SOD1 Binder Peptide Design
1. Generate Binders with PepMLM
First, I retrieve the human SOD1 amino acid sequence from UniProt (https://www.uniprot.org/uniprotkb/P00441/entry):
sp|P00441|SODC_HUMAN Superoxide dismutase [Cu-Zn] OS=Homo sapiens OX=9606 GN=SOD1 PE=1 SV=2 MATKAVCVLKGDGPVQGIINFEQKESNGPVKVWGSIKGLTEGLHGFHVHEFGDNTAGCTS AGPHFNPLSRKHGGPKDEERHVGDLGNVTADKDGVADVSIEDSVISLSGDHCIIGRTLVV HEKADDLGKGGNEESTKTGNAGSRLACGVIGIAQ
Then, I induced the alanine to valine mutation at residue 4 (A4V) of SOD1 sequence, resulting in the mutated version:
sp|P00441|SODC_HUMAN Superoxide dismutase [Cu-Zn] OS=Homo sapiens OX=9606 GN=SOD1 PE=1 SV=2 MATKVVCVLKGDGPVQGIINFEQKESNGPVKVWGSIKGLTEGLHGFHVHEFGDNTAGCTS AGPHFNPLSRKHGGPKDEERHVGDLGNVTADKDGVADVSIEDSVISLSGDHCIIGRTLVV HEKADDLGKGGNEESTKTGNAGSRLACGVIGIAQ
Using the collab from https://huggingface.co/ChatterjeeLab/PepMLM-650M, I selected a peptide length of 12, a top-k value of 3 (as the literature suggests this yields better and more flexible outcomes), and set the number of binders to 4.

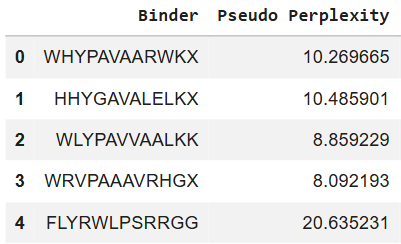
While running the Colab, I included the provided ground truth binder (FLYRWLPSRRGG) for comparison. Interestingly, I noticed that the known SOD1-binding peptide in the fourth row has an elevated perplexity as calculated by the model.

2. Evaluate Binders with AlphaFold3
I submitted the protein sequence followed by each peptide. Since three of the four peptides had ‘X’ as their last amino acid, I replaced the X with a G (Glycine) to successfully submit the jobs to AlphaFold3.
The results were as following:
Known binder FLYRWLPSRRGG: It binds to the surface near a non structured region and obtained an ipTM = 0.34.
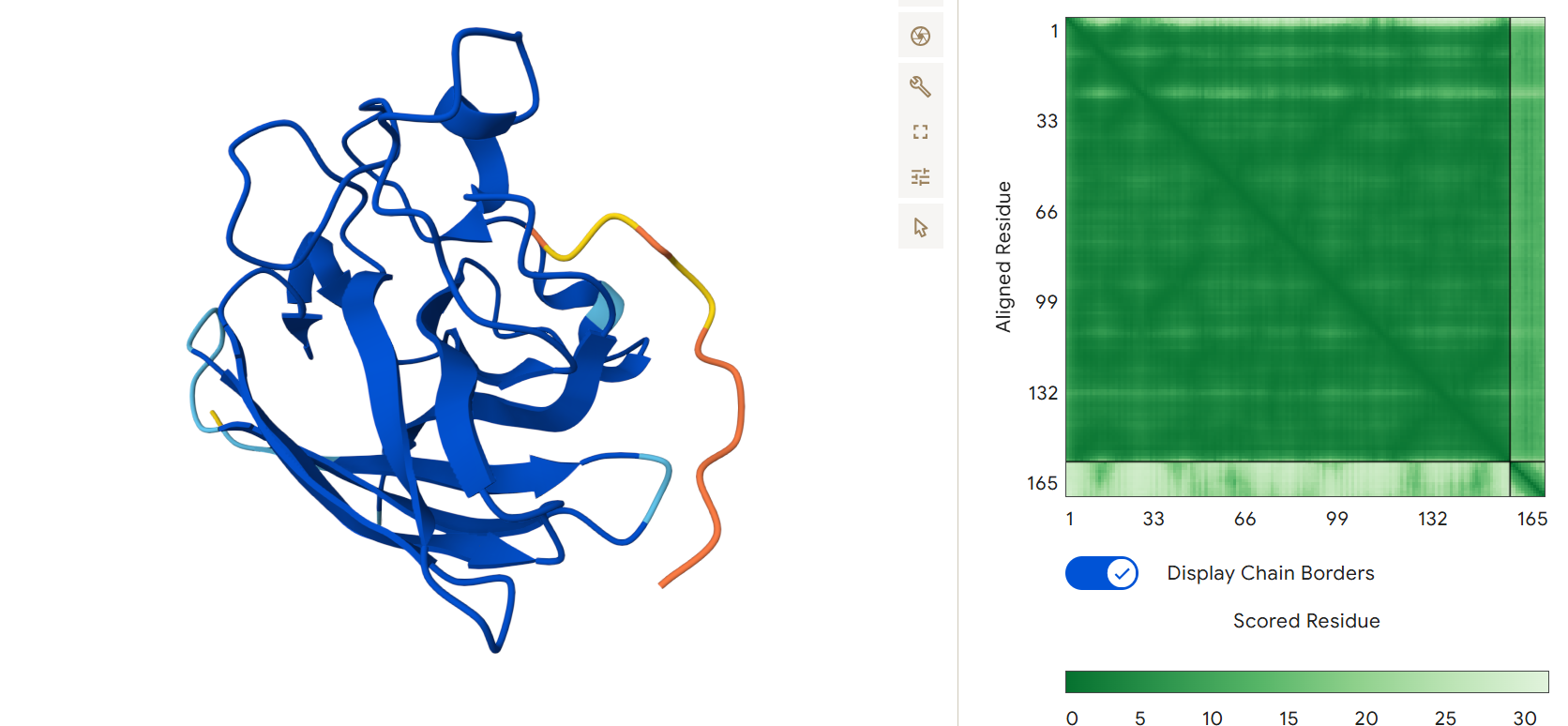

