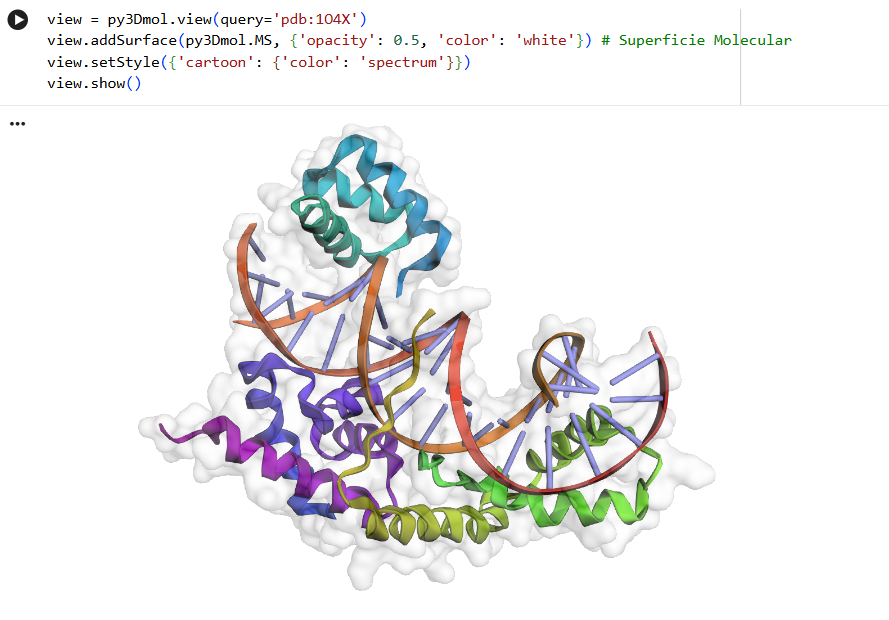
Protein analysis and visualization
Why Sox2?
This is primarily because is the next protein in the yamanaka factors and participates in a SOX2/OCT4-bound, being OCT-4 the protein I selected for week’s two homework.
In relation to Sox2 aminoacid sequence
Sequence length and composition: This protein is 317 amino acids long. Based on the frequency analysis, Serine (S) is the most frequent amino acid, appearing 36 times (11.36% of the sequence), followed closely by Glycine (G) with 35 occurrences (11.04%).
