Week 2 — DNA Read, Write, & Edit
Part 1: Benchling & In-silico Gel Art
Simulate Restriction Enzyme Digestion with the following Enzymes:
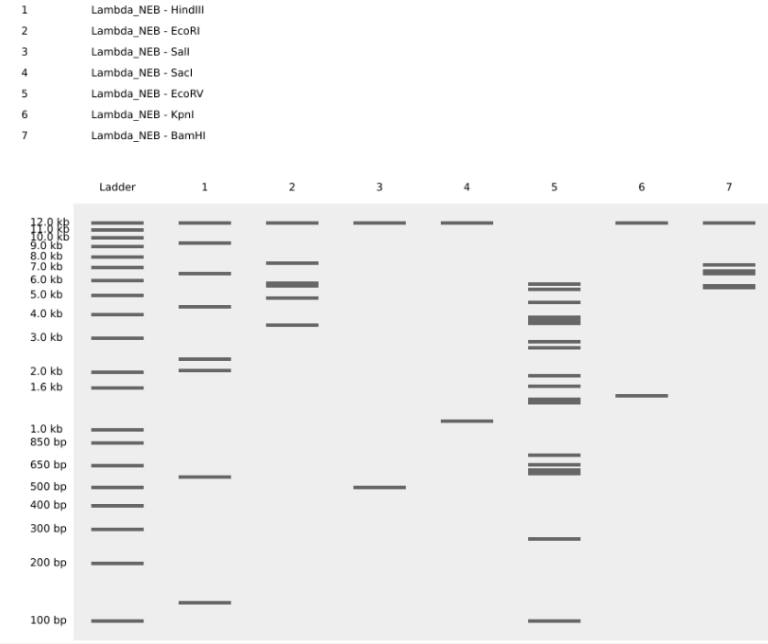


Part 3: DNA Design Challenge
3.1. Choose your protein.
In recitation, we discussed that you will pick a protein for your homework that you find interesting. Which protein have you chosen and why? Using one of the tools described in recitation (NCBI, UniProt, google), obtain the protein sequence for the protein you chose.
The protein selected for this assignment is Translation Initiation Factor IF-2 (IF2) from Escherichia coli (strain K12).
I chose IF2 because it plays a central role in the initiation of protein synthesis, a critical and highly regulated step of gene expression. IF2 is responsible for promoting the binding of the initiator tRNA to the ribosomal P site and facilitating the correct assembly of the translation initiation complex. Due to its essential function, IF2 is a key factor in controlling translational efficiency and fidelity. Additionally, IF2 is highly conserved among bacteria, making it an important target for studies in molecular biology, ribosome dynamics, and antibiotic development. Its biological relevance and mechanistic complexity make it a particularly interesting protein to study.
The amino acid sequence of IF2 was obtained from UniProt.
sp|P0A705|IF2_ECOLI Translation initiation factor IF-2 OS=Escherichia coli (strain K12) OX=83333 GN=infB PE=1 SV=1 MTDVTIKTLAAERQTSVERLVQQFADAGIRKSADDSVSAQEKQTLIDHLNQKNSGPDKLT LQRKTRSTLNIPGTGGKSKSVQIEVRKKRTFVKRDPQEAERLAAEEQAQREAEEQARREA EESAKREAQQKAEREAAEQAKREAAEQAKREAAEKDKVSNQQDDMTKNAQAEKARREQEA AELKRKAEEEARRKLEEEARRVAEEARRMAEENKWTDNAEPTEDSSDYHVTTSQHARQAE DESDREVEGGRGRGRNAKAARPKKGNKHAESKADREEARAAVRGGKGGKRKGSSLQQGFQ KPAQAVNRDVVIGETITVGELANKMAVKGSQVIKAMMKLGAMATINQVIDQETAQLVAEE MGHKVILRRENELEEAVMSDRDTGAAAEPRAPVVTIMGHVDHGKTSLLDYIRSTKVASGE AGGITQHIGAYHVETENGMITFLDTPGHAAFTSMRARGAQATDIVVLVVAADDGVMPQTI EAIQHAKAAQVPVVVAVNKIDKPEADPDRVKNELSQYGILPEEWGGESQFVHVSAKAGTG IDELLDAILLQAEVLELKAVRKGMASGAVIESFLDKGRGPVATVLVREGTLHKGDIVLCG FEYGRVRAMRNELGQEVLEAGPSIPVEILGLSGVPAAGDEVTVVRDEKKAREVALYRQGK FREVKLARQQKSKLENMFANMTEGEVHEVNIVLKADVQGSVEAISDSLLKLSTDEVKVKI IGSGVGGITETDATLAAASNAILVGFNVRADASARKVIEAESLDLRYYSVIYNLIDEVKA AMSGMLSPELKQQIIGLAEVRDVFKSPKFGAIAGCMVTEGVVKRHNPIRVLRDNVVIYEG ELESLRRFKDDVNEVRNGMECGIGVKNYNDVRTGDVIEVFEIIEIQRTIA
