Week 2 HW: DNA Read, Write, & Edit
Part 1: Benchling & In-silico Gel Art
I design two gels:
Gel A: ‘ART’
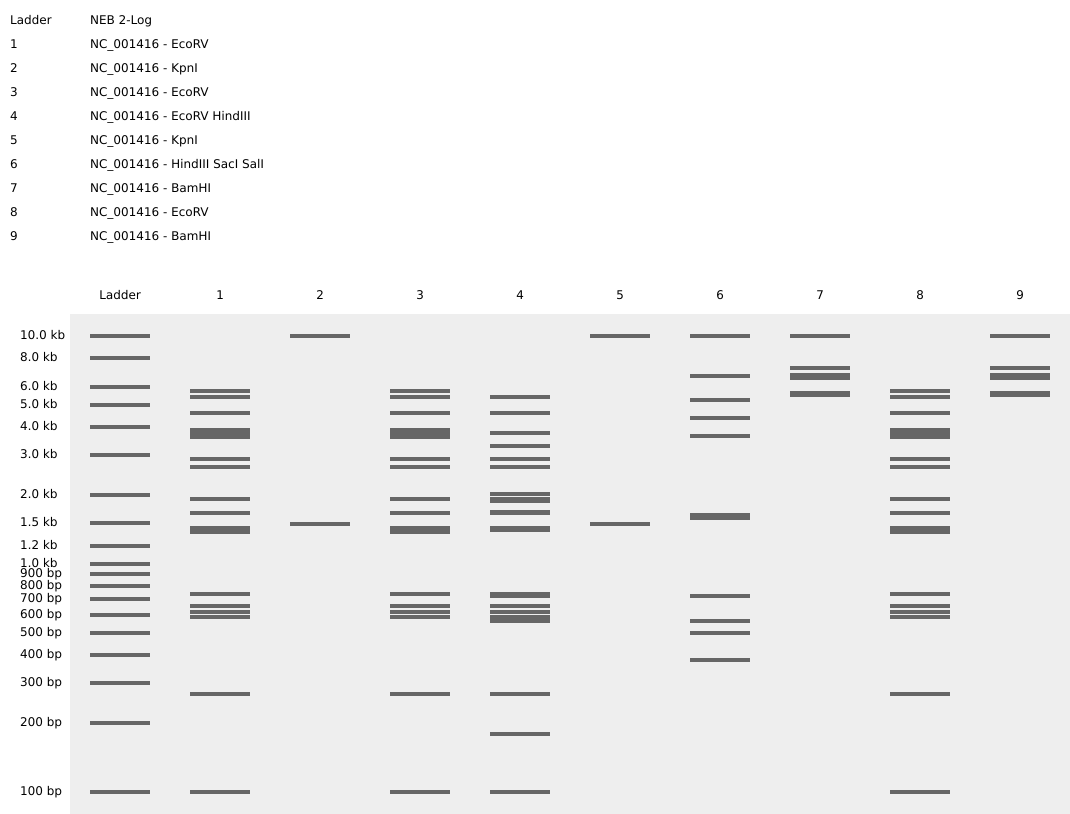

Gel B: ‘ATP’
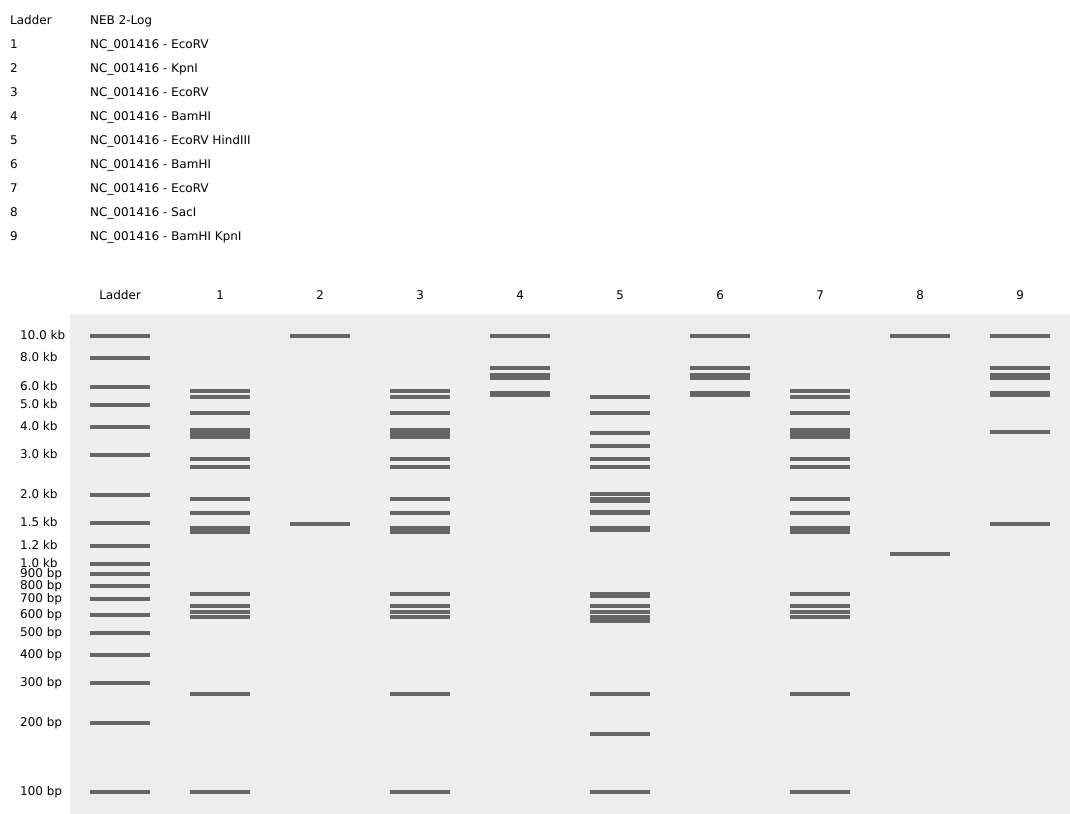


Part 3: DNA Design Challenge
3.1. Choose a protein - TcHMGB
For this homework, I chose the protein HMGB from Trypanosoma cruzi (TcHMGB), the parasite that causes Chagas disease. TcHMGB was characterized by the research group in which I carried out my undergraduate thesis. During this work, I constructed a DM28c cell line expressing TcHMGB fused to the biotin ligase TurboID, with the objective of performing a proximity labeling–based interactomics assay.
TcHMGB belongs to the HMG-B family, which plays a key role in chromatin organization and gene regulation in eukaryotic organisms. TcHMGB has been shown to modify chromatin structure, making it more accessible