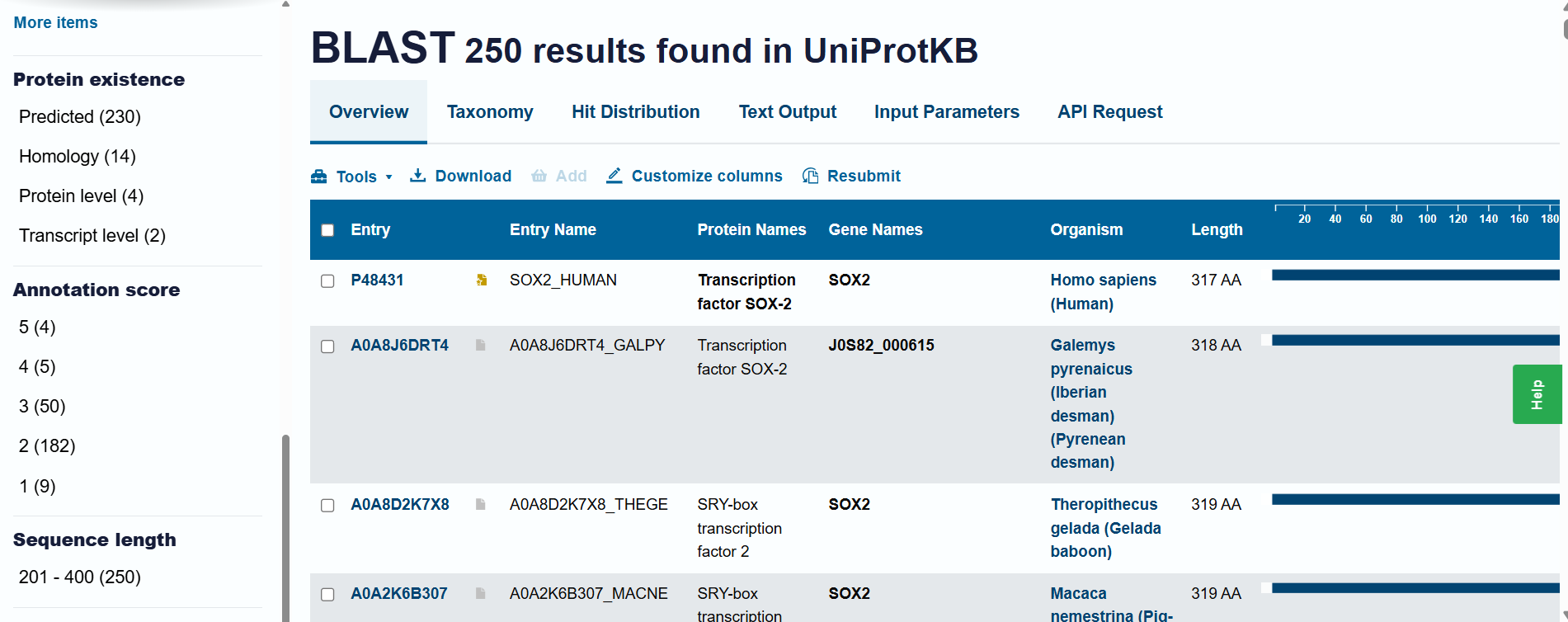
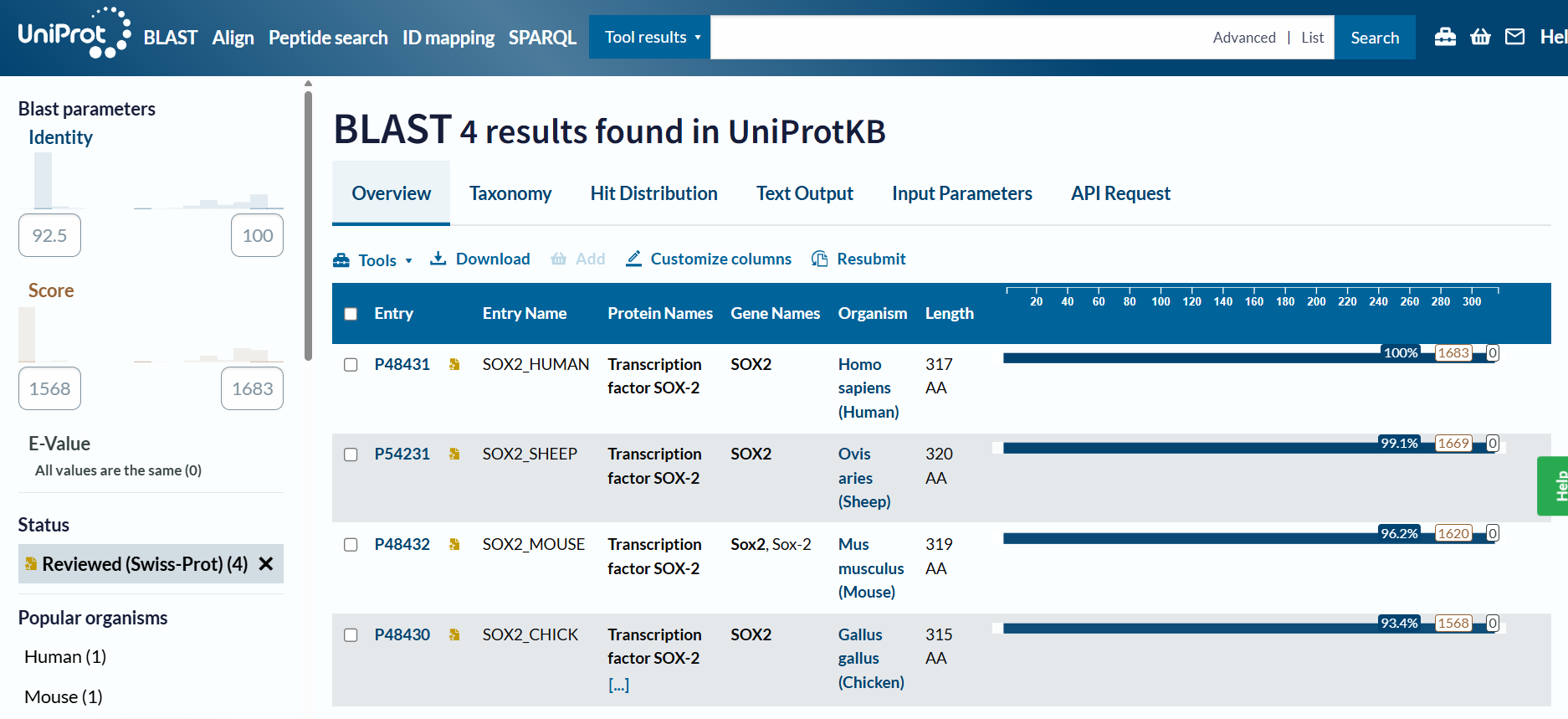
Homology and BLAST results: When including “predicted” and “homology” evidence levels, the search reached the limit of 250 results in the UniProt BLAST tool. This high number of hits reflects that Sox-2 is an extremely conserved protein across species. However, for high-confidence analysis, I primarily considered the results from the reviewed Swiss-Prot database.