3.2. Reverse Translate: Protein (amino acid) sequence to DNA (nucleotide) sequence.
The Central Dogma discussed in class and recitation describes the process in which DNA sequence becomes transcribed and translated into protein. The Central Dogma gives us the framework to work backwards from a given protein sequence and infer the DNA sequence that the protein is derived from. Using one of the tools discussed in class, NCBI or online tools (google “reverse translation tools”), determine the nucleotide sequence that corresponds to the protein sequence you chose above.
Using the reverse translation tool available in Benchling, the amino acid sequence of Translation Initiation Factor IF-2 (IF2) from Escherichia coli (strain K12) was converted into a corresponding nucleotide sequence.
IF2_ECOLI_DNA ATGACCGATGTGACCATTAAAACCCTGGCGGCGGAACGCCAGACCAGCGTGGAACGCCTGGTGCAGCAGTTTGCGGATGCGGGCATTCGCAAAAGCGCGGATGATAGCGTGAGCGCGCAGGAAAAACAGACCCTGATTGATCATCTGAACCAGAAAAACAGCGGCCCGGATAAACTGACCCTGCAGCGCAAAACCCGCAGCACCTTGAACATTCCGGGCACCGGCGGCAAAAGCAAAAGCGTGCAGATTGAAGTGCGCAAAAAACGCACCTTTGTGAAACGCGATCCGCAGGAAGCGGAACGCCTGGCGGCGGAAGAACAGGCGCAGCGCGAAGCGGAAGAACAGGCGCGCCGCGAAGCGGAAGAAAGCGCGAAACGCGAAGCGCAGCAGAAAGCGGAACGCGAAGCGGCGGAACAGGCGAAACGCGAAGCGGCGGAACAGGCGAAACGCGAAGCGGCGGAAAAAGATAAAGTGAGCAACCAGCAGGATGATATGACCAAAAACGCGCAGGCGGAAAAAGCGCGCCGCGAACAGGAAGCGGCGGAACTGAAACGCAAAGCGGAAGAAGAAGCGCGCCGCAAACTGGAAGAAGAAGCGCGCCGCGTGGCGGAAGAAGCGCGCCGCATGGCGGAAGAAAACAAATGGACCGATAACGCGGAACCGACCGAAGATAGCAGCGATTATCATGTGACCACCAGCCAGCATGCGCGCCAGGCGGAAGATGAAAGCGATCGCGAAGTGGAAGGCGGCCGCGGCCGCGGCCGCAACGCGAAAGCGGCGCGCC…
I’ve shortened the sequense because it’s very long.
3.3. Codon optimization.
Once a nucleotide sequence of your protein is determined, you need to codon optimize your sequence. You may, once again, utilize google for a “codon optimization tool”. In your own words, describe why you need to optimize codon usage. Which organism have you chosen to optimize the codon sequence for and why?
Codon optimization is necessary because, although multiple codons can encode the same amino acid, different organisms show preferences for specific codons. These preferences are related to the abundance of corresponding tRNAs and directly affect translation efficiency, protein yield, and overall expression levels.
The organism selected for codon optimization was Escherichia coli, because it is one of the most widely used hosts for recombinant protein expression. E. coli offers fast growth, low cost, well-established genetic tools, and high-level protein production.
I honestly don’t know how to represent it, so I’m going to include the evidence from Bencling.
