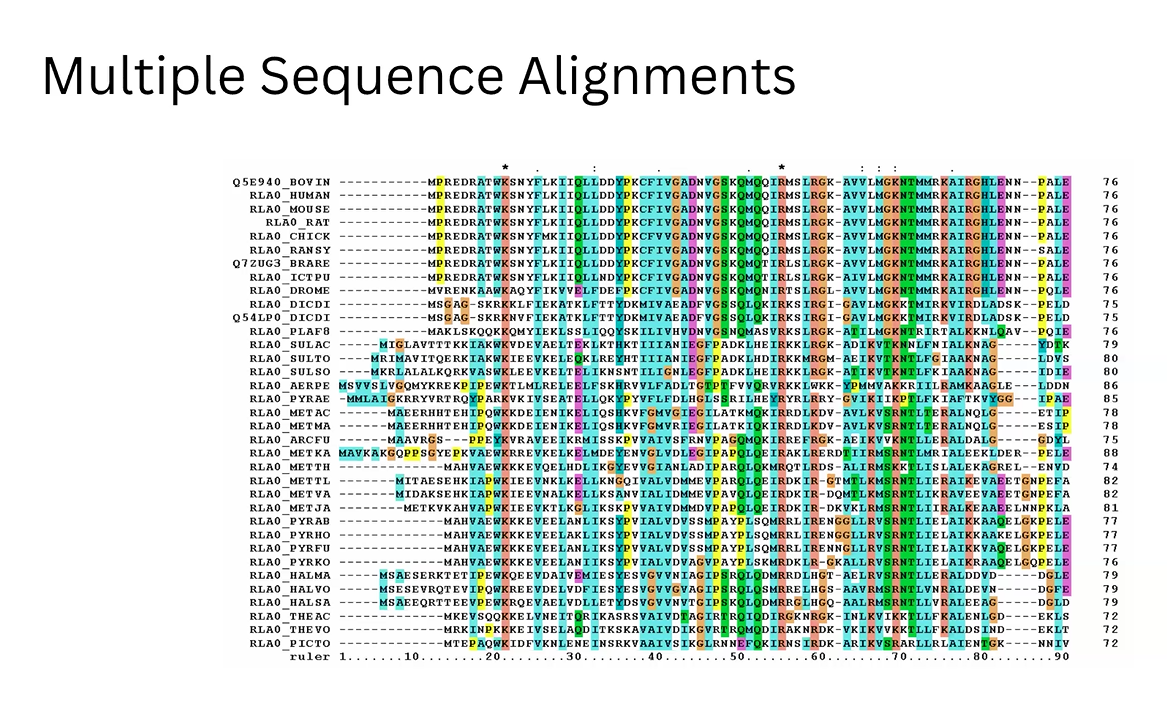
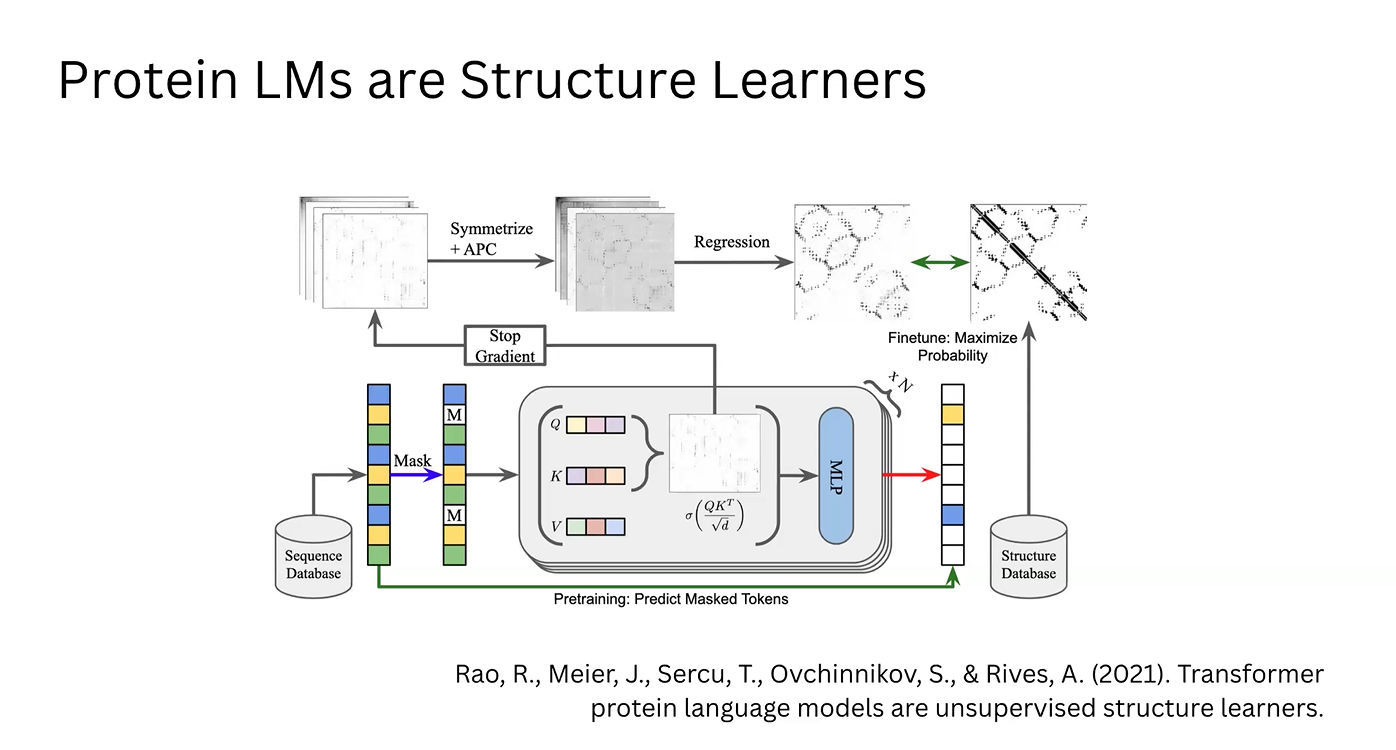
Likelihood Generation: We utilize the fundamental property of the ESM (Masked Language Modeling) to calculate amino acid probabilities based on the sequence context.