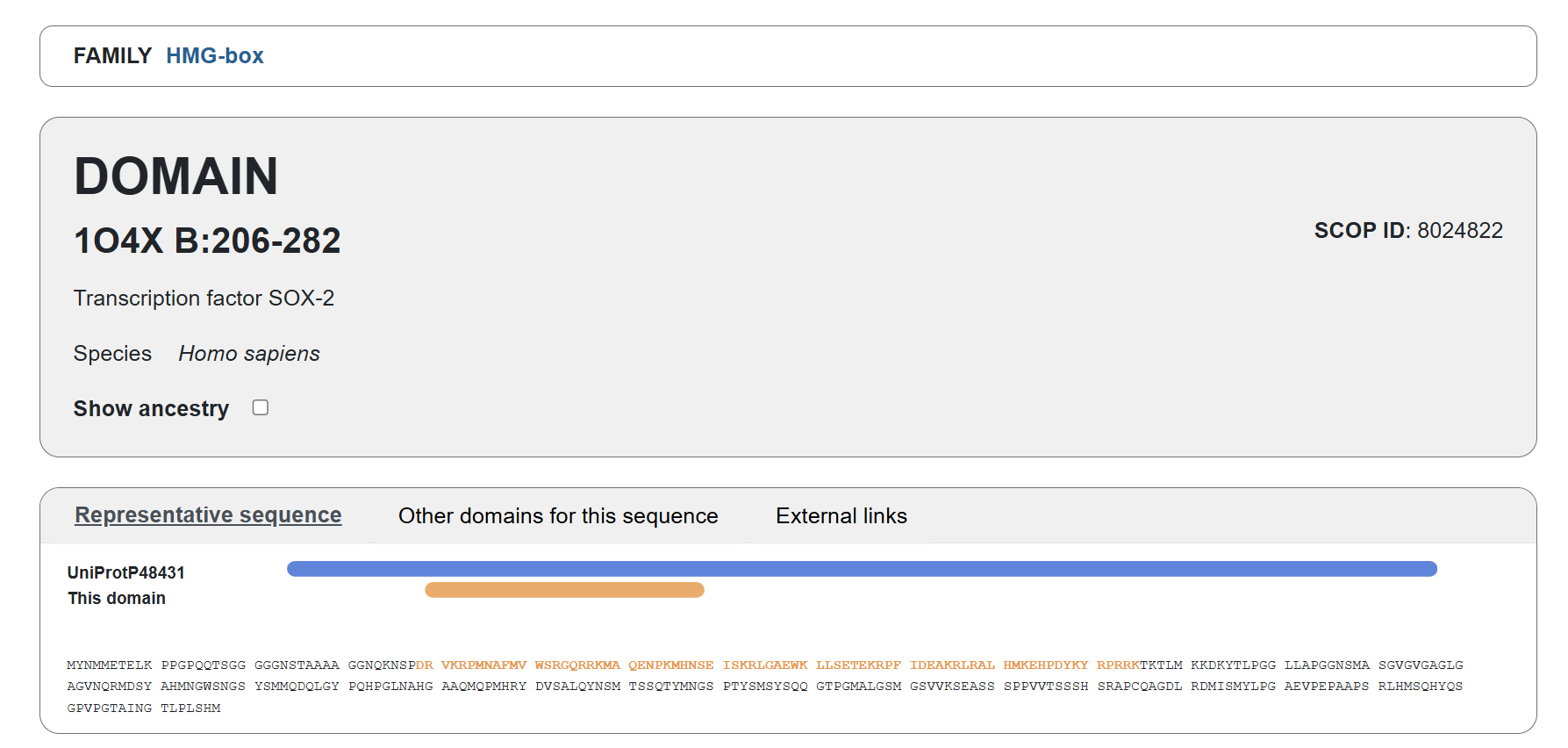
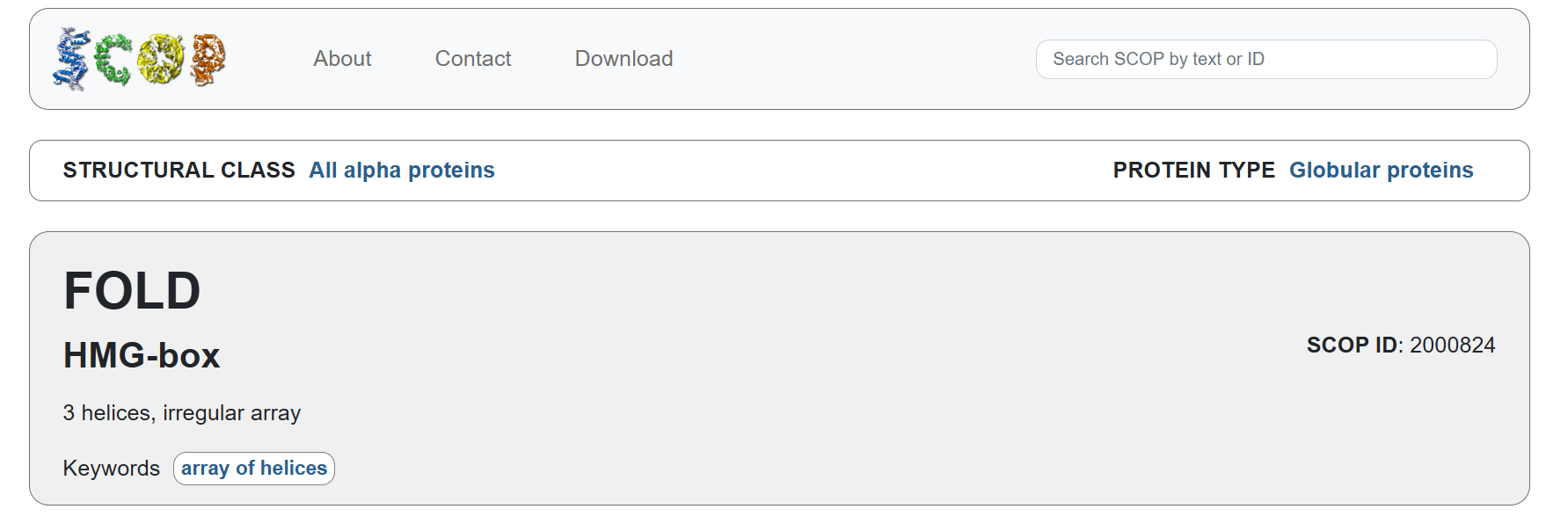
Familiy Classification: SCOP results showed that Sox-2 contains a High Mobility Group (HMG) box structural domain located between residues 206-282. This domain is characterized by its L-shaped triple α-helix fold that binds to the minor groove of DNA. While it shares this structural “HMG-box” fold with other architectural proteins, Sox-2 functionally belongs to the SOX transcription factor family. Despite recognizing the same DNA consensus motif ( 5-(A/T)(A/T)CAA(A/T)G-3 ) different Sox factors achieve target gene selectivity through differential affinity for particular flanking sequences next to consensus Sox sites, homo- or heterodimerization among Sox proteins, posttranslational modifications of Sox factors, or interaction with other co-factors (Wegner, 2010)