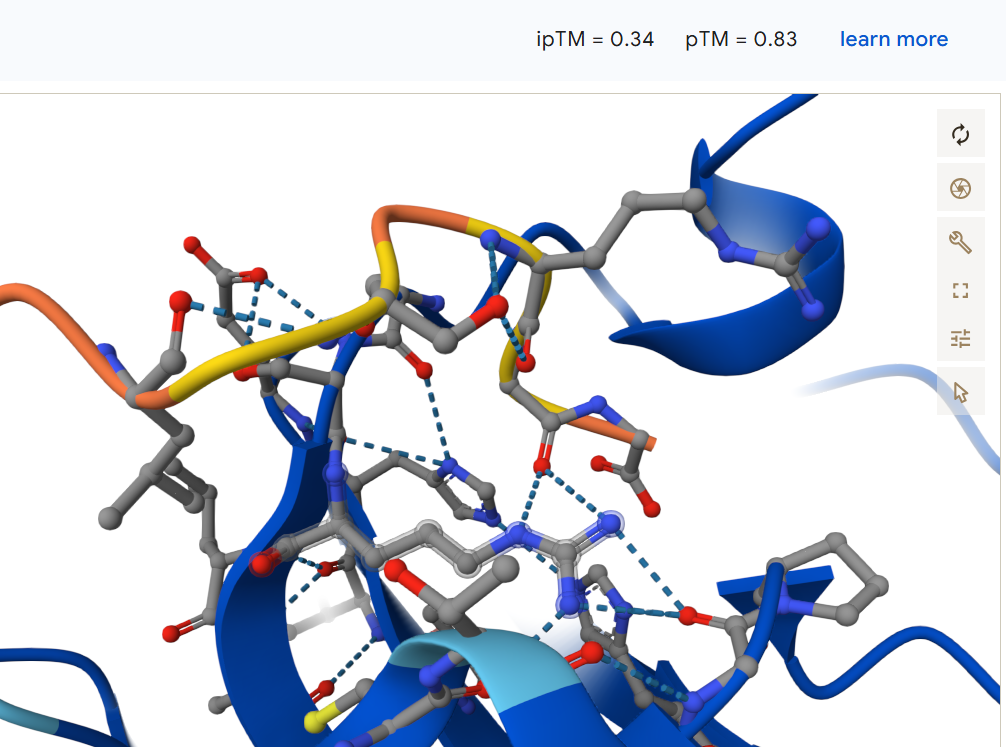

None of the generated peptides appeared to bind near any terminus; all were surface-bound and mostly located in a loop from approximately position 65 to 78.
First generated binder WHYPAVAARWKX with an ipTM = 0.31:
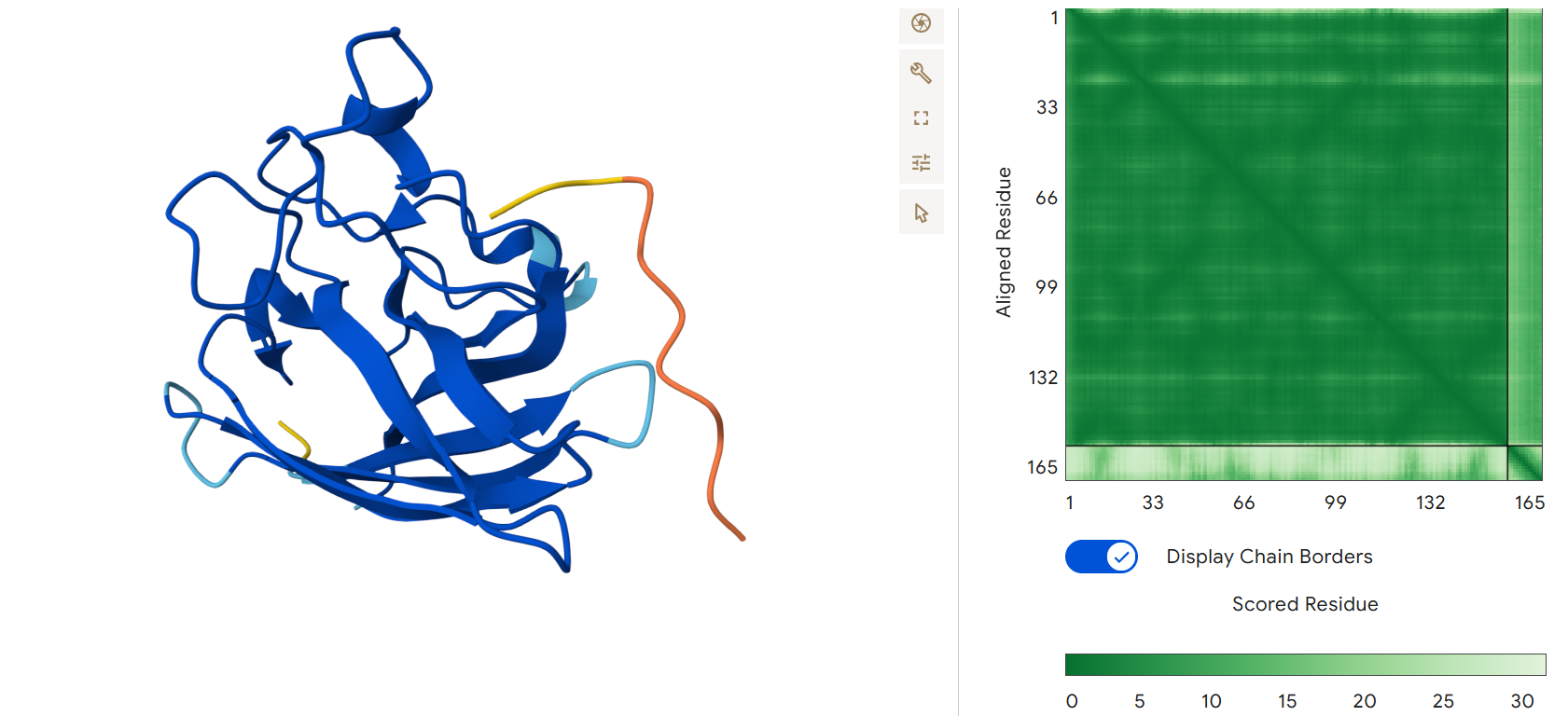