to the replication, transcription, repair, and recombination machinery. Additionally, TcHMGB is believed to be essential for T. cruzi survival and is therefore considered a potential therapeutic target.
- Protein sequence
Found in UniProt
- tr|Q4D714|Q4D714_TRYCC High mobility group protein, putative OS=Trypanosoma cruzi (strain CL Brener) OX=353153 GN=Tc00.1047053504431.64 PE=4 SV=1
MSTELKSGPLPADVEEVIANIMREEGVNFLTSKILRLRLEARYRMEFTSHKAAIEGIITKLMQLPEFKKQLENAVKEEKAASSIGGKKKKRSASAAADERNAKVNKKEKKPDDYPKAALSPYILFGNDHRDKVKEQNPEMKNTEILQSLGKMWAEASDAVKEKYKKLAEDDKKRFDRELSEYKKSGGTEYKRGGGKVKAKDENAPKRSMSAYFFFVSDFRKKHPDLSVTETSKAAGAAWKELSDEMKKPYEAMAQKDKERYQREMAARAS
3.2. Reverse Translate: Protein (amino acid) sequence to DNA (nucleotide) sequence.
For the reverse trnaslate of the protein TcHMGB, i used the EMB-EBI’s tool ‘EMBOSS BACKTRNASEQ’ setting T. cruzi codon table.