In relation to protein structure
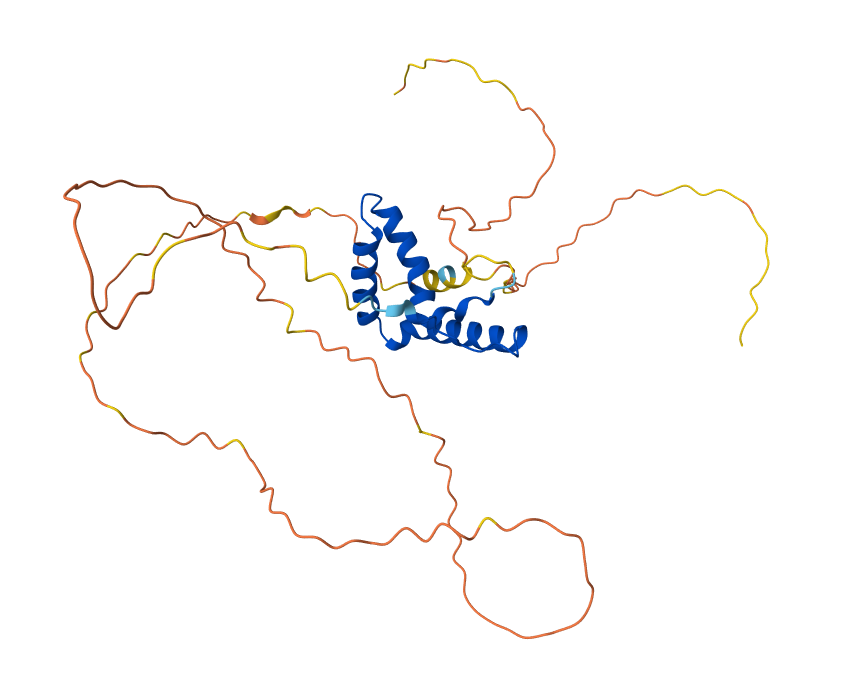
 RCSB result for P48431 : https://www.rcsb.org/groups/sequence/polymer_entity/P48431
RCSB result for P48431 : https://www.rcsb.org/groups/sequence/polymer_entity/P48431
The structure was solved and published in 2003 by Reményi et al. in the journal Genes & Development (https://doi.org/10.1101/gad.269303). Since the article its about the crystal structure of a POU/HMG/DNA ternary complex, the resolution is not below 2.70 Å, wich is not ideal for high-detail drug design, but I think the 2.70 Å resolution obtained is considered a good quality structure for analyzing protein-DNA ternary complexes.