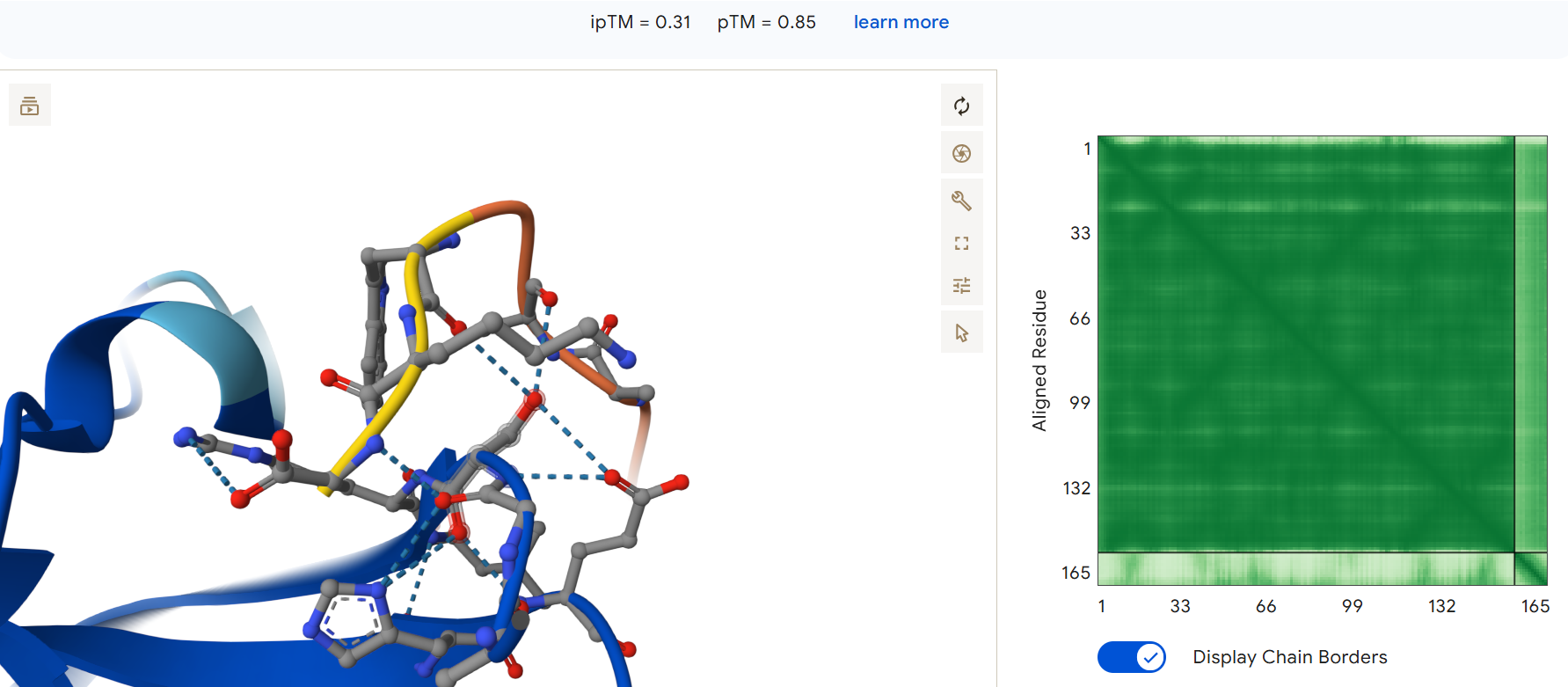

Second generated binder HHYGAVALELKX with an ipTM = 0.45:
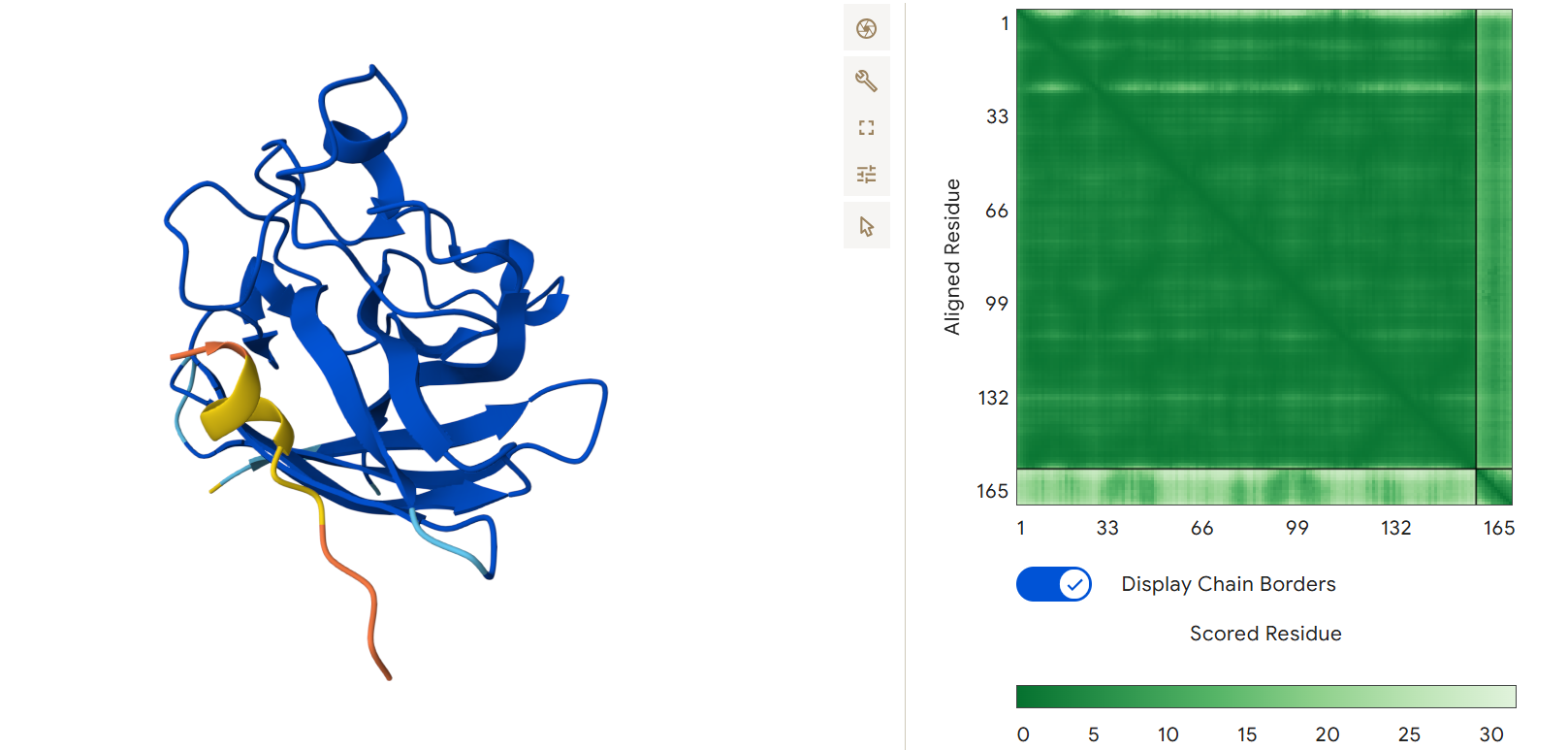