TcHMGB reverse trnaslated dna sequence
ATG AGC ACG GAG CTG AAG AGC GGC CCG CTG CCG GCG GAC GTG GAG GAG GTG ATT GCG AAC ATT ATG CGC GAG GAG GGC GTG AAC TTT CTG ACG AGC AAG ATT CTG CGC CTG CGC CTG GAG GCG CGC TAC CGC ATG GAG TTT ACG AGC CAC AAG GCG GCG ATT GAG GGC ATT ATT ACG AAG CTG ATG CAG CTG CCG GAG TTT AAG AAG CAG CTG GAG AAC GCG GTG AAG GAG GAG AAG GCG GCG AGC AGC ATT GGC GGC AAG AAG AAG AAG CGC AGC GCG AGC GCG GCG GCG GAC GAG CGC AAC GCG AAG GTG AAC AAG AAG GAG AAG AAG CCG GAC GAC TAC CCG AAG GCG GCG CTG AGC CCG TAC ATT CTG TTT GGC AAC GAC CAC CGC GAC AAG GTG AAG GAG CAG AAC CCG GAG ATG AAG AAC ACG GAG ATT CTG CAG AGC CTG GGC AAG ATG TGG GCG GAG GCG AGC GAC GCG GTG AAG GAG AAG TAC AAG AAG CTG GCG GAG GAC GAC AAG AAG CGC TTT GAC CGC GAG CTG AGC GAG TAC AAG AAG AGC GGC GGC ACG GAG TAC AAG CGC GGC GGC GGC AAG GTG AAG GCG AAG GAC GAG AAC GCG CCG AAG CGC AGC ATG AGC GCG TAC TTT TTT TTT GTG AGC GAC TTT CGC AAG AAG CAC CCG GAC CTG AGC GTG ACG GAG ACG AGC AAG GCG GCG GGC GCG GCG TGG AAG GAG CTG AGC GAC GAG ATG AAG AAG CCG TAC GAG GCG ATG GCG CAG AAG GAC AAG GAG CGC TAC CAG CGC GAG ATG GCG GCG CGC GCG
TcHMGB sequence from TryTrypDB
- TcCLB.507951.114
ATG TCC ACT GAA CTA AAG TCA GGT CCT CTT CCG GCA GAC GTC GAA GAG GTC ATC GCG AAT ATC ATG CGC GAA GAG GGT GTG AGC TTT CTC ACA AGT AAA ATT TTG CGT CTG CGC CTG GAG GCG AGG TAC CGC ATG GAG TTC ACC TCG CAC AAG GCT GCC ATA GAG GGC ATC ATT ACG AAG CTA ATG CAG CTT CCT GAG TTT AAG AAA CAG CTG GAG AAT GCT GTC AAG GAG GAG AAG GCG GCA AGC AGT ATC GGT GGT AAA AAG AAG AAG CGC AGC GCC AGT GCT GCG GCT GAC GAG AGG AAT GCG AAG GTG AAC AAA AAG GAG AAG AAG CCA GAT GAT TAC CCA AAG GCG GCG CTC TCG CCG TAC ATT CTC TTT GGG AAT GAC CAC CGT GAT AAA GTT AAG GAA CAG AAC CCG GGA ATG AAA AAT ACG GAG ATC TTG CAA AGT TTG GGC AAA ATG TGG GCC GAA GCC TCG GAT GCG GTC AAA GAA AAG TAC AAG AAA CTC GCA GAA GAT GAC AAG AAA AGA TTC GAT CGT GAA CTC AGC GAA TAT AAA AAG AGT GGT GGG ACT GAA TAC AAG CGC GGT GGC GGA AAG GTA AAG GCC AAG GAT GAA AAT GCC CCA AAG AGG TCC ATG TCT GCC TAC TTC TTC TTT GTG AGC GAT TTC CGG AAG AAG CAT CCC GAC CTC AGC GTC ACG GAG ACC TCC AAG GCA GCC GGC GCT GCG TGG AAA GCG CTC TCT GAT GAT ATG AAA AAG CCA TAC GAG GCC ATG GCA CAG AAG GAC AAG GAG CGG TAT CAG AGA GAG ATG GCT GCA AGG GCG AGC TAG
3.3. Codon optimization
Codon usage differs between organisms; that is, some organisms preferentially use codons that are rare in others. As a consequence, the translation efficiency of a T. cruzi gene expressed in yeast may be limited by the low availability of the corresponding tRNAs. Therefore, codon optimization is required to improve the efficiency of expression of the protein of interest.
tchmgb encodes a protein from Trypanosoma cruzi, a unicellular eukaryotic organism. Therefore, Saccharomyces cerevisiae was selected as the expression host because it provides an eukaryotic cellular context, including post-translational modification and protein-folding machinery, which is absent in prokaryotic systems such as Escherichia coli.