As I mentioned before, the solved structure is often a ternary complex. Apart from the SOX2 protein, it includes the POU domain of the OCT4 protein and a specific DNA oligonucleotide (the FGF4 enhancer)
Acording to SCOP, it belongs to the HMG-box family within the structural class all alpha protein

In relation to the molecule visualization:
Since I ran into some strange errors while trying to use Conda, I used pip to import the PyMOL libraries instead
 I used py3Dmol to visualize Sox2 in “cartoon” and “ball and stick” representations, and PyMOL for the “ribbon” view
I used py3Dmol to visualize Sox2 in “cartoon” and “ball and stick” representations, and PyMOL for the “ribbon” view
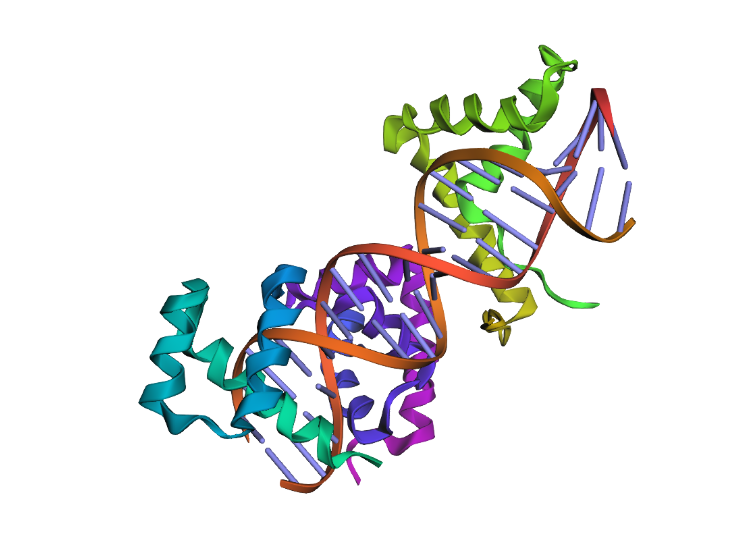

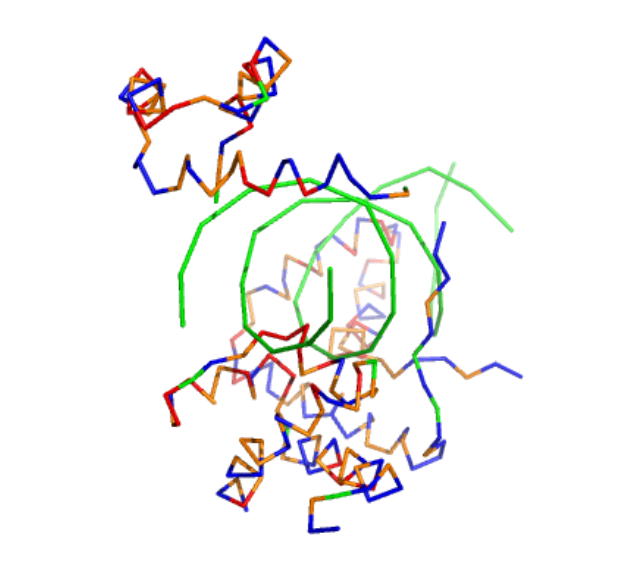

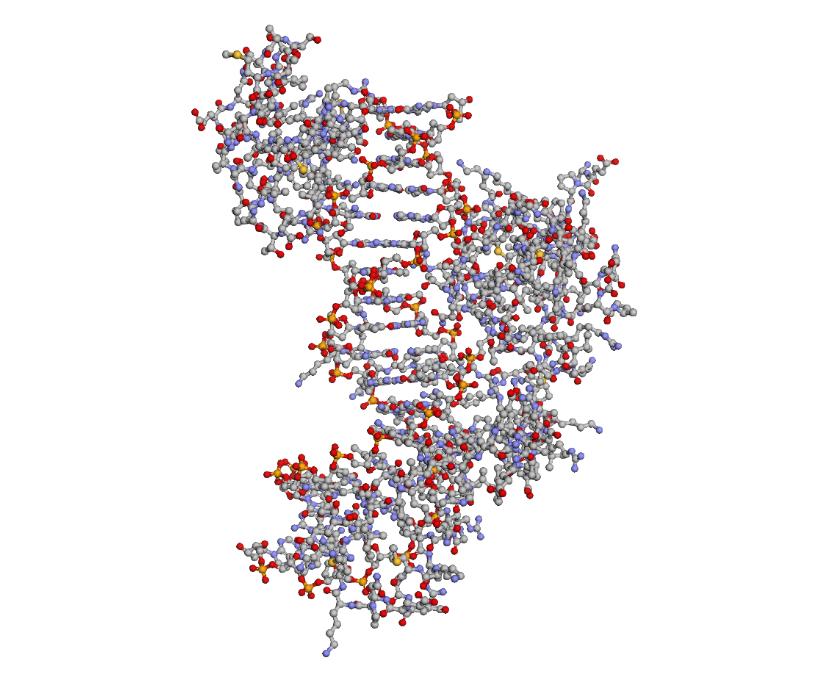

To highlight the secondary structures, I added colors: red for alpha helices, blue for beta sheets, and grey for loops