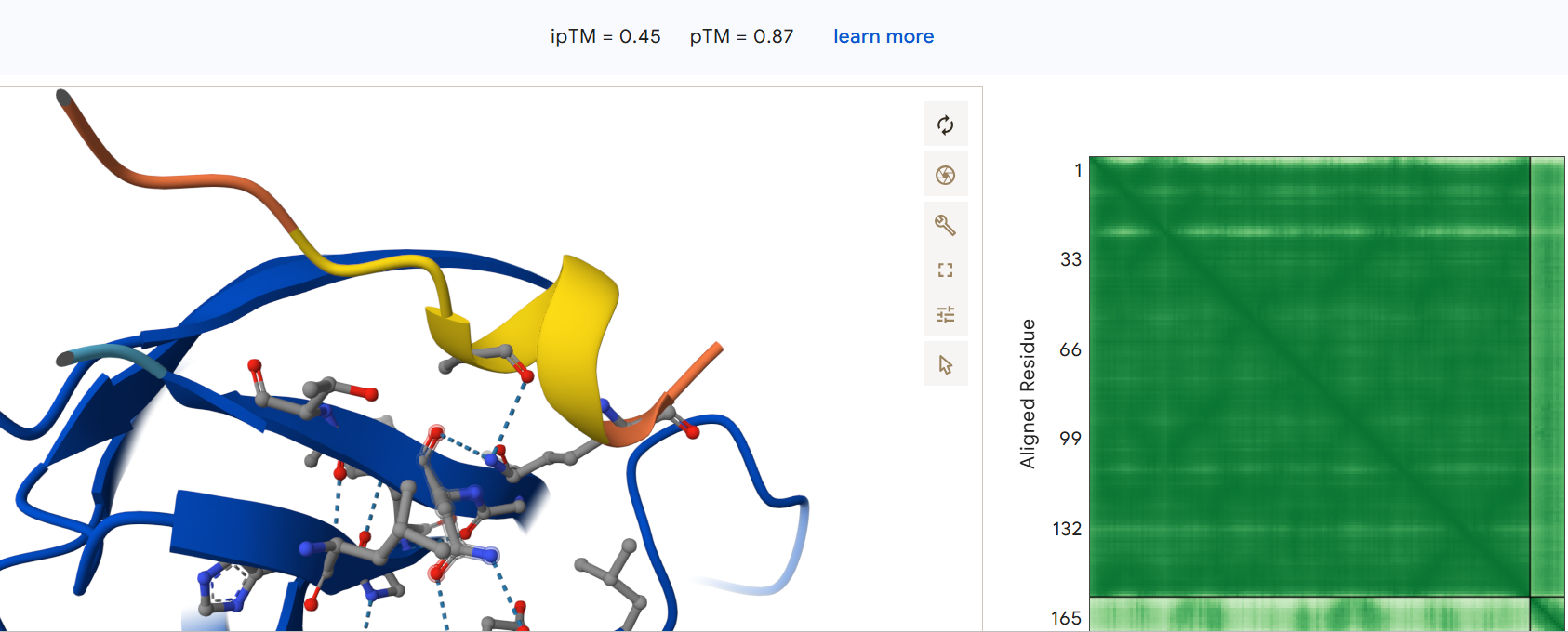

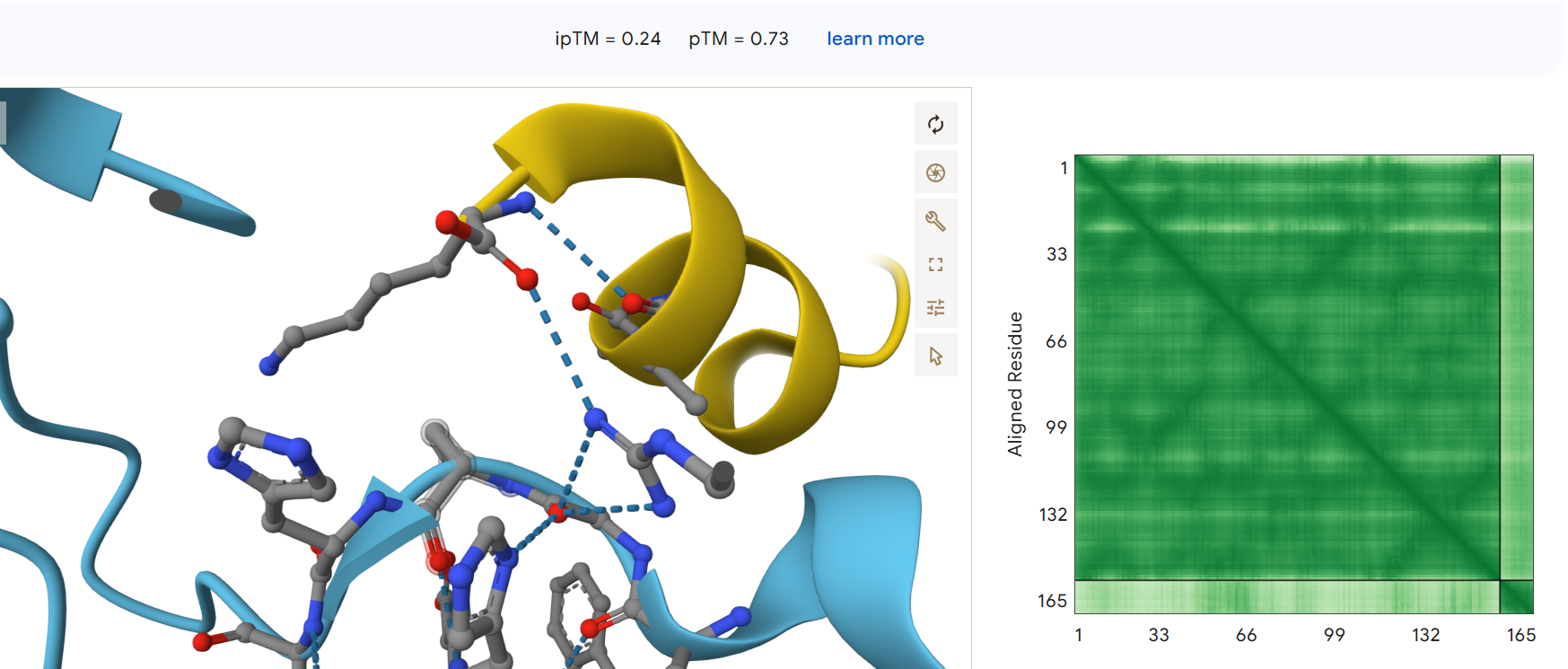
Third generated binder WLYPAVVAALKK with an ipTM = 0.24:
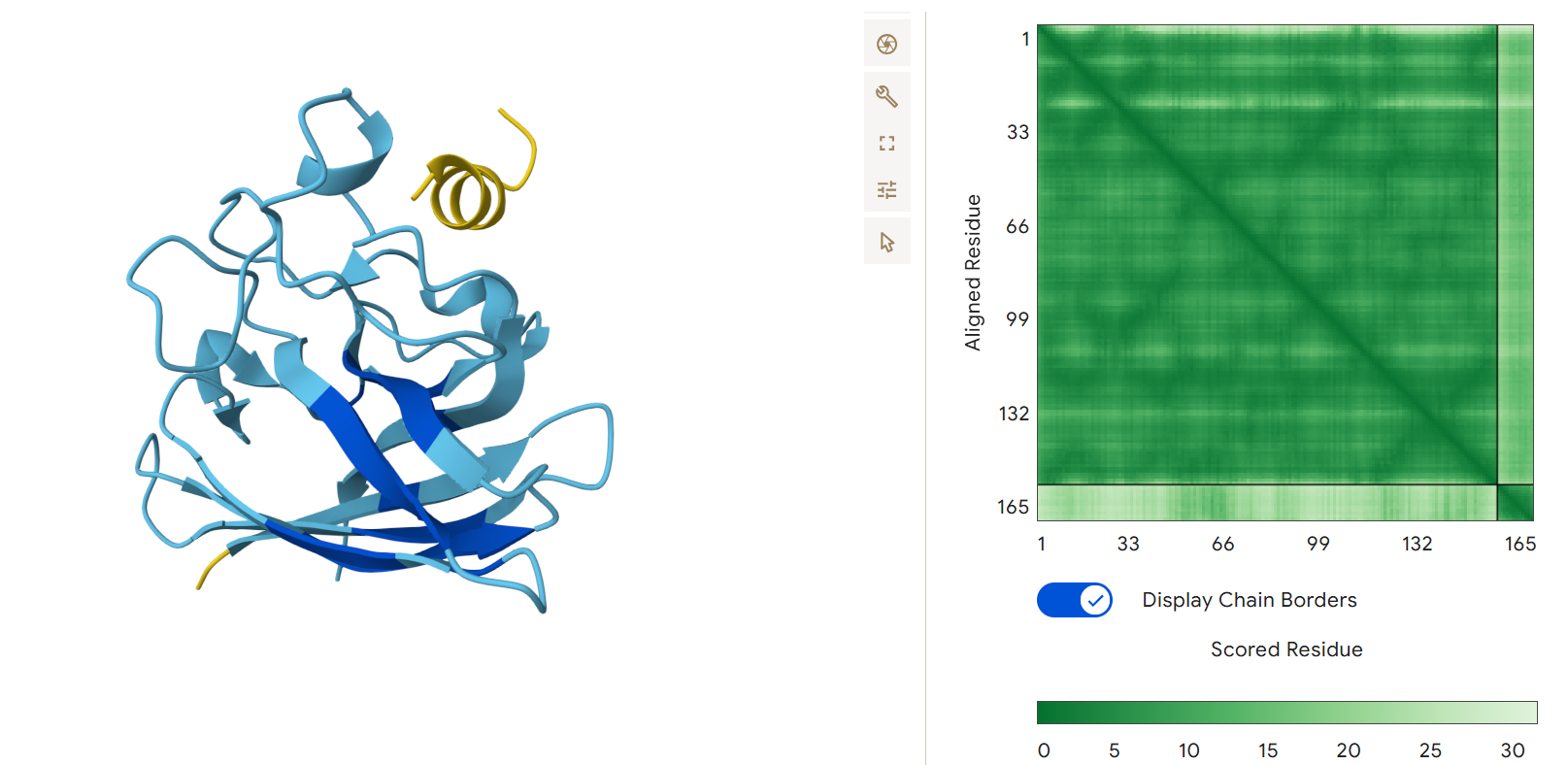


