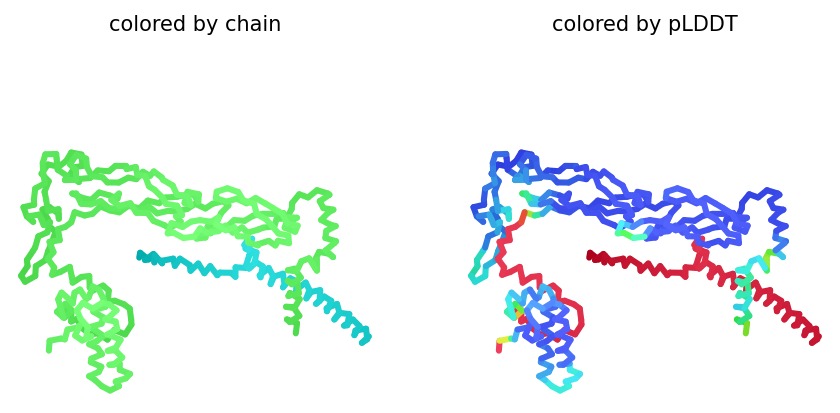


3. Proposed Engineering Strategy (5 Lead Mutants)
Based on the structural data, I have designed 5 mutations to enhance autonomous folding and lysis efficiency:
| # | Mutation | Region | Engineering Goal |
|---|---|---|---|
| 1 | L25P | Soluble | Increase structural autonomy (DnaJ independence). |
| 2 | R14G | Soluble | Stabilize the domain responsible for folding. |
| 3 | F52L | Transmembrane | Enhance membrane insertion for faster killing. |
| 4 | W45A | Transmembrane | Optimize pore stability in the bacterial wall. |
| 5 | S31A | Soluble | Stabilize the N-terminal soluble domain. |
4. Defining Success
A successful mutant is defined by its ability to maintain a high Plaque Forming Unit (PFU) count on resistant bacterial strains. Computationally, success is marked by an improved pLDDT score, indicating the protein has gained structural independence.
Final Synthesis
This project demonstrates a complete workflow in protein engineering: from designing “molecular shields” for neurodegenerative diseases (SOD1) to engineering viral proteins to bypass antibiotic resistance. Computational tools like AlphaFold and moPPIt allow us to solve critical biological challenges with high precision.