- tcHMGB was codon-optimized for expression in Saccharomyces cerevisiae using the IDT Codon Optimization Tool (Integrated DNA Technologies; https://www.idtdna.com/CodonOpt)
ATG TCA ACT GAA CTG AAG AGT GGC CCA TTG CCT GCA GAC GTA GAA GAG GTC ATA GCT AAT ATT ATG AGG GAA GAA GGA GTG AAT TTT TTG ACT TCT AAG ATT CTT AGA CTT AGA CTA GAA GCA AGA TAC AGA ATG GAA TTT ACT TCA CAT AAA GCC GCT ATT GAA GGC ATC ATT ACA AAA TTA ATG CAG TTG CCT GAA TTT AAG AAG CAA TTA GAG AAC GCA GTA AAA GAA GAG AAG GCA GCG TCA TCT ATT GGA GGT AAA AAG AAA AAG AGA TCA GCT TCT GCG GCT GCA GAC GAA AGA AAT GCT AAG GTC AAC AAG AAG GAA AAA AAA CCT GAC GAT TAT CCA AAG GCA GCT TTG AGC CCA TAT ATT TTA TTC GGT AAT GAC CAT AGA GAT AAG GTG AAG GAA CAA AAT CCA GAA ATG AAA AAT ACT GAG ATT TTG CAA TCA TTG GGA AAA ATG TGG GCC GAG GCG TCT GAC GCA GTT AAA GAG AAA TAC AAG AAA TTA GCC GAA GAT GAT AAG AAA CGT TTC GAT AGA GAA TTA TCT GAA TAC AAA AAA AGT GGT GGT ACT GAA TAT AAA CGT GGA GGT GGC AAG GTT AAA GCT AAG GAT GAG AAC GCT CCT AAG CGT AGT ATG TCT GCA TAT TTT TTT TTC GTG AGT GAT TTC AGA AAA AAG CAT CCA GAC TTA AGT GTT ACA GAA ACC TCT AAA GCC GCC GGT GCT GCT TGG AAA GAA CTA TCC GAT GAG ATG AAA AAG CCA TAC GAA GCT ATG GCA CAG AAG GAT AAA GAA AGA TAT CAA AGA GAA ATG GCT GCT AGG GCG
3.4. You have a sequence! Now what?
To produce this protein, it is first necessary to obtain a plasmid that enables expression of the protein of interest in the selected expression system, either cell-dependent or cell-free. For plasmid construction, there are two main options: (1) cloning the coding sequence into an existing expression vector, or (2) ordering a synthetic plasmid that includes regulatory elements selected according to the chosen expression system and experimental goals—such as a constitutive or inducible promoter, ribosome binding site, coding sequence, affinity tag, and transcription termination sequence—thereby improving expression efficiency and control.
Once the plasmid is obtained, it is introduced into the chosen expression system. In this case, Saccharomyces cerevisiae was selected as the host organism. The plasmid can be delivered into yeast cells by electroporation, followed by selection of successfully transformed cells. After selection, protein expression is induced under the appropriate conditions, allowing transcription of the gene and subsequent translation into the protein of interest.
Part 4: Prepare a Twist DNA Synthesis Order
4.2 Build Your DNA Insert Sequence
- Promotor: yeast GAL1 promoter ( BBa_J63006 - https://parts.igem.org/wiki/index.php?title=Part:BBa_J63006): GAGGAAACTAGACCCGCCGCCACCATGGAG
- RBS: designed yeast Kozak sequence (BBa_J63003 - https://parts.igem.org/wiki/index.php?title=Part:BBa_J63003): CCCGCCGCCACCATGGAG
- Coding sequence: tchmgb codon optimized for yeast (without start and stop codons)