Fourth generated binder WRVPAAAVRHGX with an ipTM = 0.36:
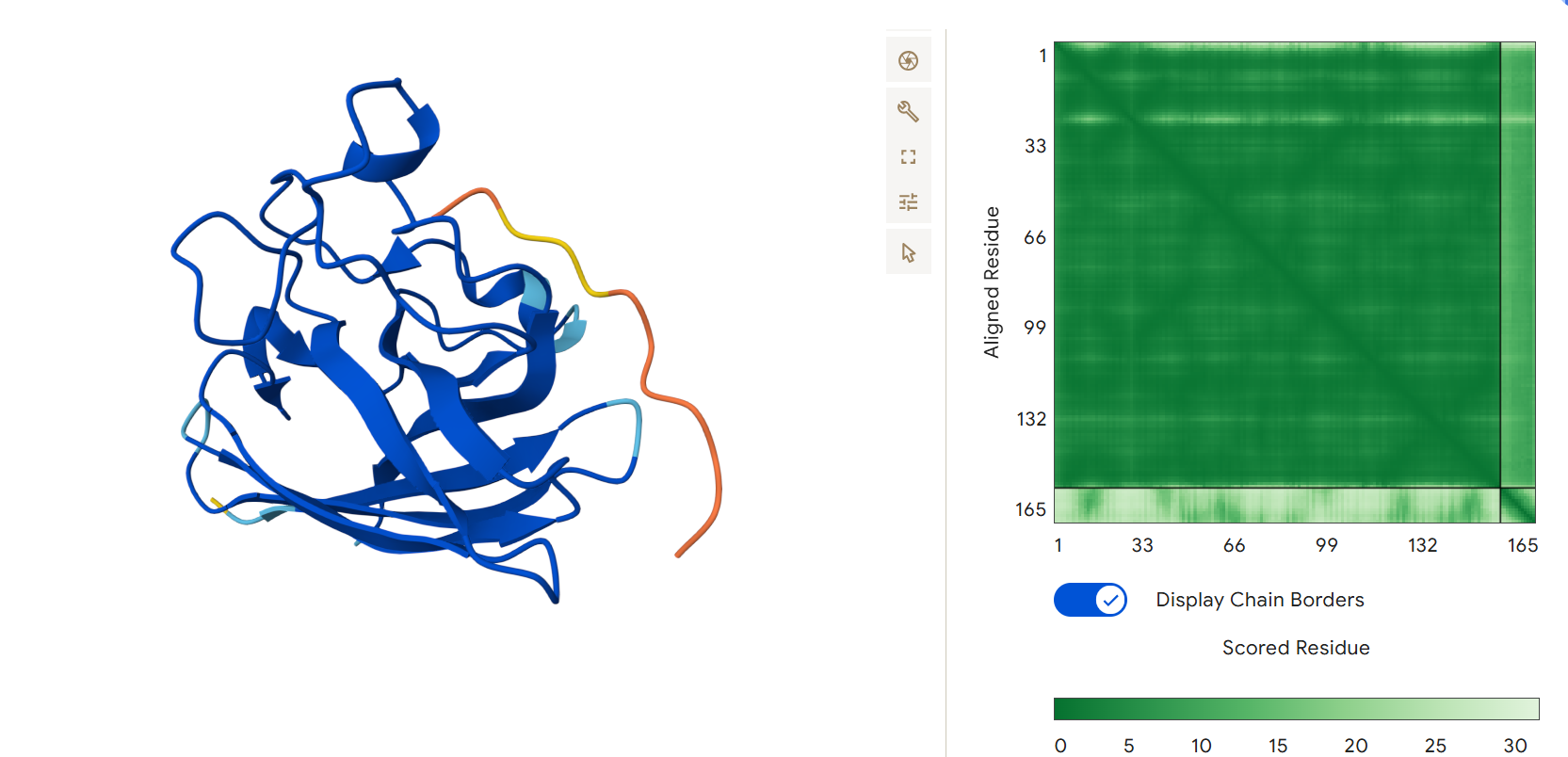

