Week 4 — Protein Design Part I
Part A. Conceptual Questions
Why do humans eat beef but not turn into cows, and eat fish but not turn into fish? Organisms do not incorporate intact proteins. Instead, they degrade them into amino acids, which are then reassembled according to the organism’s own genome. Molecular identity is determined by genetic information, not by the origin of the raw material.
Why are there only 20 natural amino acids? Only 20 were selected by early evolution because they possessed a specific chemistry that allowed for a functional balance of hydrophobicity, specific folding patterns, and various electrical charges.
Can unnatural amino acids be created? Design new ones. Yes. Thousands have already been created; the limitation is functional rather than chemical. Rational Design Examples:
Fluorinated side-chain amino acid: Increases hydrophobicity and thermal stability.
Azide or Alkyne group amino acid: Enables “click chemistry” for selective molecular labeling.
Where did amino acids come from before enzymes and life? There is solid evidence from abiotic synthesis, most notably demonstrated by the Miller–Urey experiments, which showed that organic compounds could form from inorganic precursors under simulated primitive Earth conditions.
If you build an α-helix with D-amino acids, what chirality would it have? It would form a left-handed (levogyre) helix.
Can you discover additional helices in proteins? The common assumption that only α-helices and 3 10
-helices exist is false. Others include:
π-helices
Polyproline helices (I and II)
