Week 1 HW: Principles and Practices
The halfpipe of Doom- How to grow good?
For the first weeks lecture we had an introduction to the fundamental principles of synthetic biology and the HTGAA program. The focus of the lecture was on the governance and ethics of synthetic biology. David S. Kong discussed the balance between decentralized and centralized synBio development and the importance of thrust (something we are lacking these days). As a global community we have largely agreed to certain rules (e.g. bioweapon treaty 1975) however emerging synBio technologies also allow a much broader audience to participate in the development (e.g. community labs/ biohackers) that might not necessary always align with large governmental policies. He draws the parallel to how the early governance of the internet have allowed for a decentralized scaling that have contributed to an increased “computer literacy”. This might allow us to make better (although not perfect) personal decisions for how to use this new technology. Coming from a background of community focused biolab practice this was an interesting topic and made me think of the importance for a global bio-literacy. It also got me to think about the importance to apply these principals in a simple enough way that it doesn’t stifle participation.
Questions that I tried to include in my homework:
1. Describe a biological engineering application
Programmable colors for bacterial cellulose production
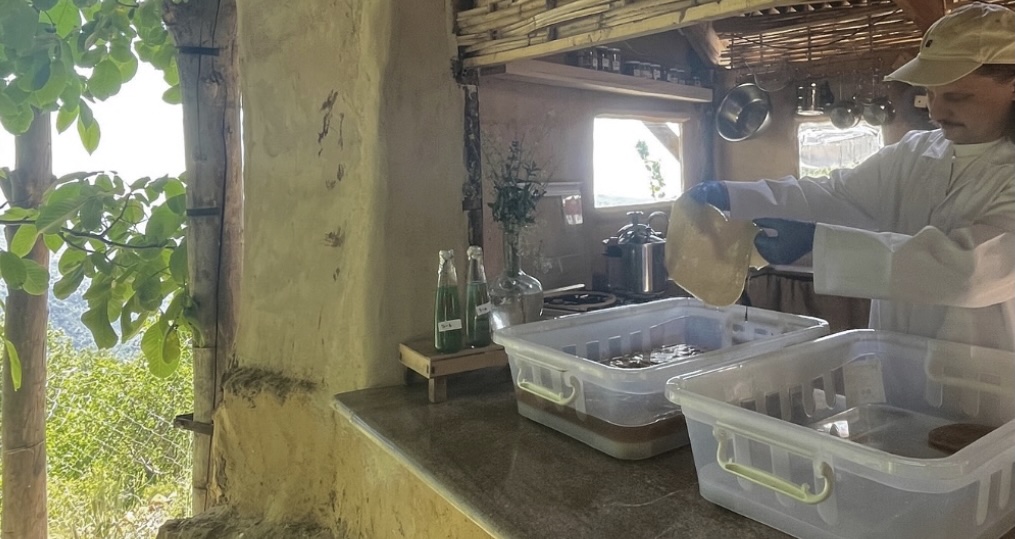
The textile dyeing industry is a major source of chemical pollution and water use. Coloration of bacterial cellulose (BC) can also be technically challenging because pigments often diffuse slowly into the material’s dense nanofibrillar network, making post-growth dyeing difficult and time consuming. This project proposes a bioengineering approach to generate color in situ during BC growth, eliminating conventional dyeing steps.
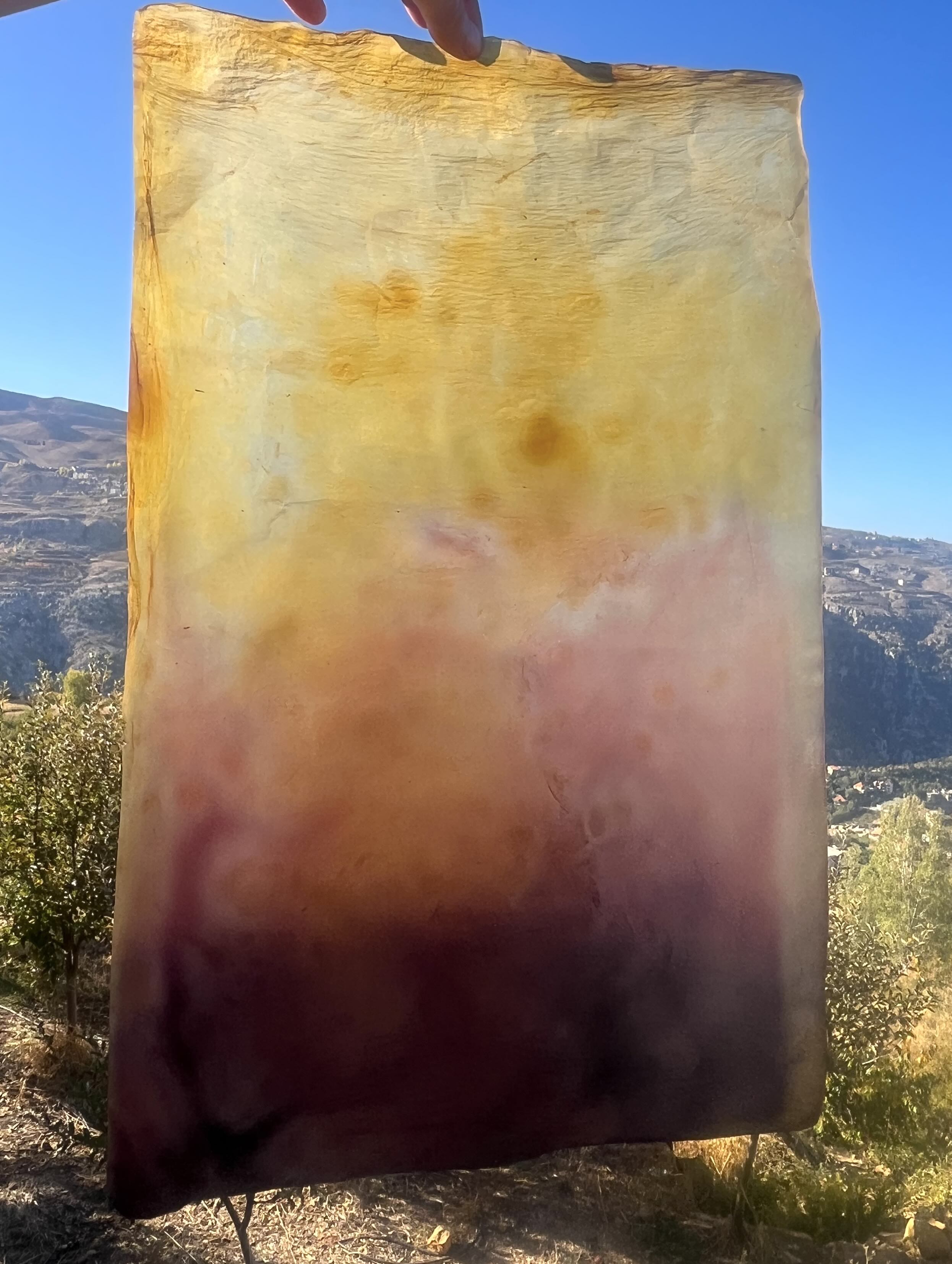

 Dyed BC I developed at TerraPods Lebanon
Dyed BC I developed at TerraPods Lebanon
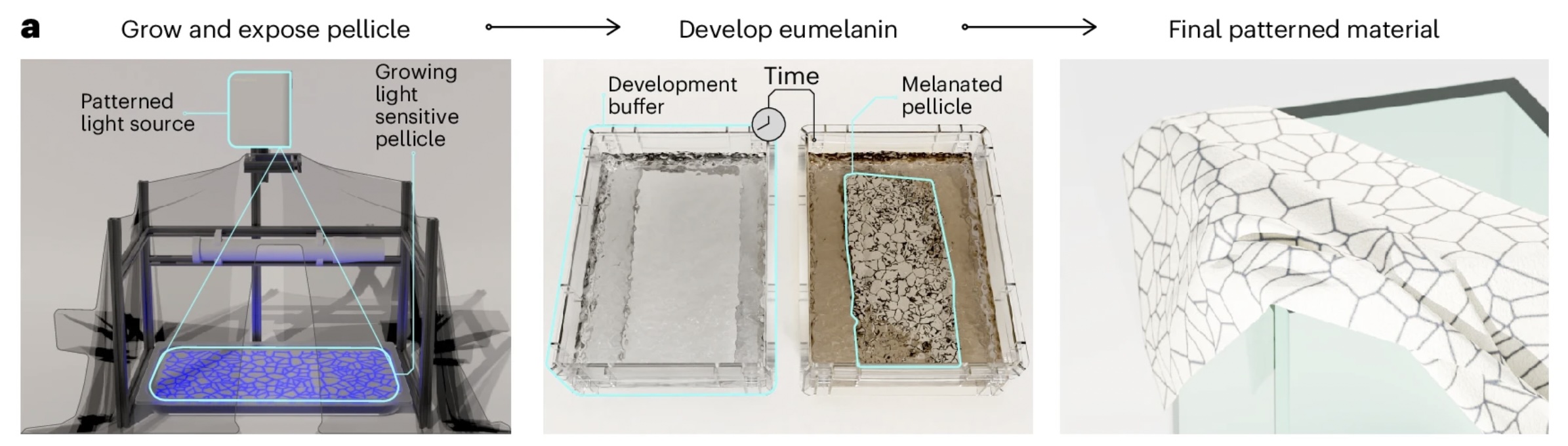
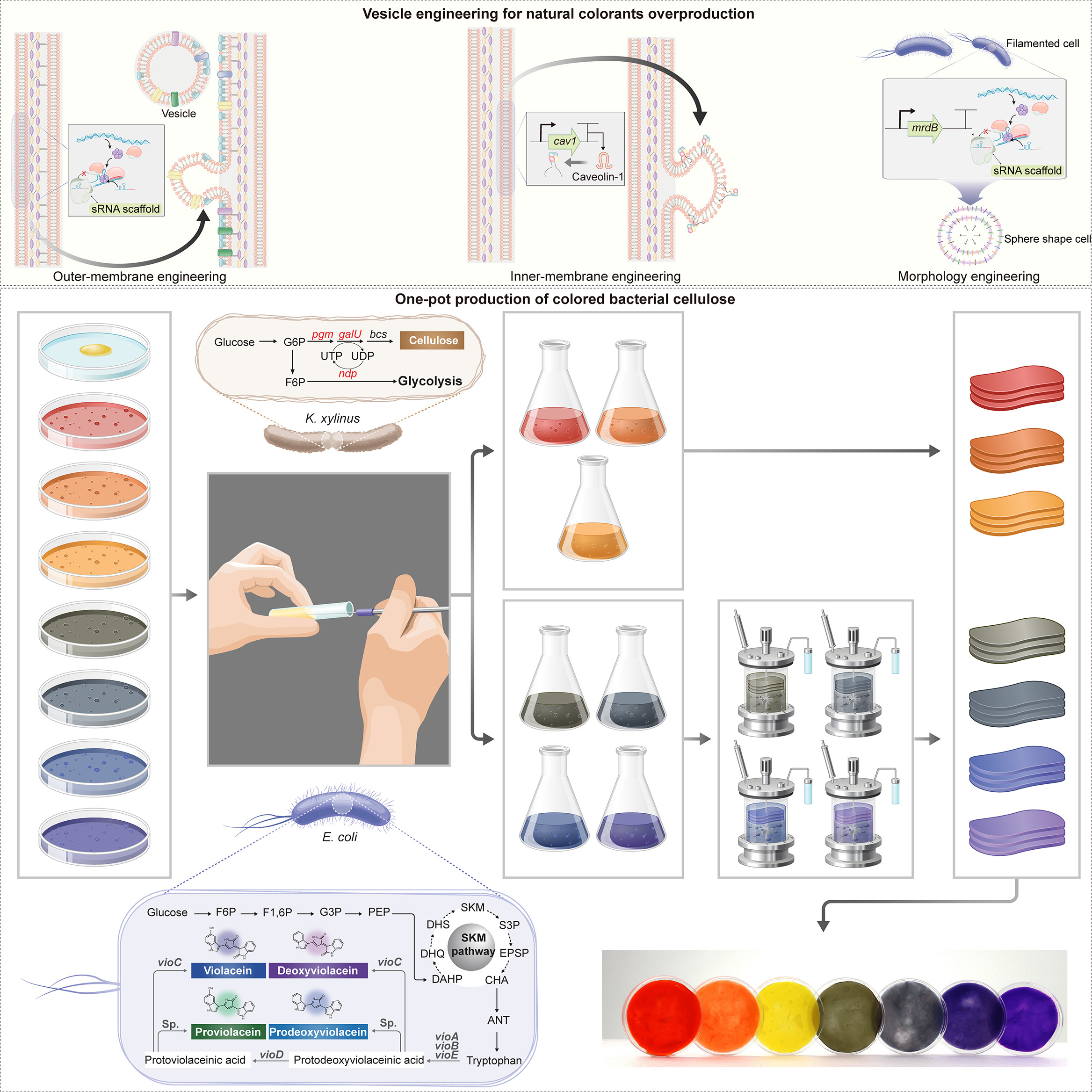
Prior work demonstrates the feasibility of embedding pigmentation into BC production. Walker et al.(2025) 1 engineered the cellulose-producing bacterium Komagataeibacter rhaeticus to generate melanin during BC growth, producing pigmented material. Zhou et al. (2025) 2 demonstrated a “one-pot” co-culture strategy coupling BC production by Komagataeibacter xylinus with pigments synthesised in engineered E. coli, enabling a broader palette by combining violacein derivatives (green/blue/navy/purple) and carotenoids (red/orange/yellow).
Building on these studies, the core concept here is light-patterned control of pigment production during BC formation. A cellulose-forming culture generates the sheet while a pigment-producing bacteria is engineered to be light-responsive, so that pigmentation occurs in illuminated regions. Patterned illumination via projection enables spatial control of coloration. Furthermore this technique would also enable varying projected patterns across growth phases that could yield multi-layer visual effects, (e.g. moiré-like effects).
Drawing from my previous experiences on working in various community biolab the project is framed as a distributed biofabrication platform for community labs, which creates governance questions around biosafety practice in a decentralized settings, concider the relative complex technique I was for this excersice imagining a centralized organization providing the framework and digital infrastructure for the community labs to safetly experiment with the protocol. Although consumer product are less ethically complicated then for example medicine or bioweapon their came up important questions concerning consumer/skin-contact safety, environmental release and waste handling, and norms for responsible dissemination of methods and bacteria strains.
2. governance/policy goals
- Actors: Community labs and networks, open-hardware designers, academic partners, funders, and (optionally) insurers.
A. Biosecurity
- A1: Reduce risk of malicious repurposing of organisms, materials, or protocols.
- A2: Improve traceability and incident reporting to support response.
B. Lab safety
- B1: Standardize safe practices (training, containment, waste handling) across labs.
- B2: Establish clear response procedures for spills, exposures, and contamination.
C. Environmental protection
- C1: Prevent release of organisms or harmful pigments/byproducts.
- C2: Enable remediation and corrective action after incidents.
D. User/consumer protection and social trust
- D1: Ensure skin-contact safety (low leaching, low irritation risk, stability).
- D2: Maintain low barrier access; avoid governance that excludes low-resource labs.
- D3: Require transparency and avoid misleading sustainability claims.
E. Feasibility and innovation
- E1: Keep requirements simple for community labs.
- E2: Avoid unnecessary friction to legitimate research and education.
3. Governance actions (three options)
➡️ Option 1 — Network baseline: certification + SOP minimums
Purpose: Reduce variability in biosafety practice across distributed labs.
Design: A lightweight participation standard for labs using the platform including training checklist; Standard operating procedure (SOP) templates for handling, contamination response, waste logs and periodic documentation checks.
Assumptions: Labs will opt in if benefits are tangible and the extra admistrive work is not to burdensome.
Risks: Uneven enforcement; exclusion of under-resourced labs if standards become to complex.
➡️ Option 2 — Pigment/material safety standard: whitelist + minimal testing + labeling
Purpose: Address the most important downstream risk for the product: skin-contact, pigment safety and environmental implications.
Design: Shared “allowable pigment classes” (whitelist) plus minimum evidence requirements for testing (basic leach, washfastness, disposal guidance, documentation of lab status). Standard labeling for intended use and safety-relevant claims.
Assumptions: Low-cost testing tools or institutional partners are available; whitelist stays current and not to restrictive.
Risks: The process to complex and hindering community engagement, or weak tests gives unreliable results, slowed innovation if the whitelist narrows too far.
➡️ Option 3 — Open-source hardware standards for safe, distributed BC biofabrication
Purpose: Reduce reliance on expensive proprietary equipment while lowering barriers to participation without lowering safety. The goal is to make safe practice easier by default through standardized, well-documented hardware and workflows suitable for community labs.
Design: an open-source “reference stack” that includes:
- Validated hardware designs for core needs (e.g., enclosed growth modules with spill containment, filtered airflow concepts, light/projection enclosures to reduce eye/UV exposure, basic sensing/logging for temperature/pH proxies where appropriate).
- A documentation package: build BOMs with substitutions, maintenance/calibration checklists, cleaning/decon compatibility notes, and safety labels.
- Inter-lab benchmarking: common test artifacts and reporting templates so labs can compare performance and identify failure modes early.
Assumptions:
- Standardizing equipment and documentation will reduce accidents and variability more effectively than rules alone.
- Community labs have enough fabrication capacity (or partner access) to build/maintain hardware.
- A shared reference design can remain adaptable across different local constraints.
Risk:
- Hardware reliability varies; incomplete documentation leads to unsafe modifications; lack of maintenance causes drift in performance.
- Lowered barriers increase scale of adoption faster than training capacity; designs are copied without safety context; fragmentation into many forks undermines standardization.
4. Score
| Does the option: | Option 1 | Option 2 | Option 3 |
|---|---|---|---|
| Enhance Biosecurity | |||
| • By preventing incidents | 1 | 2 | 2 |
| • By helping respond | 1 | 2 | 2 |
| Foster Lab Safety | |||
| • By preventing incident | 1 | 2 | 1 |
| • By helping respond | 1 | 2 | 1 |
| Protect the environment | |||
| • By preventing incidents | 2 | 1 | 2 |
| • By helping respond | 2 | 2 | 2 |
| Other considerations | |||
| • Minimizing costs and burdens to stakeholders | 2 | 3 | 1 |
| • Feasibility in community labs? | 1 | 2 | 1 |
| • Not impede research | 2 | 2 | 1 |
| • Promote constructive applications | 1 | 1 | 1 |
5. Prioritization and recommendation
I would prioritize Option 1 + Option 2 as the baseline governance package, with Option 3 as a longer-term technical pathway. Option 1 provides uniform safety culture and response capacity across labs; Option 2 directly governs consumer-contact risks and environmental externalities specific to pigment-enabled textiles. Option 3 is desirable for uniformed implementation of option 1 and 2 in a community lab setting.
Primary audiences: community lab networks and lab leads (implementation), funders/partners, and local safety/environment authorities (alignment on waste and disposal practices).
ChatGBT 5.2 was used for brainstorming bioengineering ideas for BC production in a community based setting
Aswell as searching for academic literature
and correct spelling error and double checking if I understood the research correctly
and to make the code for the governance chart:
It was also used for debugging some of the problems that I had with the website build, I am not including those prompts here…
Homework Questions from Professor Jacobson
Homework Questions from Dr. LeProust:
Homework Question from George Church:
Walker, K. T., Li, I. S., Keane, J., Goosens, V. J., Song, W., Lee, K.-Y., & Ellis, T. (2025). Nature Biotechnology, 43, 345–354. https://doi.org/10.1038/s41587-024-02194-3 ↩︎
Zhou, H., Lin, P., Jeong, K. J., & Lee, S. Y. (2025). Trends in Biotechnology. https://doi.org/10.1016/j.tibtech.2025.09.019 ↩︎