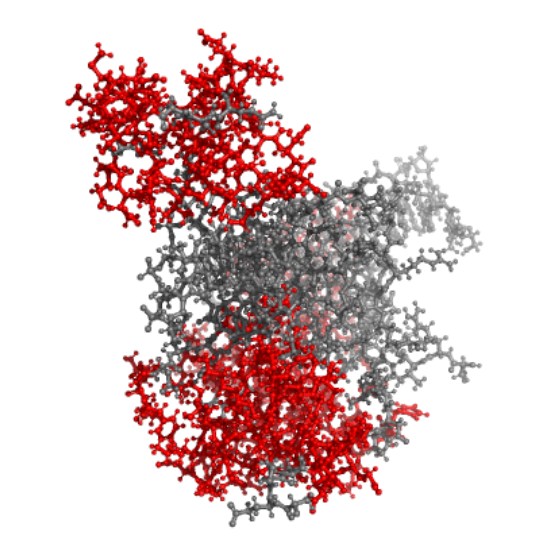

 I then used the following code to count the atoms and determine the predominant structure:
I then used the following code to count the atoms and determine the predominant structure:


By coloring the hydrophilic residues blue, the hydrophobic residues orange, and others yellow, it becomes clear how they are distributed. You can notice that the hydrophilic residues tend to interact with the phosphates in the DNA chain, while the hydrophobic ones are positioned to fit within the DNA grooves.
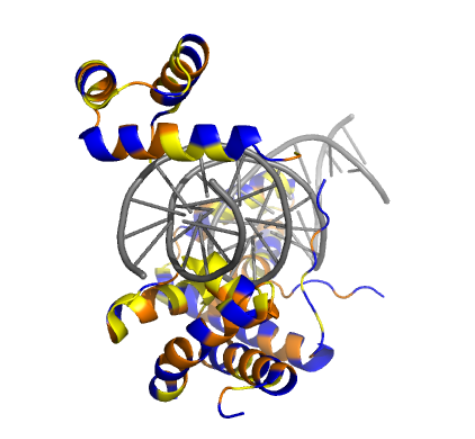

As seen in the surface visualization, the “binding pockets” of transcription factors aren’t exactly “holes.” Instead, they act more like a contoured surface that forms “hooks” around the DNA backbone.