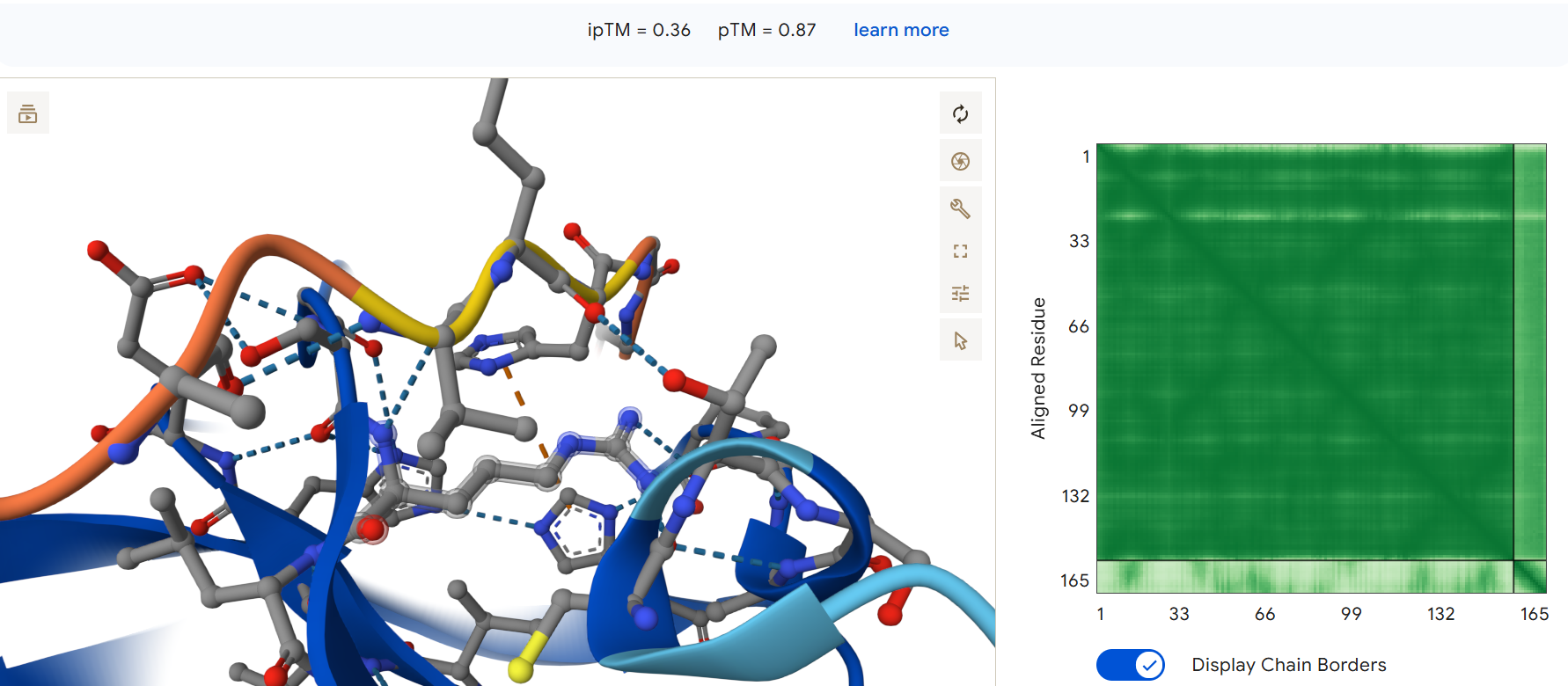

The second and fourth generated peptides achieved ipTM scores higher than that of the known binding peptide.
3. Evaluate properties of generated peptides in the PeptiVerse
I analyzed the results of the evaluation by PeptiVerse, considering predicted binding affinity, non-fouling predicted value, hemolysis, solubility, and net charge (pH 7).
FLYRWLPSRRGG: As the ground truth peptide, its properties are good, although its predicted binding affinity is considered “weak” by the model.
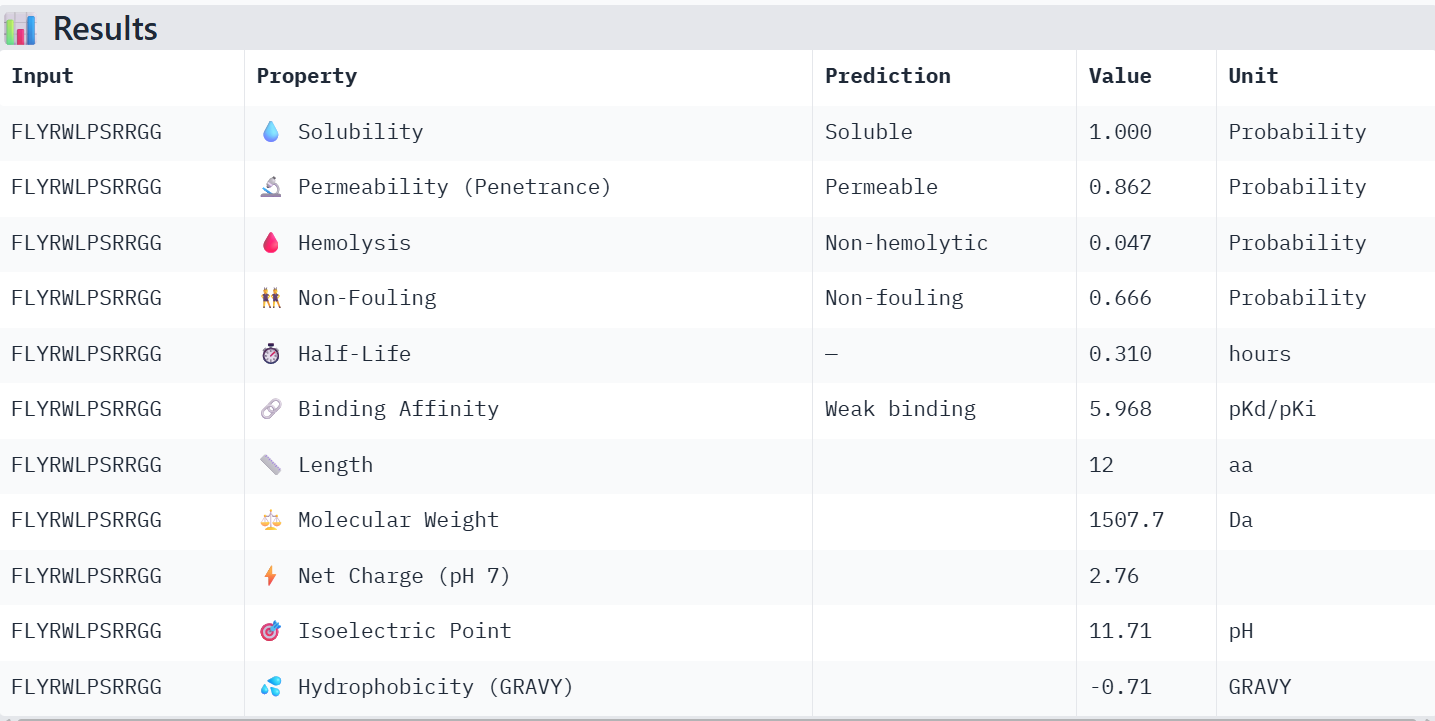

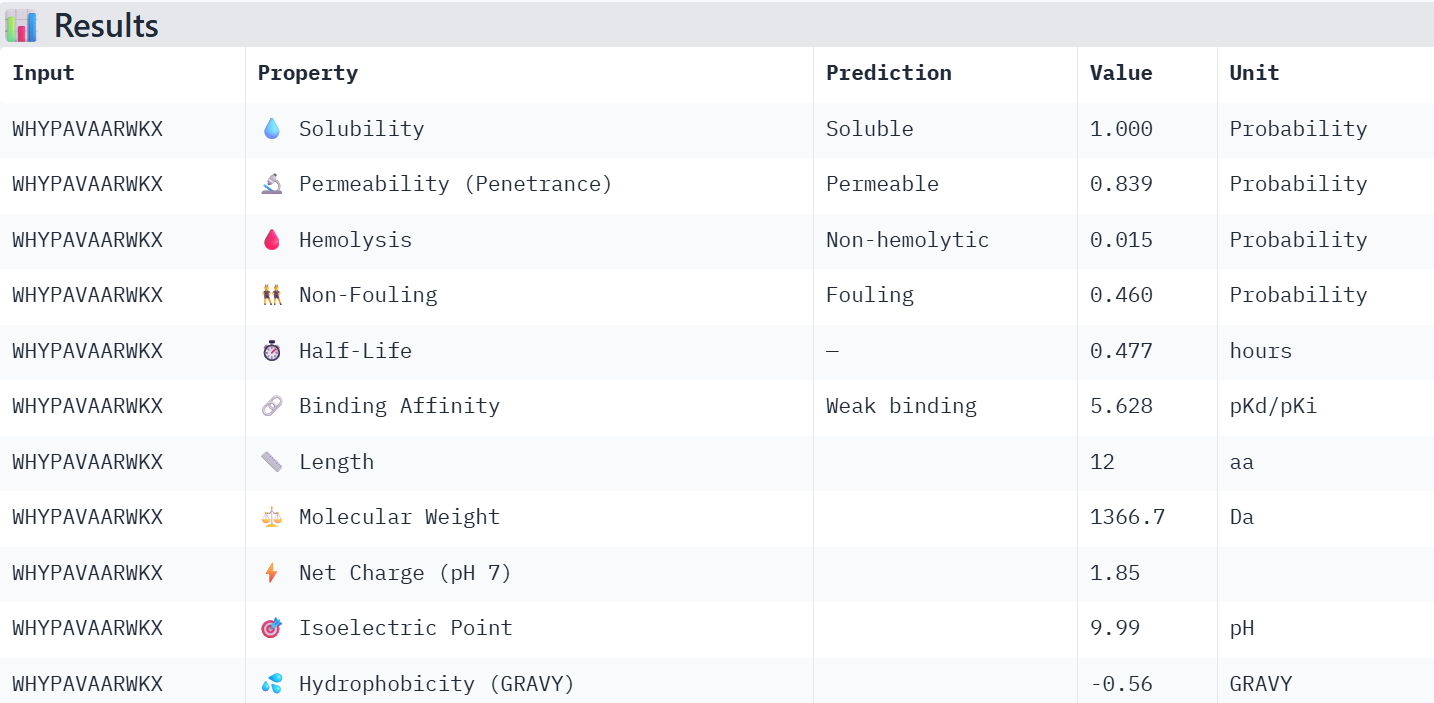
WHYPAVAARWKX: Has a relatively low predicted non-fouling value, and its binding affinity value diminished by 0.303 when changing X for G.

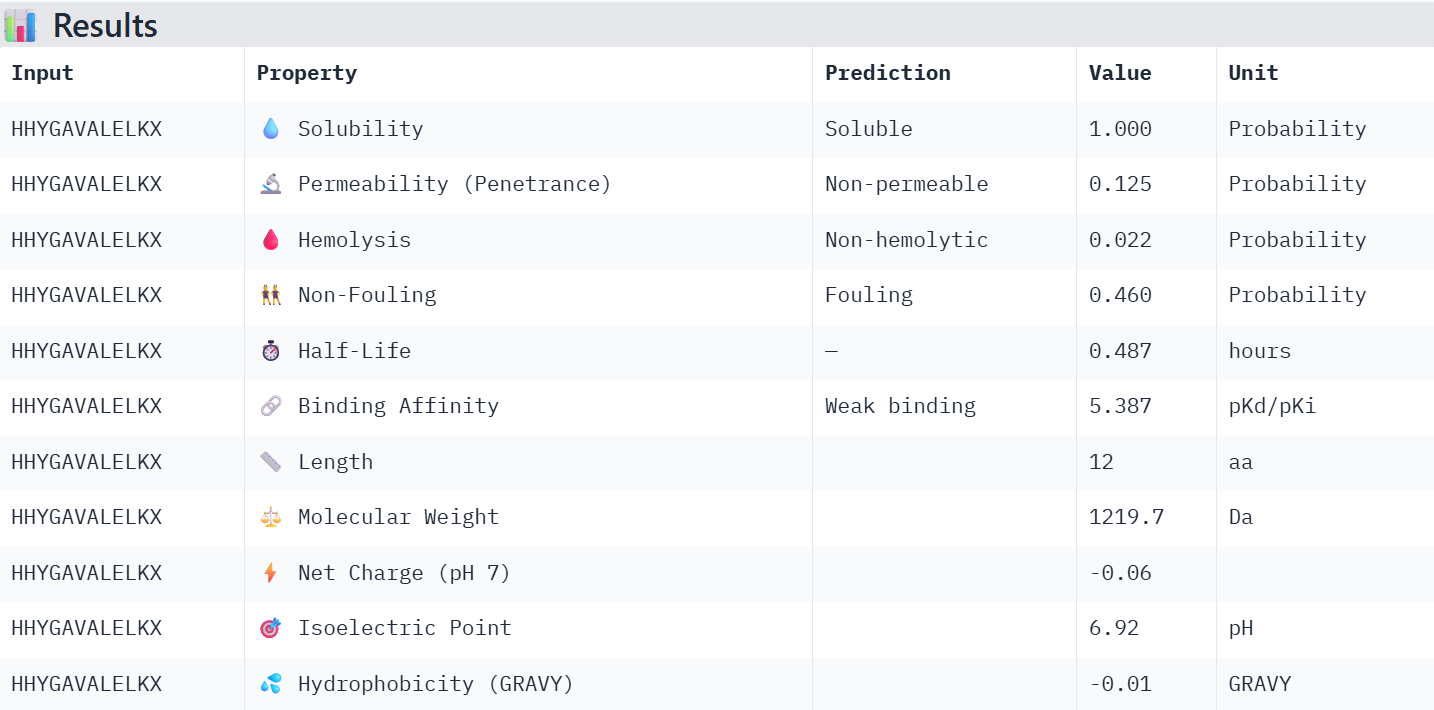
HHYGAVALELKX: When changing X for G, the binding affinity value diminished from 5.387 to 5.382, and its permeability value decreased from 0.125 to 0.41. Note that it wasn’t predicted to be permeable from the start.