Mixed helices
Transient dynamic helices
- Why are most molecular helices right-handed? Biological homochirality (the prevalence of L-amino acids) imposes a geometry that:
Minimizes steric collisions.
Optimizes hydrogen bonding.
Maximizes overall structural stability.
Why do β-sheets tend to aggregate? In physical terms, aggregation reduces free energy, making the clustered state more thermodynamically favorable.
What is the driving force behind β-aggregation? It is not a single force, but rather a balance of:
Cooperative hydrogen bonding.
The hydrophobic effect.
Entropy gain from released water molecules.
Steric “zipper” stacking.
- Can β-amyloid sheets be used as materials? Yes, this is a key emerging field. Applications include:
Mechanically resistant nanofibers.
Scaffolds for tissue engineering.
Part B. Protein Analysis and Visualization1
- Amino Acid Sequence IdentificationProtein:
- Nipah virus G glycoprotein
- Length: 602 amino acids
- Most frequent amino acid: Leucine (L)
- Homologs: 146 hits found in UniProtKB.
- Family: Attachment glycoproteins (Paramyxoviridae / Henipavirus).
- Structural Analysis (RCSB PDB: 2VSM)
- Structure Name: Nipah virus attachment glycoprotein in complex with human cell surface receptor ephrinB2.
- Resolution and Quality: 1.80 Å (High quality, resolved on May 20, 2008).
- Other Molecules: Contains NAG (N-acetylglucosamine) ligands and the human ephrin-B2 receptor.
- Structural Classification: Classified as a Hydrolase.
- 3D Visualization Guide
- Cartoon Representation:
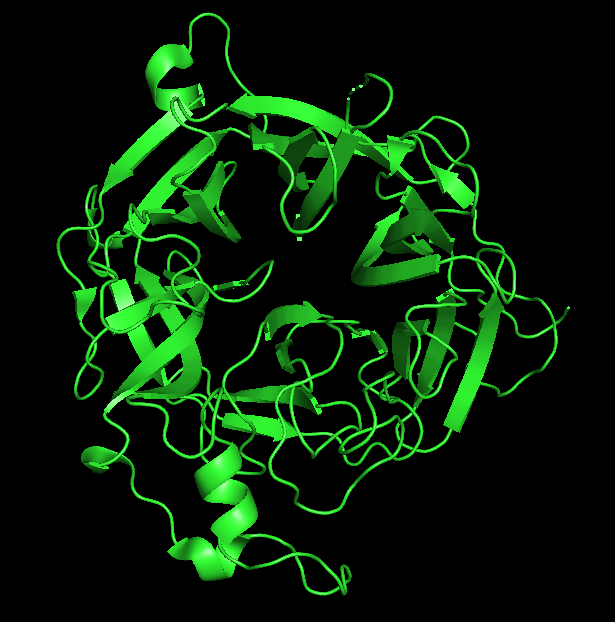

- Ribbon Representation:
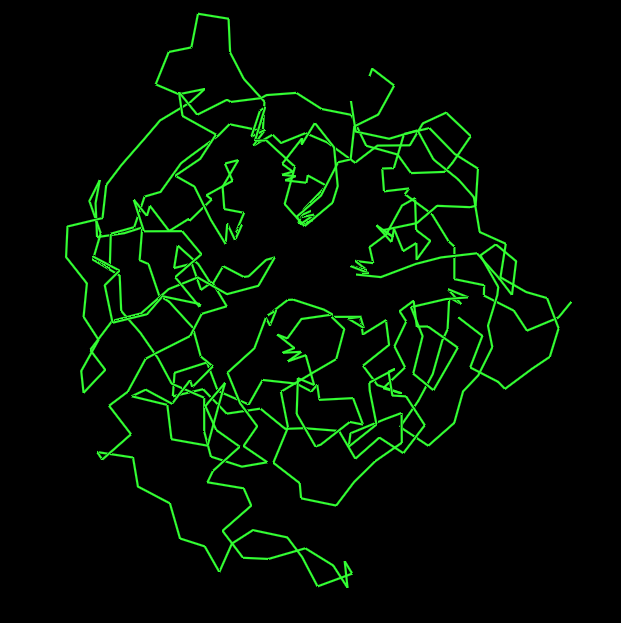

- Ball and Stick Representation:
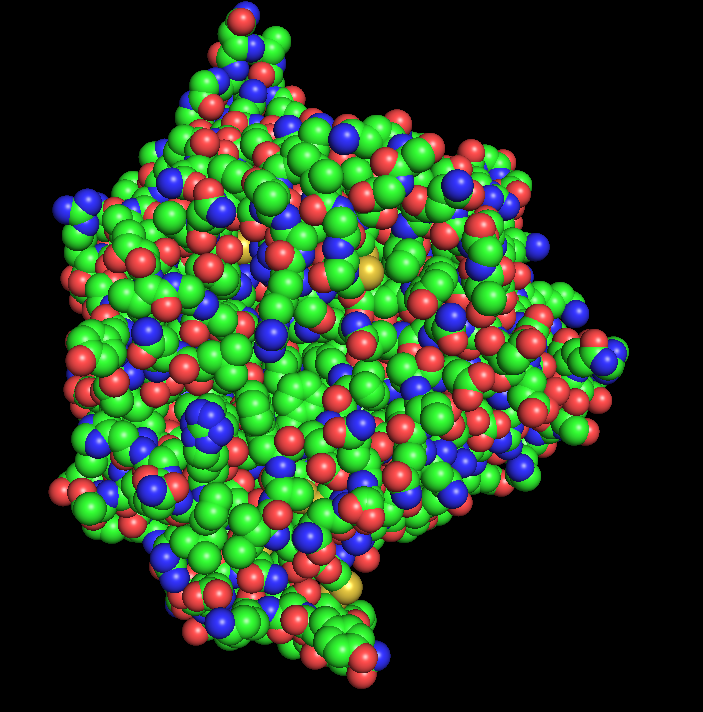




Part C. Using ML-Based Protein Design Tools
C1. Protein Language Modeling
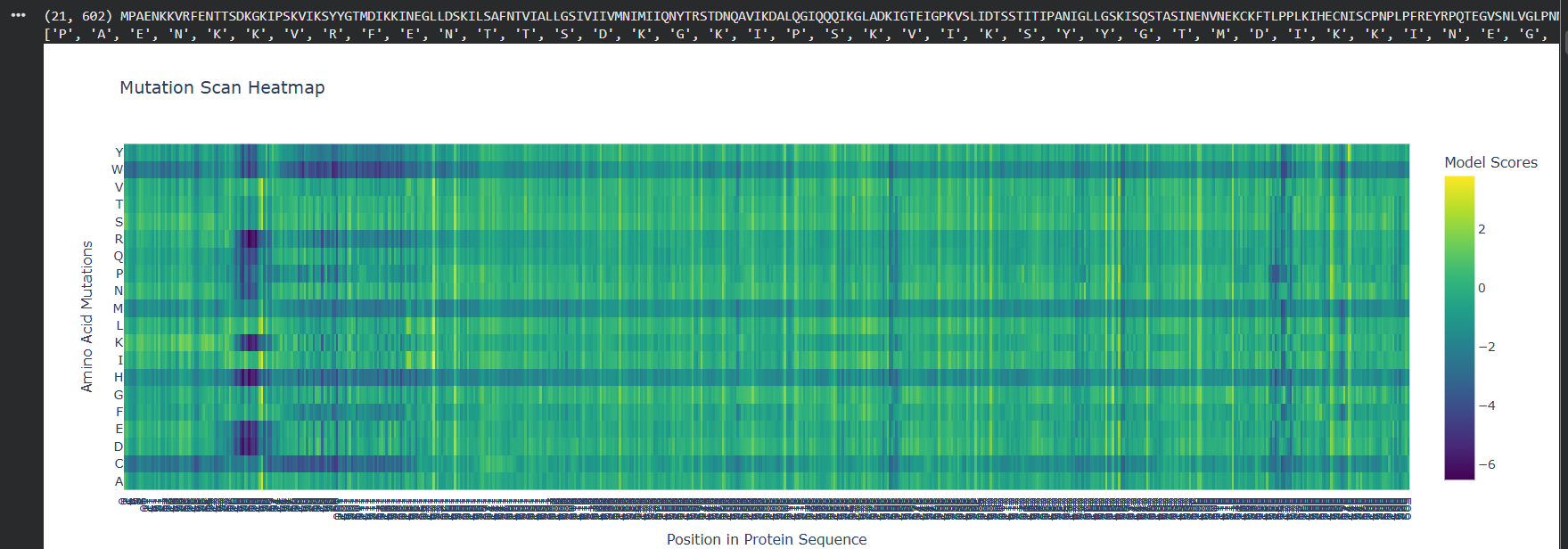
Deep Mutational ScansTo analyze the functional robustness of the Nipah virus G glycoprotein, the ESM2 language model was used to generate an unsupervised deep mutational scan (DMS) based on sequence likelihoods.
Heatmap Patterns: The mutation scan reveals distinct vertical bands of high conservation (dark purple), indicating positions where any amino acid substitution is highly penalized.
Critical Residues: A specific sensitive region is observed near the beginning of the sequence (approx. positions 100–120), where substitutions with large hydrophobic or charged residues significantly lower the model score, suggesting a critical structural core.
Functional Insights: These high-sensitivity areas likely correspond to the $\beta$-propeller folds required for ephrin-B2 receptor binding.
Latent Space Analysis
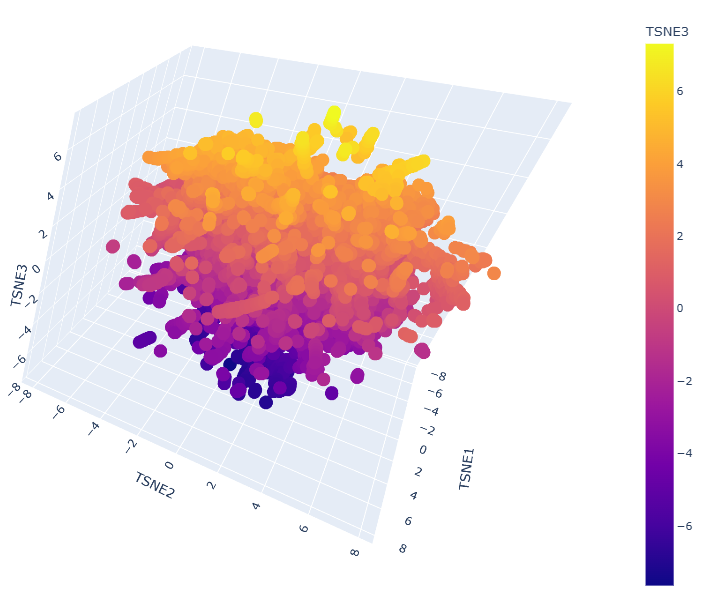
A latent space analysis was performed by embedding the protein sequence dataset into a reduced dimensionality using t-SNE.
Neighborhood Clusters: The 3D map shows clear clusters where proteins are organized by structural and evolutionary similarity.
Protein Positioning: The Nipah G protein is located within a specific neighborhood alongside other Henipavirus homologs (such as Hendra virus), confirming that the model captures biological relationships without explicit labels.

C2. Protein Folding
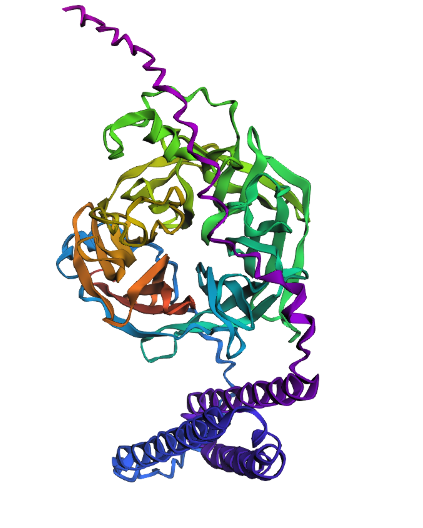
Folding with ESMFold The atomic-level structure of the protein was predicted from its primary sequence using ESMFold.
Coordinate Matching: The predicted coordinates show a high degree of structural alignment with the original experimental structure 2VSM.
Structural Features: The model accurately recovers the six-bladed $\beta$-propeller globular domain responsible for receptor recognition.
Mutation Resilience: While the protein core is resilient to single-point mutations suggested by the DMS, large segment deletions lead to a collapse in structural confidence (low pLDDT) in those regions.

C3. Protein Generation
Inverse-Folding with ProteinMPNN
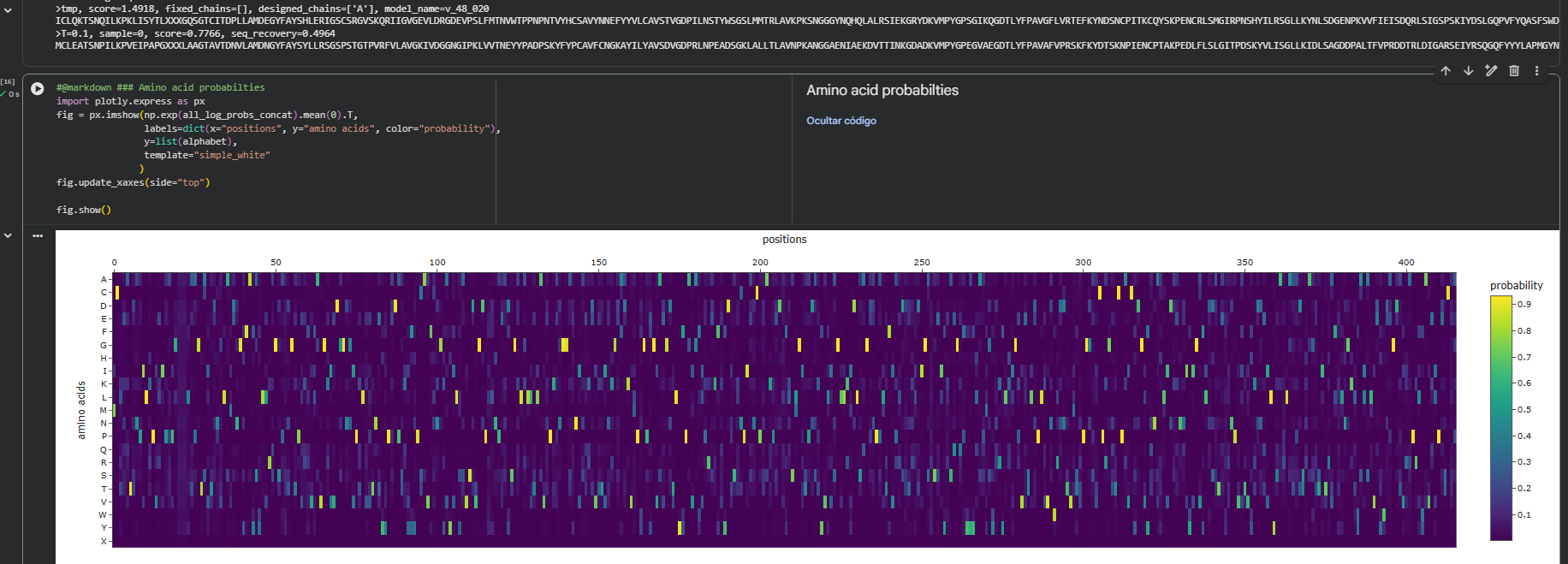
Using the backbone of the 2VSM structure, we utilized ProteinMPNN to propose new sequence candidates that could adopt the same fold.
Sequence Probabilities: The probability map indicates that the model has high confidence in specific “anchor” residues necessary to maintain internal packing.
Sequence Recovery: A sequence recovery of 49.6% was achieved (seq_recovery = 0.4964), showing that the model retains the essential native core while exploring variations on the protein surface.
Structural Validation: When the designed sequence was re-folded using ESMFold, the resulting structure matched the original backbone, validating the success of the inverse-folding design.

Part D. Group Brainstorm on Bacteriophage Engineering
Main Goals Stabilize the Lysis Protein (Protein L): Specifically focused on enhancing the stability of the transmembrane domain. Maintain Functional Motif Interaction: Ensuring that any modifications do not disrupt the critical interaction of the Leu48–Ser49 (LS) motif with its membrane target.
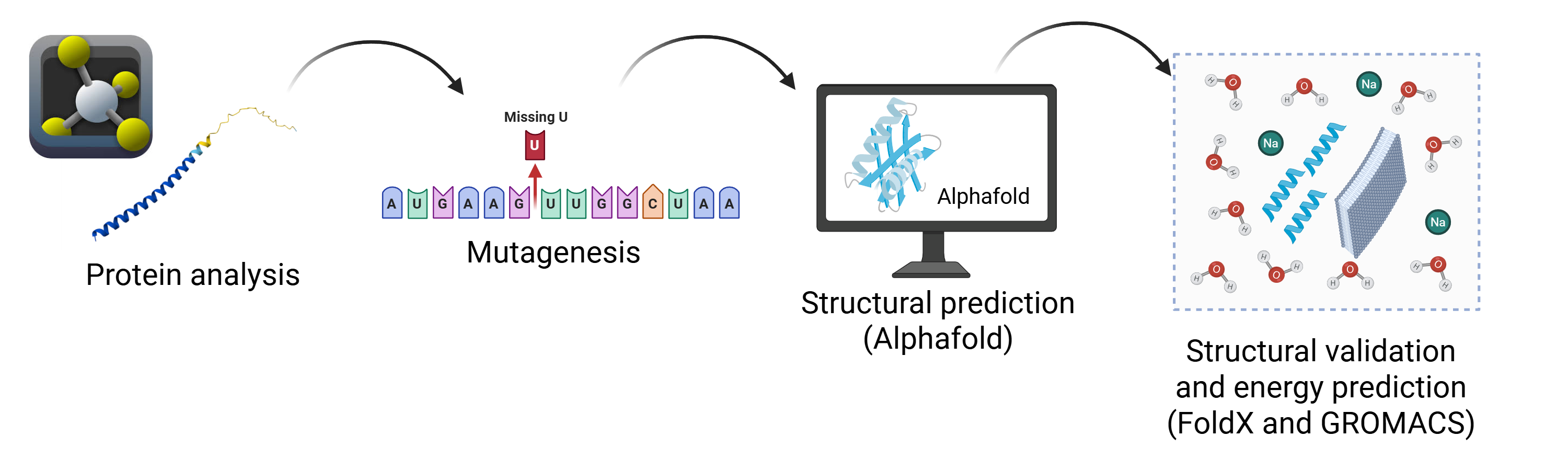
Proposed Tools and Approaches (Pipeline) The strategy combines evolutionary, structural, and physical perspectives to optimize the protein: Protein Language Models (pLMs): Used for directed in silico mutagenesis to generate variants consistent with evolutionary constraints. AlphaFold2: Employed for 3D structural prediction to verify that the transmembrane topology and functional orientation of the LS motif remain intact. FoldX / Rosetta: Used to estimate $\Delta\Delta G$ (Folding Free Energy), allowing for the prioritization of mutants with high thermodynamic stability. GROMACS: Utilized for Molecular Dynamics (MD) in a membrane environment to evaluate structural stability and flexibility within a realistic bacterial lipid bilayer.
Why These Tools? This integrated approach allows for a rational exploration of mutations while minimizing the risk of disrupting lytic function: pLMs ensure designed variants are likely to remain properly folded and functional. AlphaFold2 enables rapid structural screening of the transmembrane region. Energy-based predictions prioritize candidates, reducing the need for exhaustive downstream analysis. MD simulations provide a physiological context (bacterial membrane) necessary for validating transmembrane protein behavior.
Potential Pitfalls Prediction Accuracy: Current tools may have limited reliability for small, membrane-associated, and partially disordered proteins. Stability-Function Trade-off: Increasing thermodynamic stability might reduce the conformational flexibility required to interact with membrane targets, potentially impairing lytic activity.
Schematic of the Pipeline