Attention Mechanism: These data points are processed as Queries (Q), Keys (K), and Values (V). This generates an Attention Matrix, where vectors for each amino acid indicate the relevance and position of other residues in the chain.
Structural Extraction (Simple ESM): The raw attention matrix is symmetrized to ensure physically plausible distances and refined using Average Product Correction (APC) to eliminate correlation noise. Through regression, a 2D contact map is obtained.
Model Optimization: This map is compared against structural databases (like the PDB) to adjust the model. This feedback loop updates the model’s weights, maximizing the log-likelihood of real-world protein structures found in nature.
Enrichment via MLP: In parallel, the correlation vectors from the attention matrix are processed by a Multi-Layer Perceptron (MLP). This generates enriched vectors that describe the physicochemical implications of the attention data.
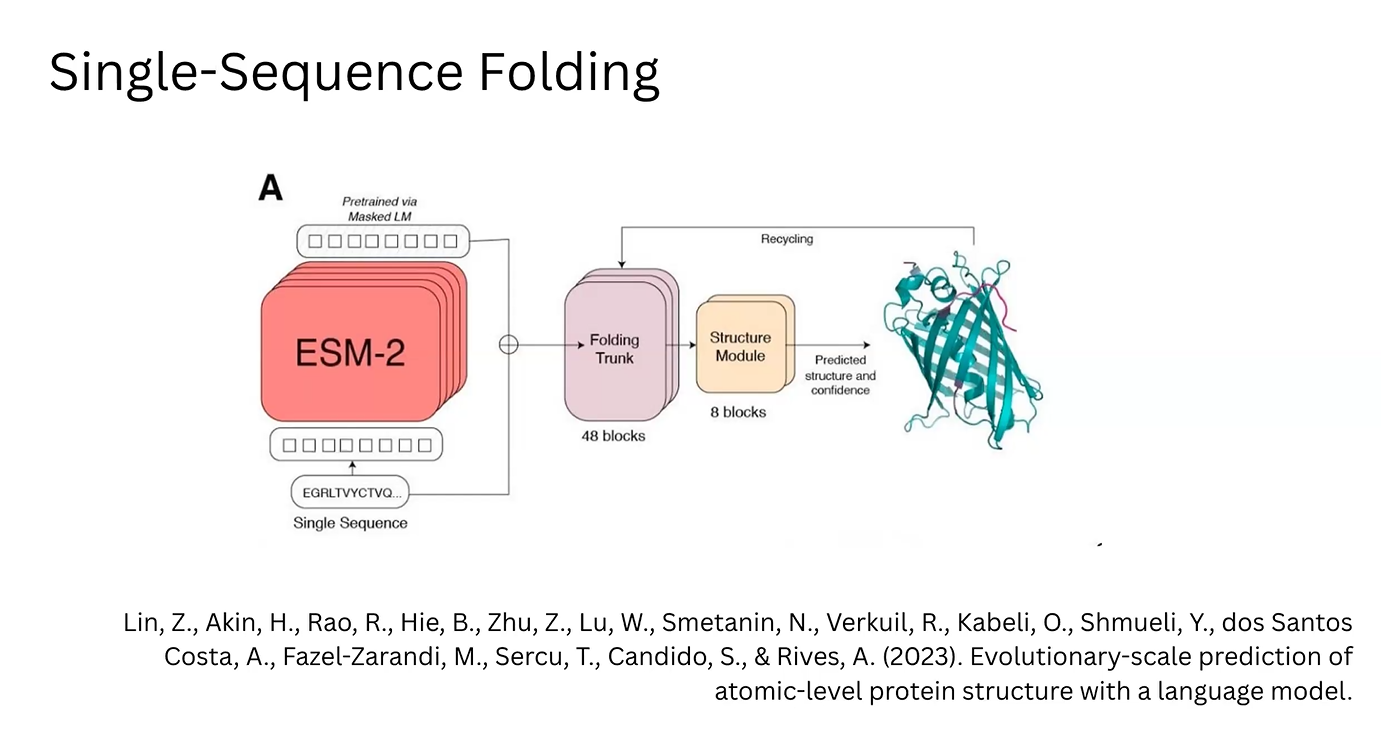
3D Prediction (ESM-2/ESMFold): In the ESM-2 architecture, these MLP-enriched vectors are fed into a “Folding Trunk” module. This module applies learned rules to predict the 3D structure, including bond angles and atomic coordinates.