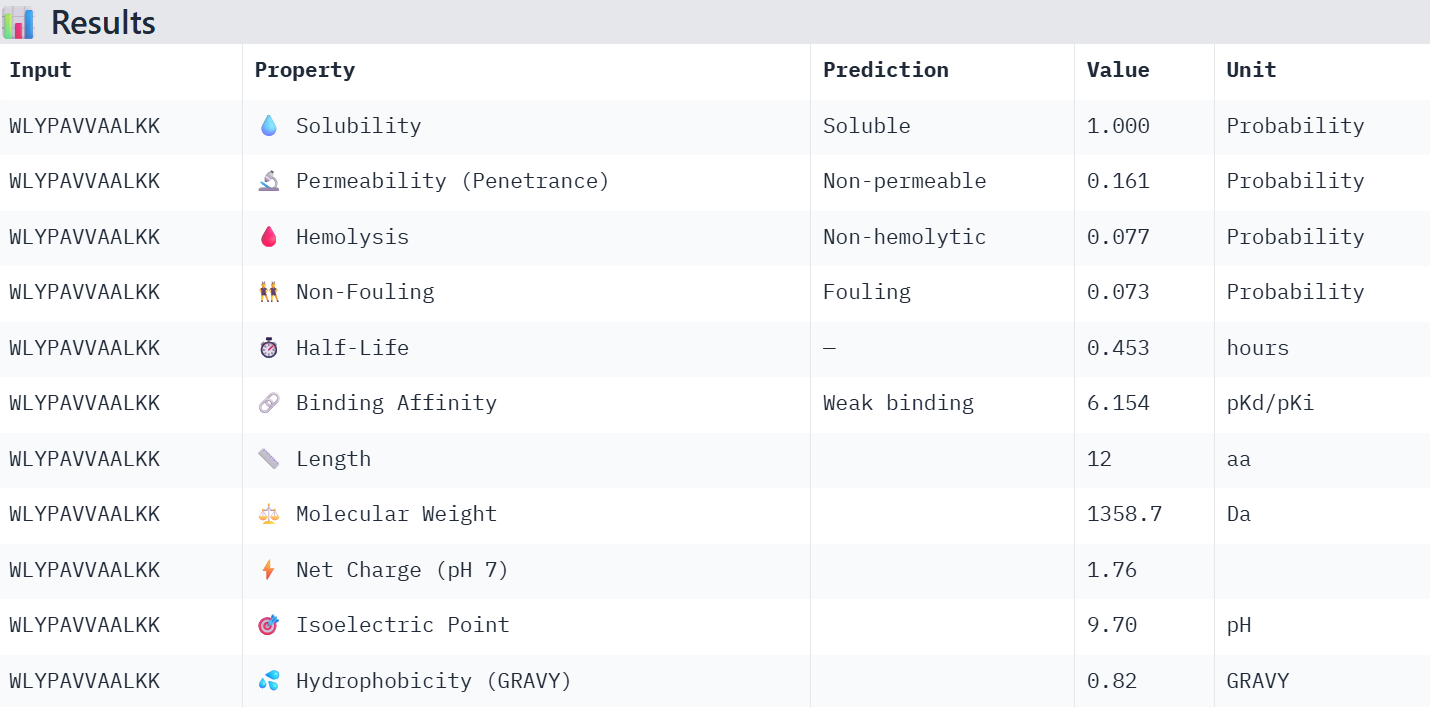
WLYPAVVAALKK: It achieved the highest binding affinity among the generated and known binding peptides; however, its fouling value is too high, and it is not permeable.

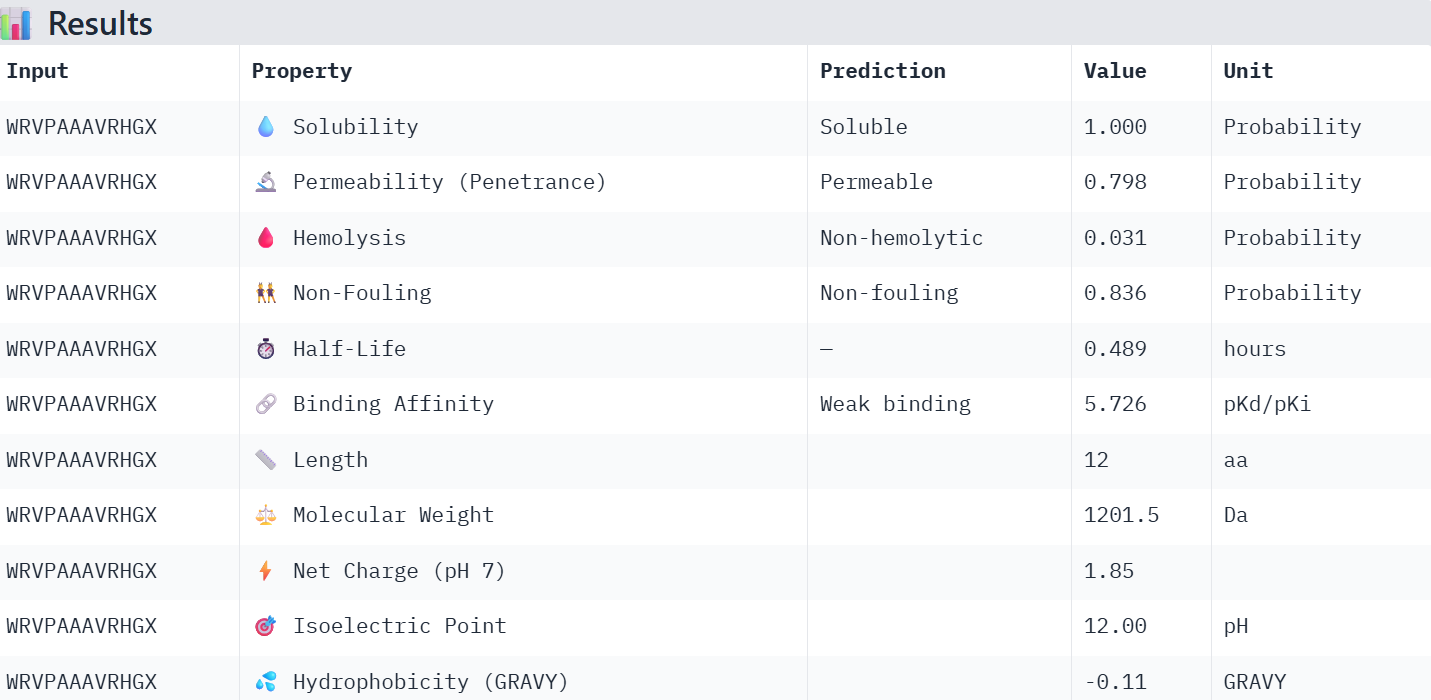
WRVPAAAVRHGX: Has good properties and a binding affinity near that of the known binding peptide.

In AlphaFold the second (HHYGAVALELKX) and fourth (WRVPAAAVRHGX) generated peptide had the higher ipTM. However, the PeptiVerse model considered the third generated peptide (WLYPAVVAALKK) and the known binding peptide (FLYRWLPSRRGG) as the ones with the the highest binding affinity.
4. Generate optimized peptides with moPPIt
I used the provided moPPit Colab, limiting the optimization parameters to work within a GPU T4 execution environment without running into computational power issues.
 I obtained a single sequence (as generating four sequences exceeded the environment’s capacity) and analyzed it.
I obtained a single sequence (as generating four sequences exceeded the environment’s capacity) and analyzed it.

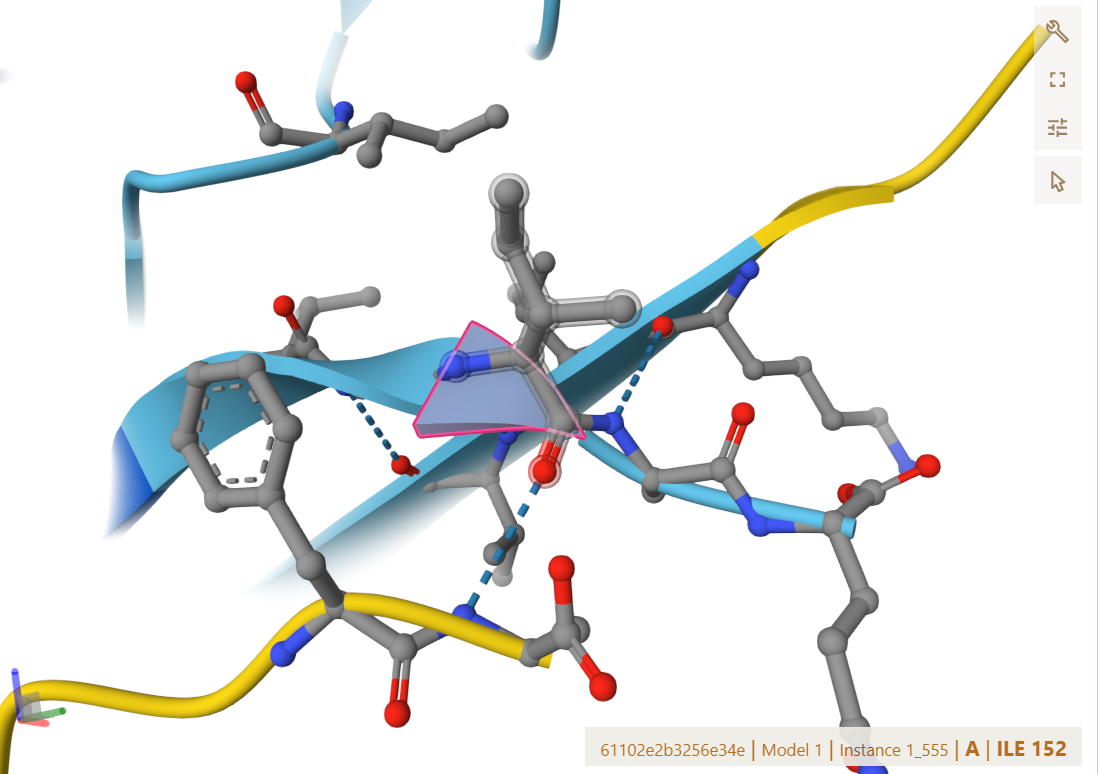
The AlphaFold 3 predicted ipTM of 0.34 was not considerably different from the non-optimized binders; however, the binding location changed significantly, appearing near the terminus this time.


The predicted structure shows that the peptide localizes near the C-terminus of the protein, within the dimer interface. Although it does not bind directly to the N-terminus, it interacts with it through the C-terminus beta-strand.

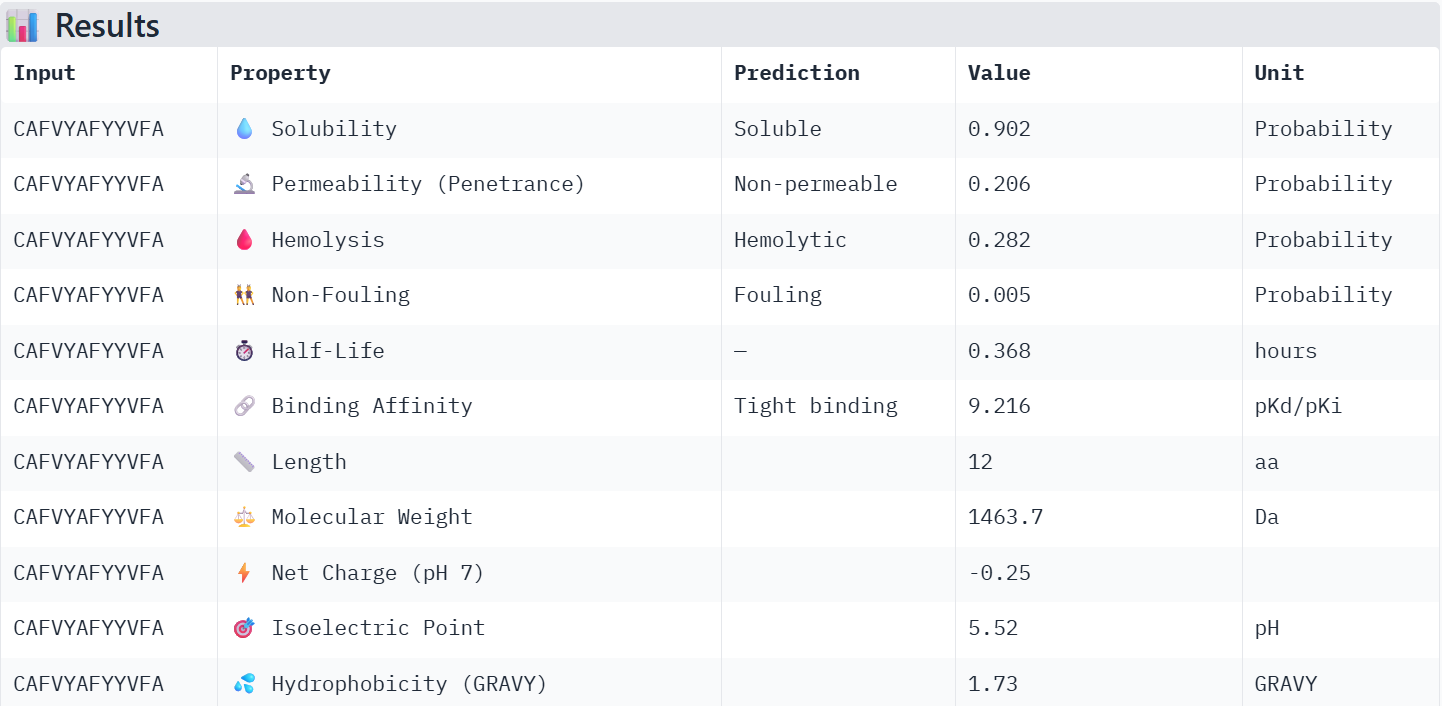
The PeptiVerse model predicted a tight binding affinity, but also a higher foulingness.

Bibliography:
- Saeed, M., Yang, Y., Deng, H.-X., Hung, W.-Y., Siddique, N., Dellefave, L., Gellera, C., Andersen, P. M., & Siddique, T. (2009). Age and founder effect of SOD1 A4V mutation causing ALS. Neurology, 72(19), 1634–1639. https://doi.org/10.1212/01.wnl.0000343509.76828.2a