Week 3 HW: Lab automatation
Python Script for Opentrons Artwork
To create the Python file to run on an Opentrons liquid-handling robot, I used the scripts from the file downloaded from http://opentrons-art.rcdonovan.com/ as guide, to write the scripts unthe the ‘#YOUR CODE HERE’ line. I also edited the ‘well_color’ library to add some colors.
Google colab cell link: https://colab.research.google.com/drive/1Ke57DkO8O2jpCUDaYyK3krv4kUYFVgn8#scrollTo=pczDLwsq64mk&line=9&uniqifier=1
Simulation visualisation:
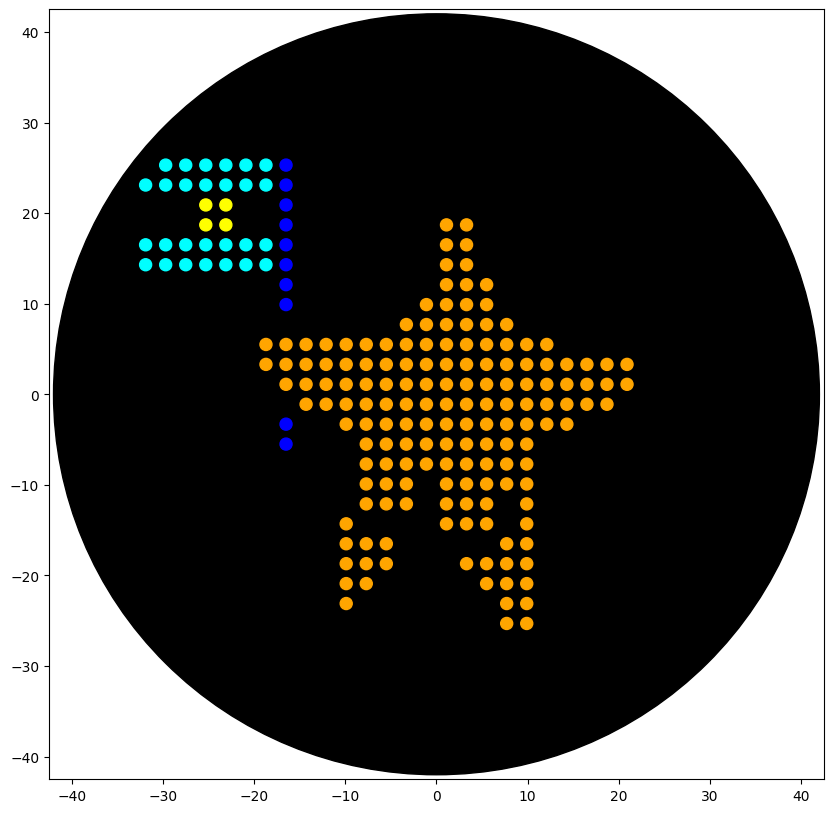

Post-Lab Questions
Descrbe 3 proyects
Proyect 1: Engineered Miniproteins to Disrupt and Prevent Bacterial Biofilm Formation
THE PROBLEM
Biofilms are structured microbial communities embedded in a self-produced extracellular polymeric substance (EPS) matrix composed of polysaccharides, proteins, extracellular DNA and lipids. They form on:
- Medical devices
- Tissues
- Industrial surfaces
- Environmental niches.
Clinically, biofilms are highly problematic:
EPS matrix limits antibiotic penetration and there is a reduced metabolic activity of inner-layer cells leading to Antibiotic tolerance
Quorum sensing (QS) coordinates: cell-density (dependent gene regulation), virulence and matrix biosynthesis.
In pathogens such as Pseudomonas aeruginosa, QS regulators like LasR orchestrate biofilm maturation and virulence programs.
As a consequence, biofilm-associated infections are often chronic, recurrent, and difficult to eradicate.
THE VISION
Instead of bactericidal pressure (which accelerates resistance evolution), this project proposes a molecular destabilization strategy using de novo engineered miniproteins.
These programmable miniproteins will:
- Bind and inhibit QS regulators
- Target transcription factors such as LasR-type proteins
- Block coordinated gene expression
- Prevent structured biofilm formation
- Interfere with EPS assembly
- Bind structural matrix proteins
- Disrupt polysaccharide–protein interactions
- Destabilize mature biofilms
- Be modular and retargetable
- Scaffold-based design
- Sequence redesign for new pathogens
- Compatible with automated high-throughput screening
The goal is to convert biofilms from a protected, resistant state into a destabilized, antibiotic-sensitive state.
AUTOMATIZATION
Automation will rely on an Opentrons OT-2 liquid-handling robot to execute the design–build–test cycle. Miniprotein variants will be assembled through automated cloning workflows using liquid-handling robotics, the the constructed plasmids will be transformed and cultured in multiwell plate formats and finally biofilm formation and inhibition assays will be performed using automated pipetting and plate-based quantification.