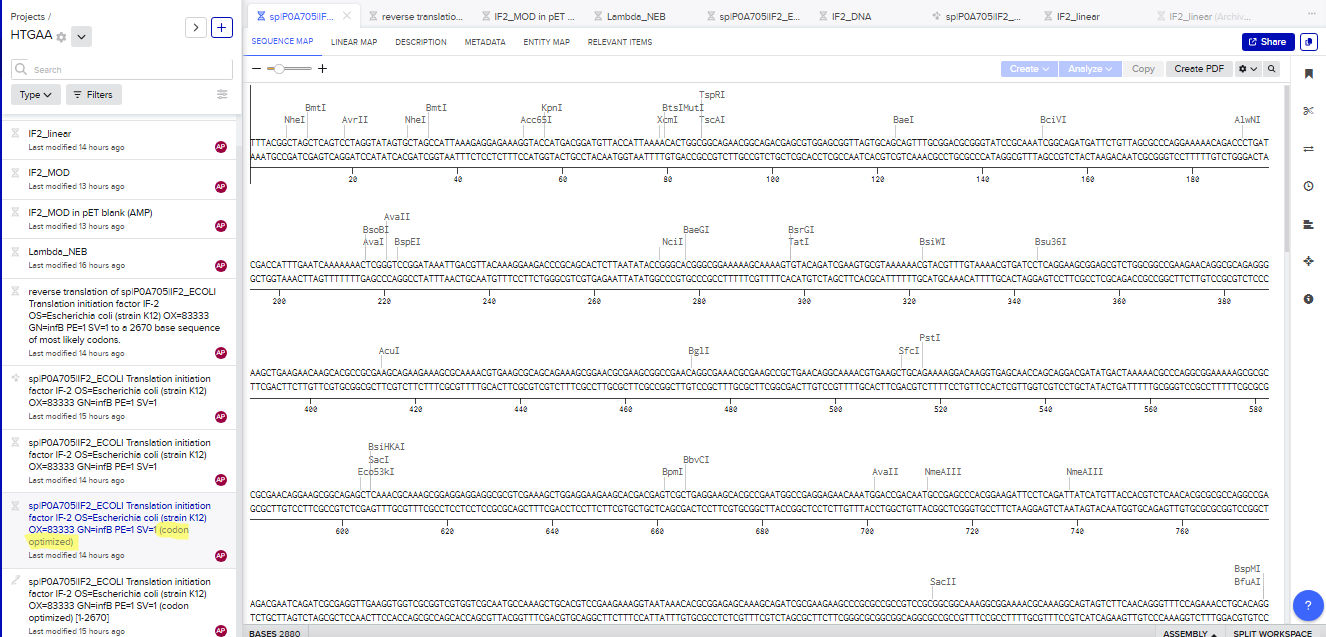

3.4. You have a sequence! Now what?
What technologies could be used to produce this protein from your DNA? Describe in your words the DNA sequence can be transcribed and translated into your protein. You may describe either cell-dependent or cell-free methods, or both.
In a cell-dependent system, the optimized DNA is cloned into an expression plasmid and introduced into E. coli. Inside the cell, the DNA is transcribed into mRNA and then translated by ribosomes into a protein, which folds into its functional form and can later be purified using techniques such as affinity chromatography.
From personal experience I use cell-dependent but the approach of both systems, the underlying process follows the Central Dogma of Molecular Biology, where DNA is transcribed into mRNA and then translated into protein.
Part 4: Prepare a Twist DNA Synthesis Order
4.1. and 4.2. Build Your DNA Insert Sequence
The final DNA construct includes the essential regulatory and coding elements required for protein expression: a promoter to initiate transcription, a ribosome binding site (RBS) to enable efficient translation, a start codon (ATG), the codon-optimized coding sequence of the target protein, a C-terminal 7×His tag to facilitate protein purification, a stop codon, and a transcription terminator to properly end transcription. Together, these components ensure efficient transcription, translation, and purification of the recombinant protein in E. coli.
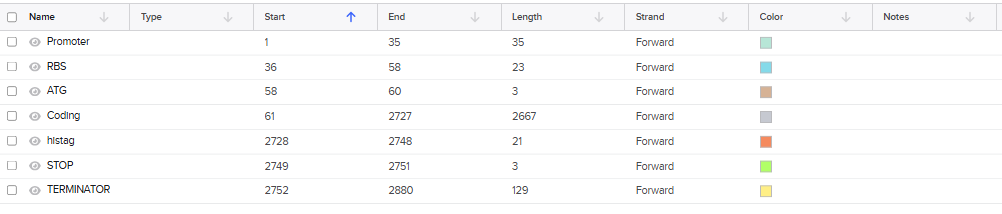



4.3. to 4.6.