TCA ACT GAA CTG AAG AGT GGC CCA TTG CCT GCA GAC GTA GAA GAG GTC ATA GCT AAT ATT ATG AGG GAA GAA GGA GTG AAT TTT TTG ACT TCT AAG ATT CTT AGA CTT AGA CTA GAA GCA AGA TAC AGA ATG GAA TTT ACT TCA CAT AAA GCC GCT ATT GAA GGC ATC ATT ACA AAA TTA ATG CAG TTG CCT GAA TTT AAG AAG CAA TTA GAG AAC GCA GTA AAA GAA GAG AAG GCA GCG TCA TCT ATT GGA GGT AAA AAG AAA AAG AGA TCA GCT TCT GCG GCT GCA GAC GAA AGA AAT GCT AAG GTC AAC AAG AAG GAA AAA AAA CCT GAC GAT TAT CCA AAG GCA GCT TTG AGC CCA TAT ATT TTA TTC GGT AAT GAC CAT AGA GAT AAG GTG AAG GAA CAA AAT CCA GAA ATG AAA AAT ACT GAG ATT TTG CAA TCA TTG GGA AAA ATG TGG GCC GAG GCG TCT GAC GCA GTT AAA GAG AAA TAC AAG AAA TTA GCC GAA GAT GAT AAG AAA CGT TTC GAT AGA GAA TTA TCT GAA TAC AAA AAA AGT GGT GGT ACT GAA TAT AAA CGT GGA GGT GGC AAG GTT AAA GCT AAG GAT GAG AAC GCT CCT AAG CGT AGT ATG TCT GCA TAT TTT TTT TTC GTG AGT GAT TTC AGA AAA AAG CAT CCA GAC TTA AGT GTT ACA GAA ACC TCT AAA GCC GCC GGT GCT GCT TGG AAA GAA CTA TCC GAT GAG ATG AAA AAG CCA TAC GAA GCT ATG GCA CAG AAG GAT AAA GAA AGA TAT CAA AGA GAA ATG GCT GCT AGG
- 7x His Tag: CATCACCATCACCATCATCAC
- STOP: TAA
- Terminatior: Saccharomyces cerevisiae CYT1 (BBa_K2637017 - https://parts.igem.org/wiki/index.php?title=Part:BBa_K2637017): TCGCCCGTACATTCATGTAATTAGTTATGTCACGCTTACATTCACGCCCTCCTCCCACATCCGCTCTAACCGAAAAGGAAGGAGTTAGACAACCTGAAGTCTAGGTCCCTATTTATTTTTTTTAATAGTTATGTTAGTATTAAGAACGTTATTTATATTTCAAATTTTTCTTTTTTTTCTGTACAAACGCGTGTACGCATGTAACATTATACTGAAAACCTTGCTTGAGAAGGTTTTGGGACGCTCGAAGGCTTTAGATCT
Expression casset:
GAGGAAACTAGACCCGCCGCCACCATGGAGCCCGCCGCCACCATGGAGTCAACTGAACTGAAGAGTGGCCCATTGCCTGCAGACGTAGAAGAGGTCATAGCTAATATTATGAGGGAAGAAGGAGTGAATTTTTTGACTTCTAAGATTCTTAGACTTAGACTAGAAGCAAGATACAGAATGGAATTTACTTCACATAAAGCCGCTATTGAAGGCATCATTACAAAATTAATGCAGTTGCCTGAATTTAAGAAGCAATTAGAGAACGCAGTAAAAGAAGAGAAGGCAGCGTCATCTATTGGAGGTAAAAAGAAAAAGAGATCAGCTTCTGCGGCTGCAGACGAAAGAAATGCTAAGGTCAACAAGAAGGAAAAAAAACCTGACGATTATCCAAAGGCAGCTTTGAGCCCATATATTTTATTCGGTAATGACCATAGAGATAAGGTGAAGGAACAAAATCCAGAAATGAAAAATACTGAGATTTTGCAATCATTGGGAAAAATGTGGGCCGAGGCGTCTGACGCAGTTAAAGAGAAATACAAGAAATTAGCCGAAGATGATAAGAAACGTTTCGATAGAGAATTATCTGAATACAAAAAAAGTGGTGGTACTGAATATAAACGTGGAGGTGGCAAGGTTAAAGCTAAGGATGAGAACGCTCCTAAGCGTAGTATGTCTGCATATTTTTTTTTCGTGAGTGATTTCAGAAAAAAGCATCCAGACTTAAGTGTTACAGAAACCTCTAAAGCCGCCGGTGCTGCTTGGAAAGAACTATCCGATGAGATGAAAAAGCCATACGAAGCTATGGCACAGAAGGATAAAGAAAGATATCAAAGAGAAATGGCTGCTAGGCATCACCATCACCATCATCACTAATCGCCCGTACATTCATGTAATTAGTTATGTCACGCTTACATTCACGCCCTCCTCCCACATCCGCTCTAACCGAAAAGGAAGGAGTTAGACAACCTGAAGTCTAGGTCCCTATTTATTTTTTTTAATAGTTATGTTAGTATTAAGAACGTTATTTATATTTCAAATTTTTCTTTTTTTTCTGTACAAACGCGTGTACGCATGTAACATTATACTGAAAACCTTGCTTGAGAAGGTTTTGGGACGCTCGAAGGCTTTAGATCT
4.6. Choose Your Vector
Since I could not use Twist directly for this exercise, I downloaded the pTwist Amp High Copy cloning vector map and imported it into Benchling. I then inserted the previously designed TcHMGB expression cassette into this backbone using Benchling’s sequence editing tools, allowing visualization and validation of the complete circular plasmid.