Automated Modules:
- Robotic cloning workflows (Golden Gate / Gibson / Gateway-type systems)
- Multiwell bacterial transformation
- Automated culture normalization
- Plate-based biofilm quantification (crystal violet assays)
- Reporter-based QS inhibition screens
WORK FLOW
%%{init: {'themeVariables': { 'fontSize': '15px' }}}%%
flowchart LR
A["<b>1. Computational Design.</b><br/>Use de novo protein design algorithms to design small, stable binders targeting QS regulators or matrix components"]
B["<b>2. Gene Synthesis and Expression</b><br/>DNA synthesis and cloning<br/>Expresion in bacteria or-cellfree systems for screening"]
C["<b>3. Functional Validation"]
D["<b>4. Delivery Strategy"</b><br/>A. Engineered probiotic chassis for local secretion.
B. Virus-like particles for targeted protein delivery.]
A --> B --> C --> DProyect 2: Biomarkers for Endometriosis Detection
(still developing this idea)
THE PROBLEM
Endometriosis is a chronic, estrogen-dependent inflammatory disease affecting ~10% of reproductive-age women and causing chronic pelvic pain and infertility. Despite its prevalence, diagnosis is delayed by 7–10 years due to social, clinical, and technical factors, including normalization of severe menstrual pain and the lack of validated non-invasive screening biomarkers. This delay contributes to disease progression, chronic pain sensitization, reduced fertility, increased surgical burden, and significant psychological distress — highlighting the need for early, accessible detection tools.
THE VISION
Develop a standardized, automated, non-invasive screening tool — such as a blood-based microRNA panel and/or a combined inflammatory profile — to enable earlier risk identification of endometriosis through circulating molecular biomarkers. The goal is not to provide a definitive diagnosis, but to develop a tool capable of indicating the probability of endometriosis. This would help identify women at higher risk, support earlier referral to specialists, reduce diagnostic delay, and ultimately improve long-term clinical outcomes.
FIRST APROACH
Circulating microRNAs (miRNAs) are among the most promising non-invasive biomarker candidates for endometriosis. Additionally, altered inflammatory markers such as IL-6, IL-8, and TNF-α further support the presence of a detectable circulating molecular signature. However, no single biomarker has demonstrated sufficient sensitivity and specificity for clinical implementation due to inter-study variability and lack of methodological standardization.
Therefore, the initial aim is to standardize and automate a reproducible biomarker panel, primarily based on previously reported circulating miRNAs and potentially complemented by selected inflammatory markers. By integrating these candidates into a controlled, automated laboratory workflow, the project seeks to reduce technical variability, improve reproducibility, and enable scalable early risk stratification for endometriosis.
Proyect 3: Paper-Based Biosensor for Early Detection of Plant Water Stress
THE PROBLEM
Water stress is one of the main causes of crop yield loss worldwide. Current monitoring strategies rely primarily on soil moisture sensors, satellite imaging, or visible plant symptoms. However:
- Soil moisture does not necessarily reflect plant physiological status.
- Visual symptoms appear after stress is already advanced.
- Molecular stress markers require laboratory-based analysis. There is currently no simple, field-deployable tool to detect early physiological stress directly from plant tissue.
THE VISION
Develop a portable, paper-based biosensor capable of detecting early molecular markers of water stress from leaf extracts. Instead of measuring environmental parameters, this system will assess the plant’s internal physiological response. The device will:
- Detect stress-associated biomarkers (e.g., ROS-related signals or osmotic stress markers).
- Produce a visible colorimetric output.
- Operate without laboratory equipment.
- Be deployable directly in agricultural settings.
Possible implementations:
Whole-cell biosensor:
- Engineered E. coli
- Stress-responsive promoter → colorimetric reporter (e.g., LacZ, chromoprotein)
Cell-free transcription-translation (TXTL) system
- Freeze-dried on paper
- Synthetic promoter responsive to biomarker
- Faster, safer, more stable in field conditions
Paper microfluidics advantages
- Capillary-driven flow (no pumps)
- Wax-printed channel definition
- Low cost
- Disposable
- Compatible with freeze-dried reagents
Readout could be:
- Visible color change
- Smartphone-quantifiable signal
AUTOMATIZATION
First sensor constructs will be designed and prototyped using automated cloning workflows on an Opentrons liquid-handling platform. Then variants will be assembled in parallel and expressed in bacterial or cell-free systems. Screening of biosensor performance will be performed in 96-well plate formats prior to integration into the paper-based device. Finally the experimental data will guide iterative redesign of sensing circuits.
Opentrons-Based Pipeline
- Automated Golden Gate / Gibson assembly
- Parallel promoter variant library construction
- Transformation or TXTL assembly
- 96-well high-throughput screening
- Quantitative readout (plate reader)
WORK FLOW
%%{init: {'themeVariables': { 'fontSize': '15px' }}}%%
flowchart LR
A["<b>1. Biomarker Selection</b><br/>Identify and select early molecular indicators of plant water stress."]
B["<b>2. Biosensor Design</b>
a. Genetic circuit design
b. Automated assembly and screening
c. Sensitivity and specificity validation<br/>"]
C["<b>3. Prototype Development</b>
a. Freeze-drying optimization
b. Paper device fabrication
c. Device integration"]
D["<b>4. Field Validation</b><br/>Correlate sensor output with physiological plant stress measurements"]
A --> B --> C --> D