Week 1 HW: Principles and Practices
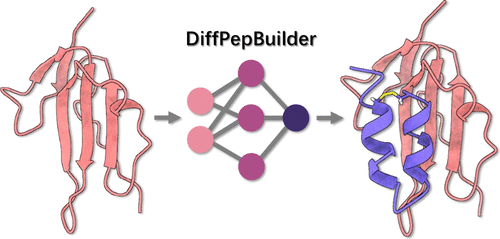

Governance of AI-Driven Biological Design
1. Biological Engineering Application or Tool
Description
As the core idea of my project, I would like to develop a concept that has been under discussion in my laboratory for some time: a computational platform for the de novo design of peptide and protein ligands capable of inhibiting essential microbial processes, such as translation, with the goal of suppressing or controlling microbial growth.
This platform would integrate computational biochemistry, focused on deep-learning-based structure prediction, together with generative sequence design and molecular dynamics simulations to generate ligands that selectively bind to key molecular interactions in microbial metabolism, including protein–protein and protein–nucleic acid interactions. Tools such as RosettaFold Diffusion and BindCraft have demonstrated remarkable capabilities for designing ligands with high affinity and specificity, consolidating this approach as a promising strategy for the rational development of new antimicrobial agents.
2. Governance and Policy Goals
General Governance Objective
Ensure that computational platforms for antimicrobial ligand design are developed and implemented in ways that maximize public health benefits while minimizing risks of misuse, accidental harm, and unethical applications.
Objective 1: Prevent Malicious or Irresponsible Use of Molecular Design Platforms
Sub-goals:
Limit the use of ligand design tools to prevent the generation of molecules that could increase microbial virulence, toxicity, immune evasion, or environmental persistence.
Establish mechanisms to identify, evaluate, and manage cases of dual-use research of concern (DURC) arising from computational molecular engineering.
Implement educational mechanisms focused on responsible AI use, aimed at preventing unintentionally harmful applications.
Objective 2: Strengthen Biosecurity Throughout the Research Process
Sub-goals:
- Ensure that computational design workflows incorporate early biological risk assessment, and promote rigorous experimental validation protocols that evaluate toxicity and potential ecological impact before advancing projects.
Objective 3: Promote Responsible Innovation and Transparency in AI-Driven Bioengineering
Sub-goals:
- Encourage documentation, traceability, and auditability of molecular design decisions, supported by open scientific communication and the sharing of best practices related to risk prevention and mitigation.
3. Governance Actions
Action 1 — Integrated Biosecurity Filters in Molecular Design Platforms
Purpose: Current molecular design tools optimize binding and stability without systematic safety analysis. The proposed change is to integrate mandatory biosecurity filters that detect or log potentially dangerous sequences or misuse, particularly those related to pathogens or viral components.
Design: This action would involve collaboration with the laboratories and organizations responsible for developing these software platforms to implement built-in safety checks.
Assumptions: It is assumed that harmful biological functions can be predicted and flagged computationally with sufficient accuracy.
Risks of Failure & Success:
Failures include false negatives, where harmful designs are not detected. Additionally, many platforms operate locally and offline, limiting centralized monitoring.
Action 2 — Institutional Oversight for AI-Driven Molecular Engineering
Purpose: Establish specialized review processes to evaluate dual-use risks before experimental implementation.
Design:
Universities, ethics committees, funding agencies, and regulatory bodies would implement multidisciplinary review panels and mandatory risk assessments prior to project approval and funding.
Assumptions: This approach assumes institutional capacity for technical risk evaluation and researcher compliance.
Risks of Failure & Success:
Excessive regulation may discourage exploratory research*, particularly in low-resource environments.
Action 3 — Tiered Access and Licensing of Advanced Molecular Design Platforms
Purpose:
Implement user identification and credential-based access models to monitor and deter misuse.
Design:
Platform developers, regulatory agencies, and academic consortia would establish access levels, authentication systems, and activity monitoring.
Assumptions:
This approach assumes credential-based access control is enforceable and accepted by researchers.
Risks of Failure & Success: Failures include excluding under-resourced researchers and the emergence of unregulated alternative tools.
4. Governance Option Evaluation Matrix
| Does the option: | Option 1 | Option 2 | Option 3 |
|---|---|---|---|
| Enhance Biosecurity | |||
| • By preventing incidents | 1 | 2 | 1 |
| • By helping respond | 2 | 1 | 2 |
| Foster Lab Safety | |||
| • By preventing incident | 2 | 1 | 2 |
| • By helping respond | 2 | 1 | 2 |
| Protect the environment | |||
| • By preventing incidents | 2 | 1 | 2 |
| • By helping respond | 3 | 1 | 2 |
| Other considerations | |||
| • Minimizing costs and burdens to stakeholders | 2 | 3 | 3 |
| • Feasibility? | 1 | 2 | 3 |
| • Not impede research | 2 | 3 | 1 |
| • Promote constructive applications | 1 | 2 | 2 |
5. Governance Prioritization and Recommendation
Based on the scoring and overall evaluation, I would prioritize interinstitutional oversight mechanisms and tiered-access systems as the most effective governance options. The main reason for this prioritization is that the primary risk factor for misuse lies in the broad open access to these software tools, combined with limited monitoring of their actual use. Since many AI-based biological design platforms are freely accessible, traceability, accountability, and early risk detection are currently limited, substantially increasing the risk of accidental misuse or deliberate malicious exploitation.
The administrative burden, slower research workflows, and barriers for resource-limited institutions were considered key trade-offs. However, the significant improvements in risk mitigation, accountability, and governance transparency outweigh these disadvantages. Furthermore, these limitations can be reduced through careful system design, including fast-track approval procedures and international collaboration frameworks.
This recommendation assumes that institutions possess the organizational and technical capacity to implement oversight systems and that researchers will comply with access regulations. Major uncertainties remain regarding global regulatory harmonization, consistent enforcement, and the adaptability of governance frameworks in response to the rapid evolution of AI capabilities.
Target Audience: This recommendation is primarily directed at major research institutions, international scientific organizations, and national regulatory agencies, aiming to establish coordinated governance structures that balance innovation, safety, and public protection.
Homework Questions & Answers
Homework Questions from Professor Jacobson
Questions
Nature’s machinery for copying DNA is called polymerase. What is the error rate of polymerase? How does this compare to the length of the human genome? How does biology deal with that discrepancy?
How many different ways are there to code (DNA nucleotide code) for an average human protein? In practice, what are some of the reasons that all of these different codes don’t work to code for the protein of interest?
Answers
DNA polymerase has an intrinsic error rate of approximately 1 mistake per 10⁶ nucleotides incorporated. Given that the human genome is roughly 3 × 10⁹ base pairs long, this would result in thousands of errors per replication cycle in the absence of correction mechanisms. Biology addresses this discrepancy through the proofreading activity of DNA polymerase and post-replicative mismatch repair systems, which dramatically reduce the final mutation rate.
Due to the degeneracy of the genetic code, there exists an astronomically large number of DNA sequences capable of encoding an average human protein. However, in practice, not all of these sequences are equally viable. Factors such as mRNA stability, codon usage bias, translational efficiency, secondary structure formation, and regulatory sequence constraints limit the set of functional coding sequences.
Homework Questions from Dr. LeProust
Questions
- What’s the most commonly used method for oligo synthesis currently?
- Why is it difficult to make oligos longer than 200 nt via direct synthesis?
- Why can’t you make a 2000 bp gene via direct oligo synthesis?
Answers
The most commonly used method for oligonucleotide synthesis is solid-phase chemical synthesis using phosphoramidite chemistry.
It is difficult to synthesize oligonucleotides longer than ~200 nucleotides because the coupling efficiency at each synthesis cycle is not perfect, leading to the progressive accumulation of errors and truncated products as the length increases.
A 2000 bp gene cannot be synthesized directly because the cumulative error rate and product truncation become overwhelmingly high, preventing the recovery of a correct full-length sequence in sufficient yield and purity.
Homework Question from George Church
Question
What are the 10 essential amino acids in all animals, and how does this affect your view of the “Lysine Contingency”?
Answer
The ten essential amino acids in animals are:
- Histidine
- Isoleucine
- Leucine
- Lysine
- Methionine
- Phenylalanine
- Threonine
- Tryptophan
- Valine
- Arginine
title: ‘Week 2’ weight: 10
Homework Questions & Answers
Homework Questions from Professor Jacobson
Questions
Nature’s machinery for copying DNA is called polymerase. What is the error rate of polymerase? How does this compare to the length of the human genome? How does biology deal with that discrepancy?
How many different ways are there to code (DNA nucleotide code) for an average human protein? In practice, what are some of the reasons that all of these different codes don’t work to code for the protein of interest?
Answers
DNA polymerase has an intrinsic error rate of approximately 1 mistake per 10⁶ nucleotides incorporated. Given that the human genome is roughly 3 × 10⁹ base pairs long, this would result in thousands of errors per replication cycle in the absence of correction mechanisms. Biology addresses this discrepancy through the proofreading activity of DNA polymerase and post-replicative mismatch repair systems, which dramatically reduce the final mutation rate.
Due to the degeneracy of the genetic code, there exists an astronomically large number of DNA sequences capable of encoding an average human protein. However, in practice, not all of these sequences are equally viable. Factors such as mRNA stability, codon usage bias, translational efficiency, secondary structure formation, and regulatory sequence constraints limit the set of functional coding sequences.
Homework Questions from Dr. LeProust
Questions
- What’s the most commonly used method for oligo synthesis currently?
- Why is it difficult to make oligos longer than 200 nt via direct synthesis?
- Why can’t you make a 2000 bp gene via direct oligo synthesis?
Answers
The most commonly used method for oligonucleotide synthesis is solid-phase chemical synthesis using phosphoramidite chemistry.
It is difficult to synthesize oligonucleotides longer than ~200 nucleotides because the coupling efficiency at each synthesis cycle is not perfect, leading to the progressive accumulation of errors and truncated products as the length increases.
A 2000 bp gene cannot be synthesized directly because the cumulative error rate and product truncation become overwhelmingly high, preventing the recovery of a correct full-length sequence in sufficient yield and purity.
Homework Question from George Church
Question
What are the 10 essential amino acids in all animals, and how does this affect your view of the “Lysine Contingency”?
Answer
The ten essential amino acids in animals are:
- Histidine
- Isoleucine
- Leucine
- Lysine
- Methionine
- Phenylalanine
- Threonine
- Tryptophan
- Valine
- Arginine