Week 6 HW: Genetic Cirucuits - Part 1
Questions about the "The Chromophore Color Cloning Quest" lab protocol
Building constructs using the Kernel interface
Questions about the "The Chromophore Color Cloning Quest" lab protocol
Building constructs using the Kernel interface
While the principal components in every standard PCR reaction are a polymerase (like Taq), dNTPs, nuclease-free water, primers, buffer, and the DNA template to amplify, a Master Mix typically only contains the polymerase, dNTPs, and an optimized buffer (which includes Mg2+ as polymerases require magnesium to function). In the specific case of the Phusion High-Fidelity PCR Master Mix, we must note that the polymerase used is not Taq, but Phusion—a high-fidelity enzyme with 3’ to 5’ exonuclease (proofreading) activity. Its purpose is to amplify DNA with extreme accuracy and speed, preventing mutations during the assembly of sensitive genetic circuits.
First, the primer melting temperature (Tm). The Tm is the temperature at which 50% of the primer is bound to its complementary sequence, and the annealing temperature is typically set a few degrees below this. To calculate this melting point, we need to consider the length of the primer and the G-C content of the DNA template; G-C base pairs are bound by 3 hydrogen bonds each, so more heat is needed to separate them compared to A-T pairs. Second, the buffer conditions (specifically salt and magnesium concentrations) are a factor, as they add stability to the DNA double-stranded form by shielding the negative charges of the phosphate backbone, hence increasing the Tm and the required annealing temperature.
Protocols: PCR relies on thermal cycling (denaturation, annealing, and extension) using oligonucleotide primers and a thermostable DNA polymerase to synthesize new copies of a specific DNA fragment. In contrast, a restriction enzyme digest is an isothermal incubation (often at 37°C) where an endonuclease cuts pre-existing, double-stranded DNA at specific recognition sites, followed by heat inactivation of the enzyme.
Use Preference: PCR is preferable when you need to amplify a highly specific region from a complex template, when your starting DNA concentration is very low, or when you need to add custom overhangs (like homology arms) to the ends of your sequence. Restriction enzyme digestion is preferable for diagnostic checks (verifying the size of a cloned insert), when preparing plasmids for traditional ligation, or when you must completely avoid the risk of polymerase-induced mutations in a sequence that is already cloned.
First, in the PCR procedure, you want to end with fragments that have overlapping terminal bases (typically 15-40 base pairs) complementary to those in the backbone and the ones in the adjacent fragments, in order for them to be assembled by homology. Second, you can make sure the processed fragments are correct by doing gel electrophoresis and checking if the DNA is degraded or not by seeing if the results are consistent with the expected size. Finally, you can perform a gel extraction to purify these specific fragments from the template DNA and non-specific PCR products, ensuring a clean reaction.
Essentially, it enters through stress-induced pores in the plasmalema. It could be physical or chemical stress. This process is facilitated by cations (normally calcium cations from calcium chloride (CaCl2). These cations act as a bridge to shield and neutralize the electrostatic repulsion between the negatively charged phosphate backbone of the plasmid DNA and the negatively charged lipopolysaccharides on the bacterial plasmalema. Once the DNA is resting on the membrane, the stress pore induction (often done via heat-shock)creates a thermal imbalance that sweeps the DNA into the bacterial cell, allowing it to reach the cytoplasm. Other physical method could be electroporation, which generates pores with high electrical voltage.
There are plenty of methods, like Modular Cloning or Circular Polymerase Extentional cloning, etc. But the majority of them are based on Gibson assembly or Golden Gate Assembly. so I decided to explain the Golden Gate Assembly:
This technique leverages the capacity of specific endonucleases, called Type IIS restriction enzymes, to recognize asymmetric DNA sequences and cleave outside of this recognition site. By cutting typically 1 to 20 base pairs away, these enzymes generate customizable overhangs. The orientation of the recognition sites dictates the assembly mechanics: for the inserts, the sites are placed facing inward so the cut occurs between them, whereas in the destination vector, the sites must be outward-facing to flank the region being excised. The goal is to achieve one-pot assembly of multiple DNA fragments by designing these overhangs to be complementary (which can be generated via PCR) and using T4 DNA ligase to seal the construct. This simultaneous digestion and ligation is achievable because the restriction recognition sites are entirely removed during cleavage. Consequently, the assembled fragments and the final backbone become resistant to further digestion, driving the reaction forward seamlessly.