Part 5: DNA Read/Write/Edit
5.1 DNA Read
(i) What DNA would you want to sequence (e.g., read) and why?
I would choose to sequence DNA from hospital wastewater.Hospital effluents contain a high diversity of bacteria exposed to strong antibiotic selective pressure, making them important reservoirs of antimicrobial resistance genes. By sequencing this DNA, it is possible to identify and monitor resistance genes and their potential spread into the environment.
This information is highly relevant for public health, as it can contribute to early detection of emerging resistance patterns and support strategies to limit the dissemination of antibiotic resistance.
(ii) In lecture, a variety of sequencing technologies were mentioned. What technology or technologies would you use to perform sequencing on your DNA and why? I would use Oxford Nanopore to obtain long reads that provide structural and genetic context, and Illumina to obtain short, high-quality reads that allow more reliable genome assembly and error correction.
- Is your method first-, second- or third-generation or other? How so? Illumina is a second-generation sequencing technology, as it relies on sequencing-by-synthesis and generates large numbers of short, highly accurate reads in parallel. Oxford Nanopore is considered a third-generation sequencing technology because it sequences single DNA molecules directly and produces long reads without requiring amplificatio.
- What is your input? How do you prepare your input (e.g. fragmentation, adapter ligation, PCR)? List the essential steps. My input would be total DNA extracted from hospital wastewater, which includes mixed microbial DNA from a metagenomic sample. Essentially, Illumina requires DNA fragmentation and amplification, while Oxford Nanopore can sequence long, minimally processed DNA. The essential steps are:
- High-quality DNA extraction
- Library preparation, including fragmentation (not necessary for Nanopore), amplification, and incorporation of adapters and indices
- 3.Sequencing
- What are the essential steps of your chosen sequencing technology, how does it decode the bases of your DNA sample (base calling)?
- Illumina decodes DNA by detecting fluorescent signals emitted when labeled nucleotides are incorporated during DNA synthesis. Each base is identified based on its specific fluorescence signal.
- Oxford Nanopore decodes DNA by measuring changes in ionic current as a DNA strand passes through a nanopore. Different nucleotide sequences cause characteristic current disruptions, which are translated into base calls using computational algorithm.
- What is the output of your chosen sequencing technology? Illumina sequencing results in short, high-quality sequencing reads, while Oxford Nanopore produces long reads of variable length. In both sequencing technologies, the output consists of FASTQ files containing all the reads, which can be processed using multiple platforms for downstream analysis
5.2 DNA Write
(i) What DNA would you want to synthesize (e.g., write) and why? These could be individual genes, clusters of genes or genetic circuits, whole genomes, and beyond. As described in class thus far, applications could range from therapeutics and drug discovery (e.g., mRNA vaccines and therapies) to novel biomaterials (e.g. structural proteins), to sensors (e.g., genetic circuits for sensing and responding to inflammation, environmental stimuli, etc.), to art (DNA origamis). If possible, include the specific genetic sequence(s) of what you would like to synthesize! You will have the opportunity to actually have Twist synthesize these DNA constructs! (ii) What technology or technologies would you use to perform this DNA synthesis and why?
- What are the essential steps of your chosen sequencing methods?
- What are the limitations of your sequencing method (if any) in terms of speed, accuracy, scalability?
5.3 DNA Edit
(i) What DNA would you want to edit and why? In class, George shared a variety of ways to edit the genes and genomes of humans and other organisms. Such DNA editing technologies have profound implications for human health, development, and even human longevity and human augmentation. DNA editing is also already commonly leveraged for flora and fauna, for example in nature conservation efforts, (animal/plant restoration, de-extinction), or in agriculture (e.g. plant breeding, nitrogen fixation). What kinds of edits might you want to make to DNA (e.g., human genomes and beyond) and why?
(ii) What technology or technologies would you use to perform these DNA edits and why?
- How does your technology of choice edit DNA? What are the essential steps?
- What preparation do you need to do (e.g. design steps) and what is the input (e.g. DNA template, enzymes, plasmids, primers, guides, cells) for the editing?
- What are the limitations of your editing methods (if any) in terms of efficiency or precision?