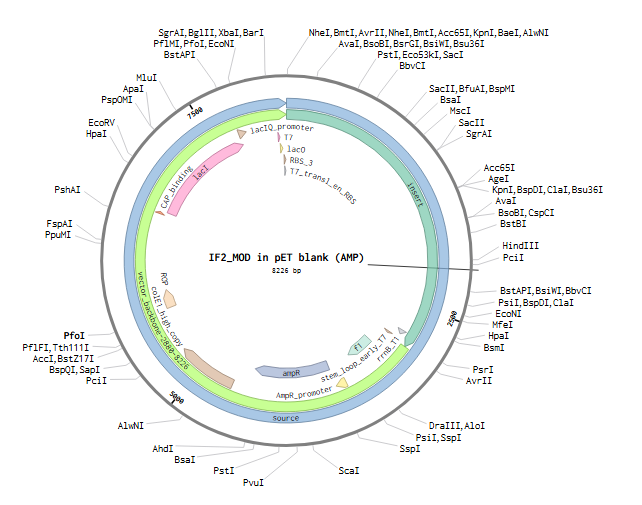

The pET target (AMP), a recombinant expression vector intended for Escherichia coli IF2 protein production, displays the IF2_MOD plasmid map in the figure. Strong, controlled transcription and translation of the inserted gene are made possible by the plasmid’s T7 promoter and T7 ribosome binding site (RBS). The lac operator (lacO) and the lacI repressor regulate expression, enabling IPTG induction. The plasmid also has a high-copy-number replication origin, which guarantees effective plasmid maintenance, and the ampicillin resistance gene (ampR), which is used to select transformed cells.
Part 5: DNA Read/Write/Edit
(i) What DNA would you want to sequence (e.g., read) and why? This could be DNA related to human health (e.g. genes related to disease research), environmental monitoring (e.g., sewage waste water, biodiversity analysis), and beyond (e.g. DNA data storage, biobank).
I would want to sequence genes related to antibiotic resistance in pathogenic bacteria, because antimicrobial resistance is a major global health problem. By sequencing these genes, we can identify resistance mechanisms, track how they spread among bacterial populations, and monitor the emergence of new resistant strains. This information is essential for improving disease surveillance, guiding treatment decisions, and developing new strategies to control infections.
(ii) In lecture, a variety of sequencing technologies were mentioned. What technology or technologies would you use to perform sequencing on your DNA and why?
For exaple: This method is a second-generation sequencing technology that is ideal for studying antibiotic resistance genes in pathogenic bacteria. The input is purified DNA extracted from bacterial or environmental samples, which is prepared by fragmentation, adapter ligation, and PCR amplification to generate a sequencing library. Sequencing is performed using sequencing-by-synthesis, where fluorescently labeled nucleotides are incorporated and detected to accurately decode each DNA base. The output consists of millions of short DNA sequence reads, which can be analyzed to identify resistance genes, detect mutations, and monitor their spread in bacterial populations.
5.2 DNA Write
(i) What DNA would you want to synthesize (e.g., write) and why? These could be individual genes, clusters of genes or genetic circuits, whole genomes, and beyond. As described in class thus far, applications could range from therapeutics and drug discovery (e.g., mRNA vaccines and therapies) to novel biomaterials (e.g. structural proteins), to sensors (e.g., genetic circuits for sensing and responding to inflammation, environmental stimuli, etc.), to art (DNA origamis). If possible, include the specific genetic sequence(s) of what you would like to synthesize! You will have the opportunity to actually have Twist synthesize these DNA constructs! :)
I would like to synthesize genes with optimized codons that encode antimicrobial peptides and inhibitors of proteins involved in bacterial translation, such as peptides that interact with IF2, as these could be used for drug discovery and the development of new antibiotics. These synthetic DNA sequences could then be inserted into expression vectors to rapidly produce and test novel therapeutic proteins.
(ii) What technology or technologies would you use to perform this DNA synthesis and why?
I would use solid-phase phosphoramidite DNA synthesis, followed by FPLC purification, because this method allows precise and reliable chemical synthesis of custom DNA sequences. The essential steps include stepwise nucleotide coupling, oxidation, capping, and deprotection to build the DNA strand, followed by cleavage from the solid support. The main limitations of this method are sequence length constraints (typically up to ~200 bp per fragment), synthesis errors that accumulate with longer sequences, and moderate scalability, which requires assembly of longer constructs from shorter fragments.
5.3 DNA Edit
(i) What DNA would you want to edit and why?
I want to edit bacterial genes involved in antibiotic resistance and translation initiation, such as infB (encoding IF2), to study their function and to develop new antimicrobial strategies. Editing this DNA would allow precise modification of key residues to understand their role in protein synthesis and to identify vulnerabilities that can be targeted for drug development.
(ii) What technology or technologies would you use to perform these DNA edits and why?
I would use CRISPR-Cas9 gene editing technology because it allows precise, efficient, and targeted modification of DNA sequences. Its high accuracy, simplicity, and adaptability make it ideal for editing bacterial genes such as infB to investigate protein function and antibiotic resistance mechanisms.