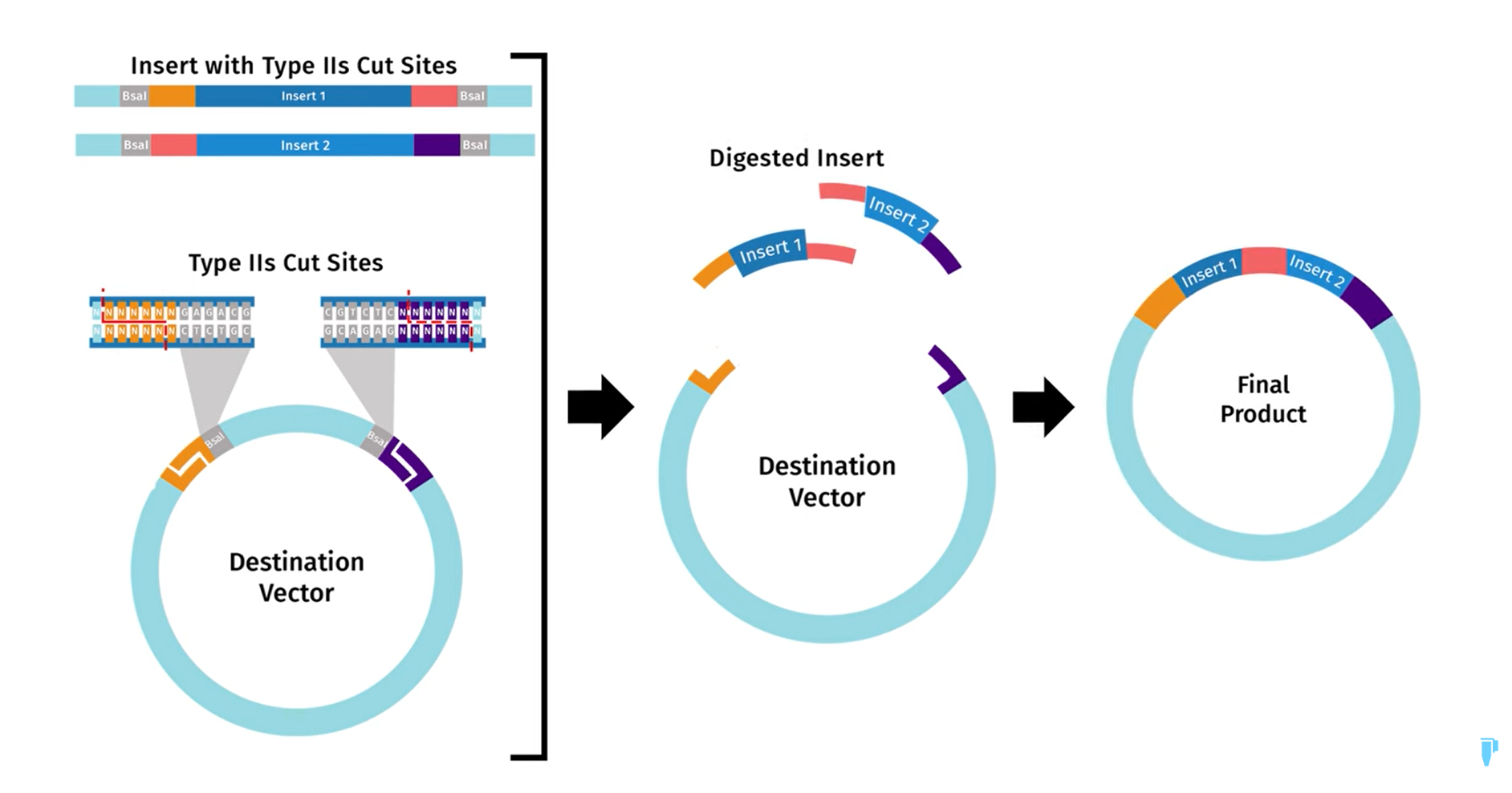
 Diagram taken from SnapGene: https://youtu.be/NzQdLQ44I7w?si=UbRcM9ca7adXYMNH
Diagram taken from SnapGene: https://youtu.be/NzQdLQ44I7w?si=UbRcM9ca7adXYMNH
Model this assembly method with Benchling: