Week 2 HW: dna read write and edit
Part 1: Benchling & In-silico Gel Art
My original idea was to make a circle, but after some trial and error I realized it would be a bit too complicated—so I settled on an arch (bridge).
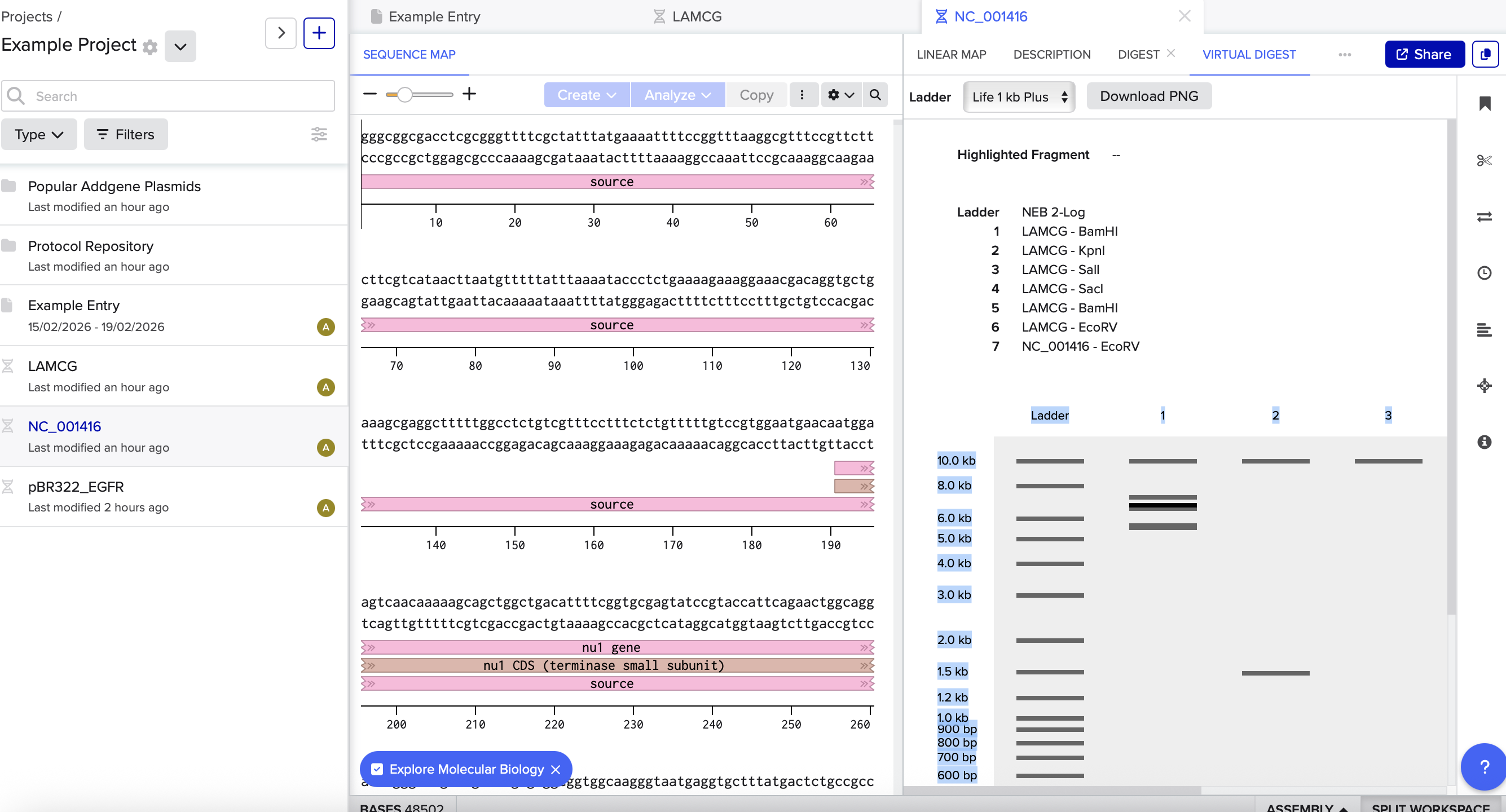
1a) I imported the sequence for lambda DNA.
1b) In Benchling, I ran all 7 restriction enzymes we had available to see which ones gave:
- a busy lane (many bands) → use as the “background” in most lanes
- a cleaner lane (fewer bands) → use to “carve out” the interior of the arch
Note:
- Lane = the vertical track DNA runs down from a single well
- Bands = the horizontal lines within a lane (different fragment sizes)
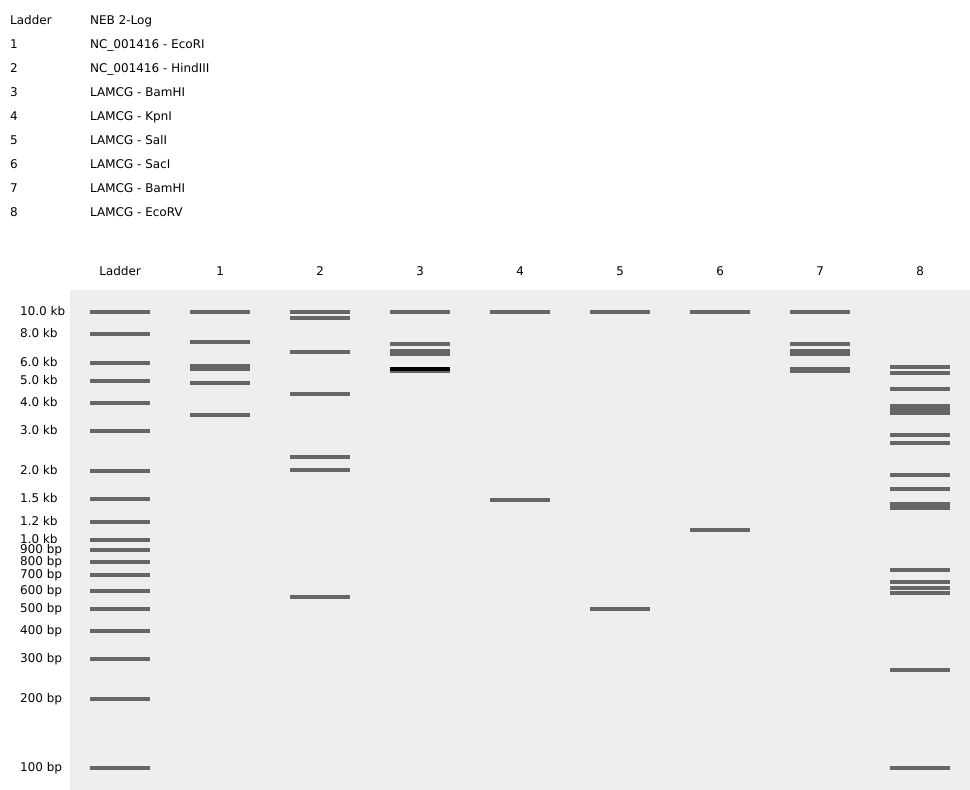
Based on the results above, I rearranged the enzymes to create the pattern:
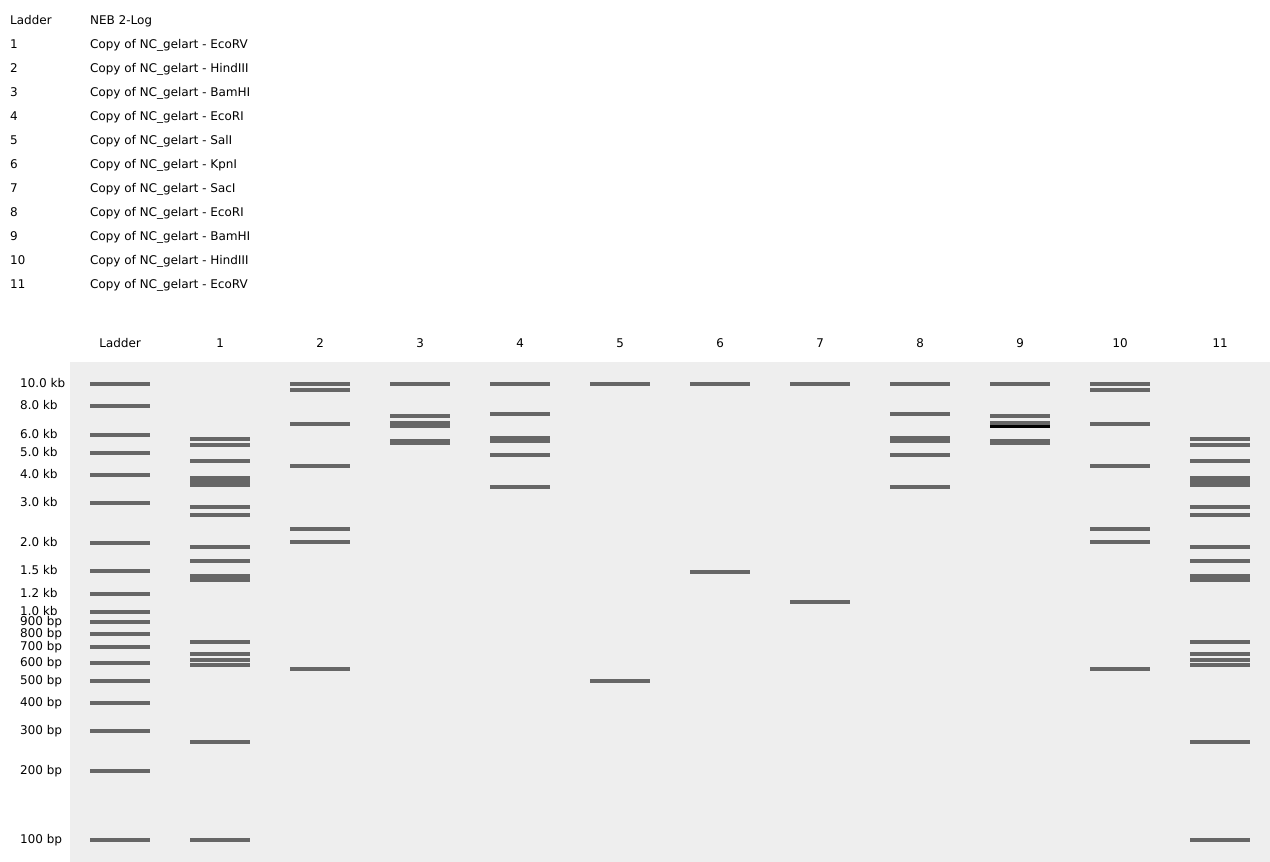


Although it’s not the most beautiful arch, this was a great exercise for understanding the basics of in-silico digests and gel band patterns.
This tool is also great for quickly iterating on gel-art layouts: https://rcdonovan.com/gel-art
3.1. Choose your protein
In recitation, we discussed picking a protein for the homework that you personally find interesting. I chose CBM3.
Why CBM3?
CBM3 is interesting because it works like a modular “cellulose anchor”: you can fuse it to other proteins so they reliably stick to cellulose (including bacterial cellulose). Beyond simple labeling, CBM fusions are used as fluorescent probes to visualize cellulose organization and dynamics, as affinity tags for low-cost purification on cellulose, and as anchoring domains to immobilize enzymes on cellulose scaffolds—turning cellulose into a reusable biocatalyst support or functional capture material.
Simply put: it’s short, often expresses well, and it sticks to cellulose.
Reference: CBM3 (example paper)
In UniProt, I searched for “carbohydrate-binding module CBM cellulose-binding protein” and got many hits. A good way to narrow the options is to pick something that is:
- Reviewed (Swiss-Prot) (more reliable annotation)
- Short / manageable (ideally ~80–250 aa)
- Clearly annotated as a CBM domain (cellulose-binding)
The UniProt entry I used was Q06851. The full protein is long, but UniProt makes it possible to extract only the domain/region relevant to the application:
- Open the UniProt entry
- Scroll to Family & Domains
- Find the feature you are interested in (domain boundaries)
I chose the CBM3 (carbohydrate-binding module family 3) from the cellulosome scaffoldin CipA, because CBM3 specifically binds cellulose and is relevant for bacterial cellulose materials.
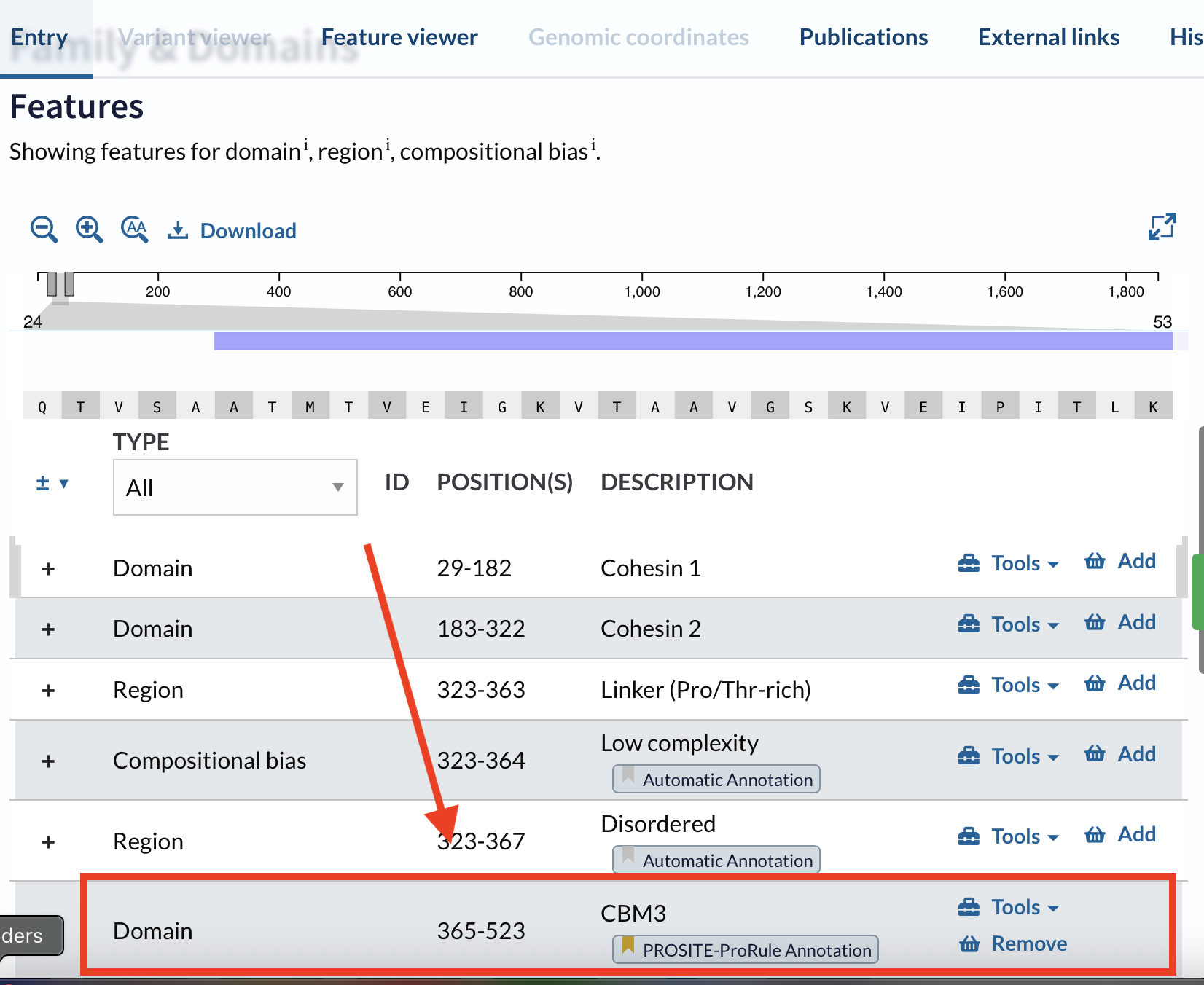

3.2. Reverse translate: Protein (amino acid) → DNA (nucleotide)
To extract only the CBM3 region, I downloaded the sequence and used the Gao Lab WebLab tool:
WebLab – range_extract_protein
I entered the range 365–523, which returned:
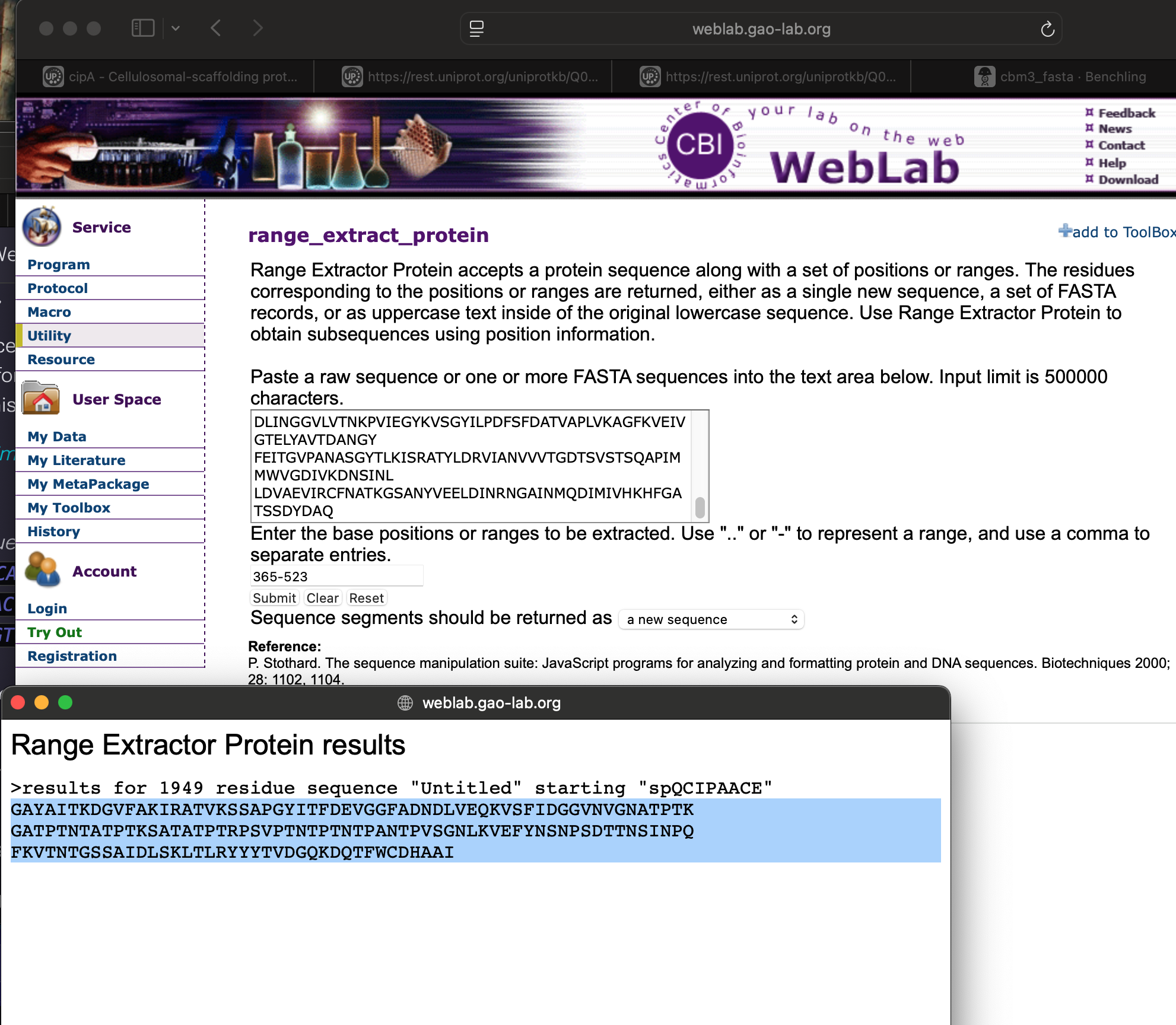

Next, I pasted the CBM3 amino-acid sequence into the Sequence Manipulation Suite reverse-translation tool: bioinformatic – Reverse Translate
Finally, I double-checked the result in Benchling by pasting the reverse-translated DNA into a new sequence and using Benchling’s Translate feature to confirm it produced the same amino-acid sequence.
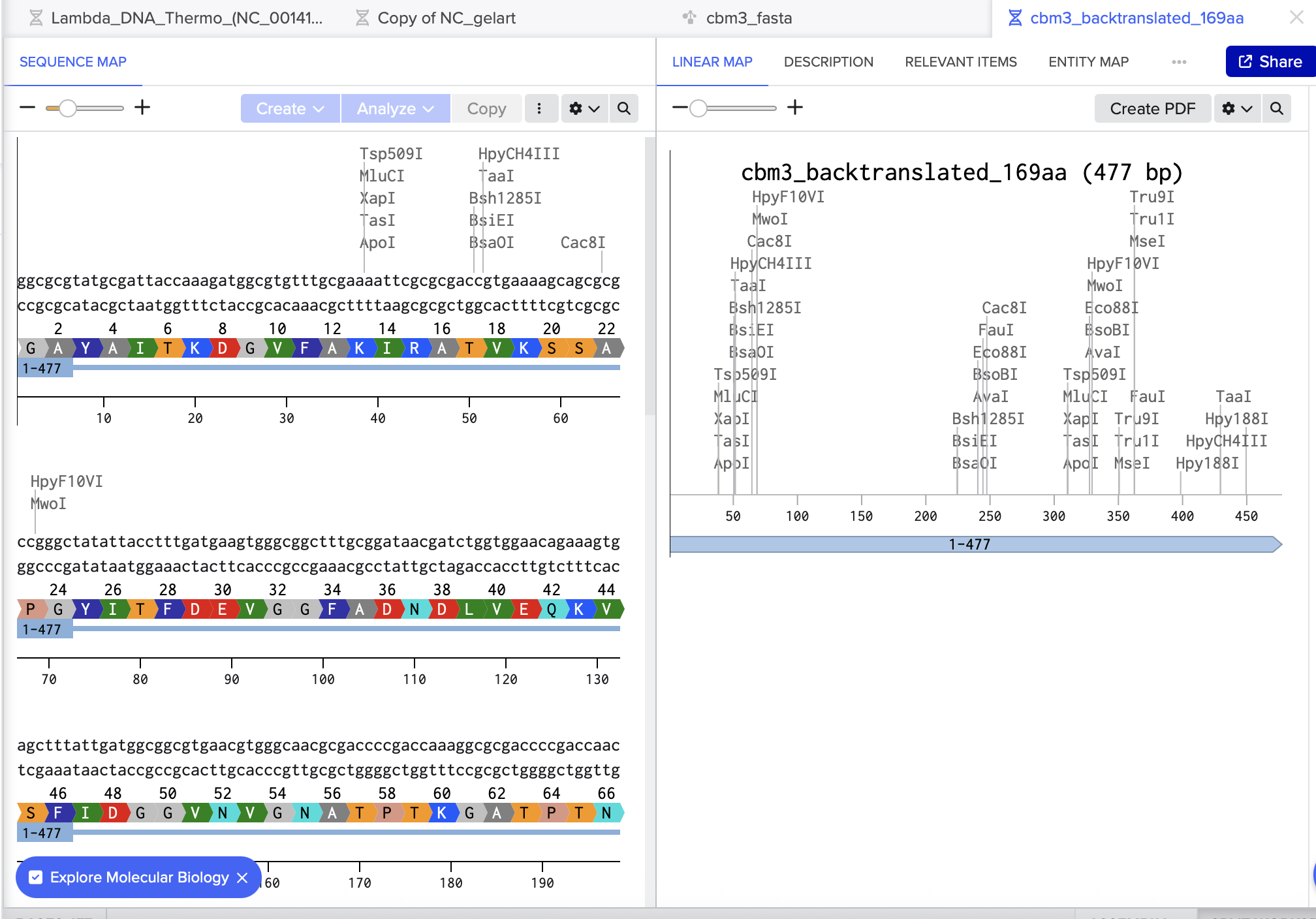

3.3. Codon optimization
I decided to codon-optimize for E. coli because it’s a common protein-expression host with well-established tools. Codon optimization matters because organisms have different codon bias / tRNA abundances, and matching preferred codons often improves translation efficiency, protein yield, and reduces stalling during expression. To do this, I used Twist’s codon-optimization workflow and selected Host: Escherichia coli. The optimization completed successfully (“Optimization was successful”) and the sequence scored Standard, indicating it is considered synthesize-able under Twist’s constraints. I then selected Use the optimized sequence and (as a sanity check) confirmed that the translated amino-acid sequence remained unchanged—only synonymous codons were swapped.
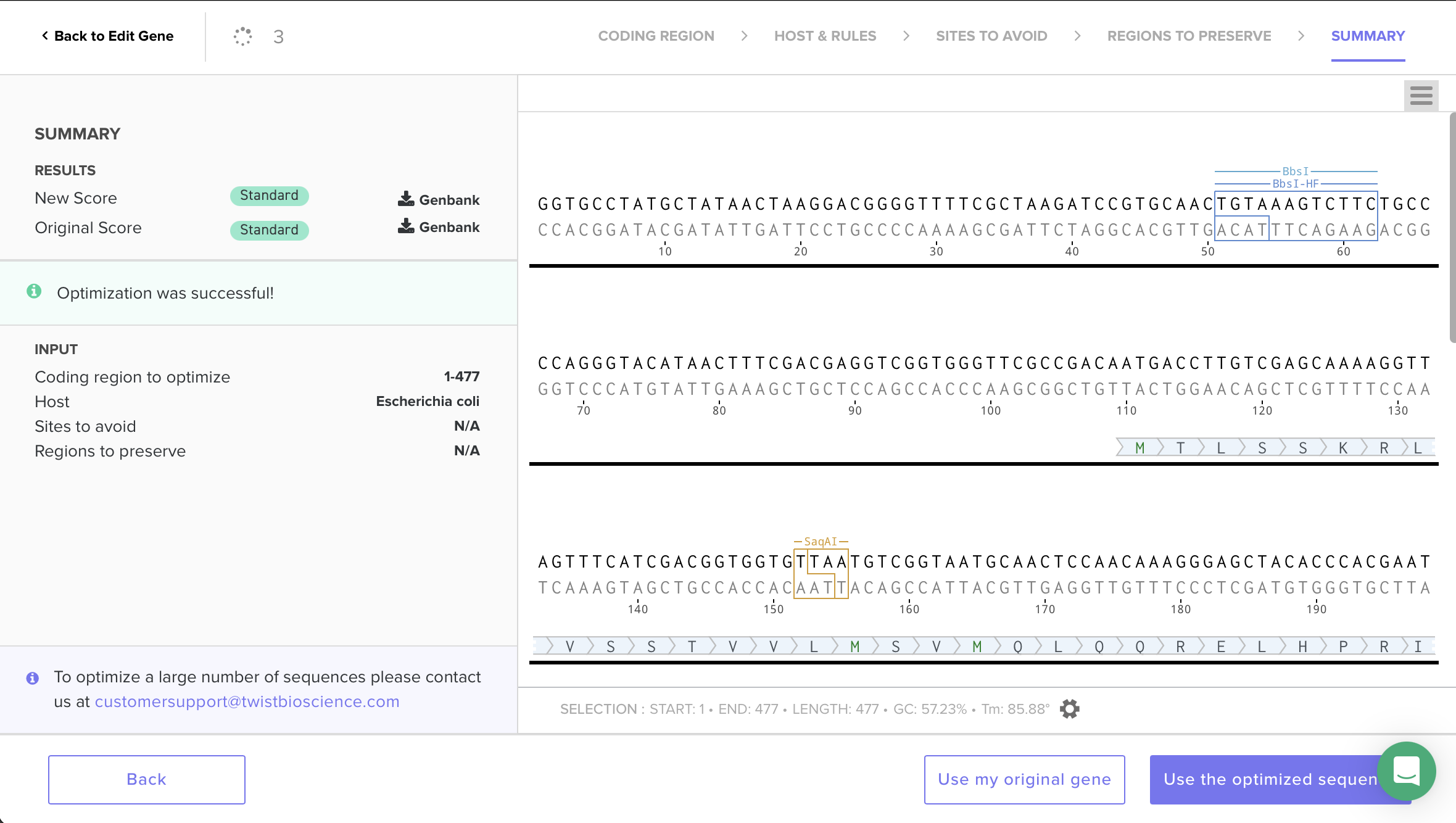

“I optimized for E. coli because it’s a common protein-expression host with well-established tools; the purified CBM can then be applied to bacterial cellulose to bind it.”
3.4. You have a sequence! Now what?
Now that I have a DNA sequence encoding CBM3, the next step is to express the protein. In a typical cell-dependent (in vivo) workflow, the codon-optimized CBM3 coding sequence is cloned into an E. coli expression plasmid under a promoter (e.g., T7/lac).
-An expression plasmid is designed to make lots of protein.
-A promoter is a DNA “on-switch” that tells the cell when to start making RNA from your gene.
-T7/lac is a common strong promoter system used to tightly control expression.
After transforming the plasmid into an expression strain, the cells are grown and expression is induced (often with IPTG).
IPTG releases repression in the lac system so the promoter becomes active, and the cells start producing CBM3.
Inside the cell, the DNA is transcribed by RNA polymerase into mRNA, and the mRNA is then translated by ribosomes into the CBM3 protein as tRNAs deliver amino acids according to the codons. The protein can then be purified (for example via an affinity tag such as His-tag) and used to bind/functionalize bacterial cellulose.
-His-tag lets you purify CBM3 using a matching resin (Ni-NTA), washing away everything else.
Alternatively, CBM3 could be produced using a cell-free expression system (TX-TL), where the DNA template (plasmid or linear) is added directly to a lysate containing RNA polymerase, ribosomes, and all required cofactors.
required cofactors: -RNA polymerase
-ribosomes
-tRNAs, amino acids
-energy + cofactors
In this setup the same steps—transcription to mRNA and translation to protein—happen in a test tube rather than inside living cells, which can be faster and easier for prototyping, though often at smaller scale.
Why do cell-free?
- Often faster for prototyping (no transformations, no growing cells).
- Convenient when testing multiple designs quickly.
- Downsides: usually more expensive per mg and often smaller scale/yield than growing E. coli.
Ethical and regulatory difference: Cell-free systems are generally considered safer because they are non-living reactions that cannot usually replicate or spread in the environment. They stop once substrates, energy, or cofactors are depleted. In contrast, in-cell genetic engineering uses living organisms, which can continue growing and may pose risks if accidentally released, such as persistence in the environment or transfer of engineered DNA to other organisms.
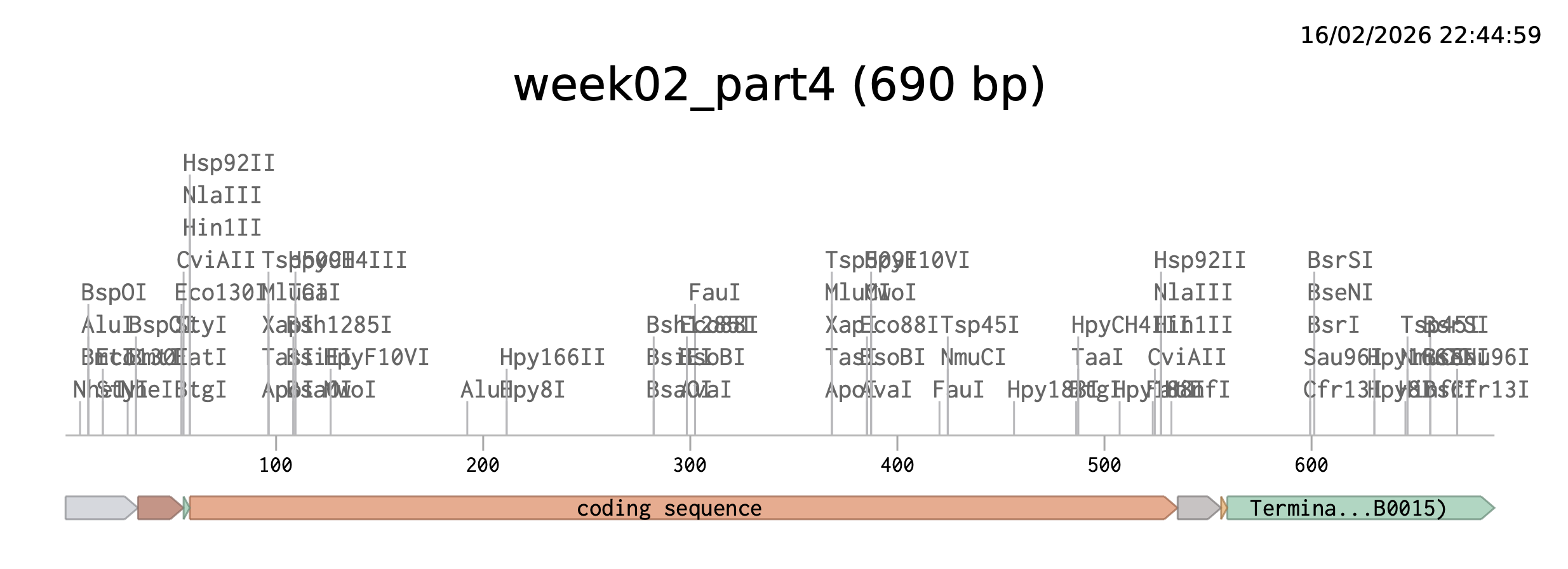
Part 4 — Build an E. coli expression cassette (Benchling → Twist-ready)
For this step I designed a complete E. coli expression DNA insert in Benchling by assembling the required genetic parts in the correct order:
- Promoter (BBa_J23106)
- RBS (BBa_B0034 + spacer)
- Start codon (ATG)
- Coding sequence: replaced the template CDS with my codon-optimized gene (from Part 3)
- C-terminal His-tag (7×His)
- Stop codon (TAA)
- Terminator (BBa_B0015)
After pasting each piece, I annotated every region (promoter, RBS, start, CDS, His-tag, stop, terminator) directly on the Benchling sequence.
I also used Benchling’s Analyze/Translate to confirm the ATG (Open Reading Frame) is in frame from the ATG (Start codon) and that the sequence ends with the His-tag followed by a stop codon.
Benchling link: https://benchling.com/s/seq-YgFm33VIxUzvPdZpyOKk?m=slm-lYfXGHAomlD9Go7bgPWh
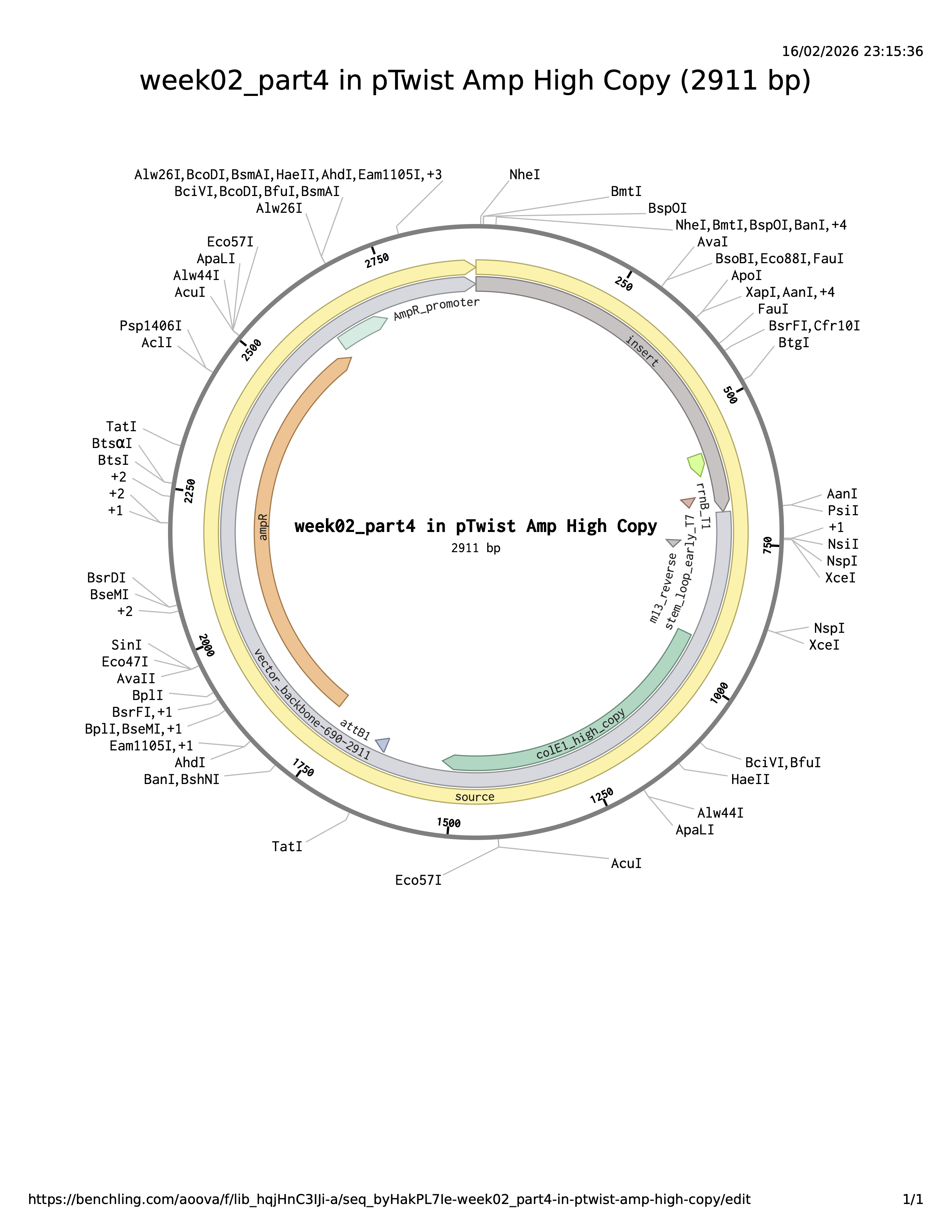

The plasmid backbone is the original vector framework containing essential elements such as the antibiotic resistance marker and origin of replication. The insert is the DNA fragment cloned into that backbone. The source annotation usually refers to the origin or overall sequence record and is not typically a functional genetic element itself.
In conclusion
- E. coli = the factory
- plasmid backbone = the delivery vehicle / operating template inside the factory
- insert = the custom cargo you added
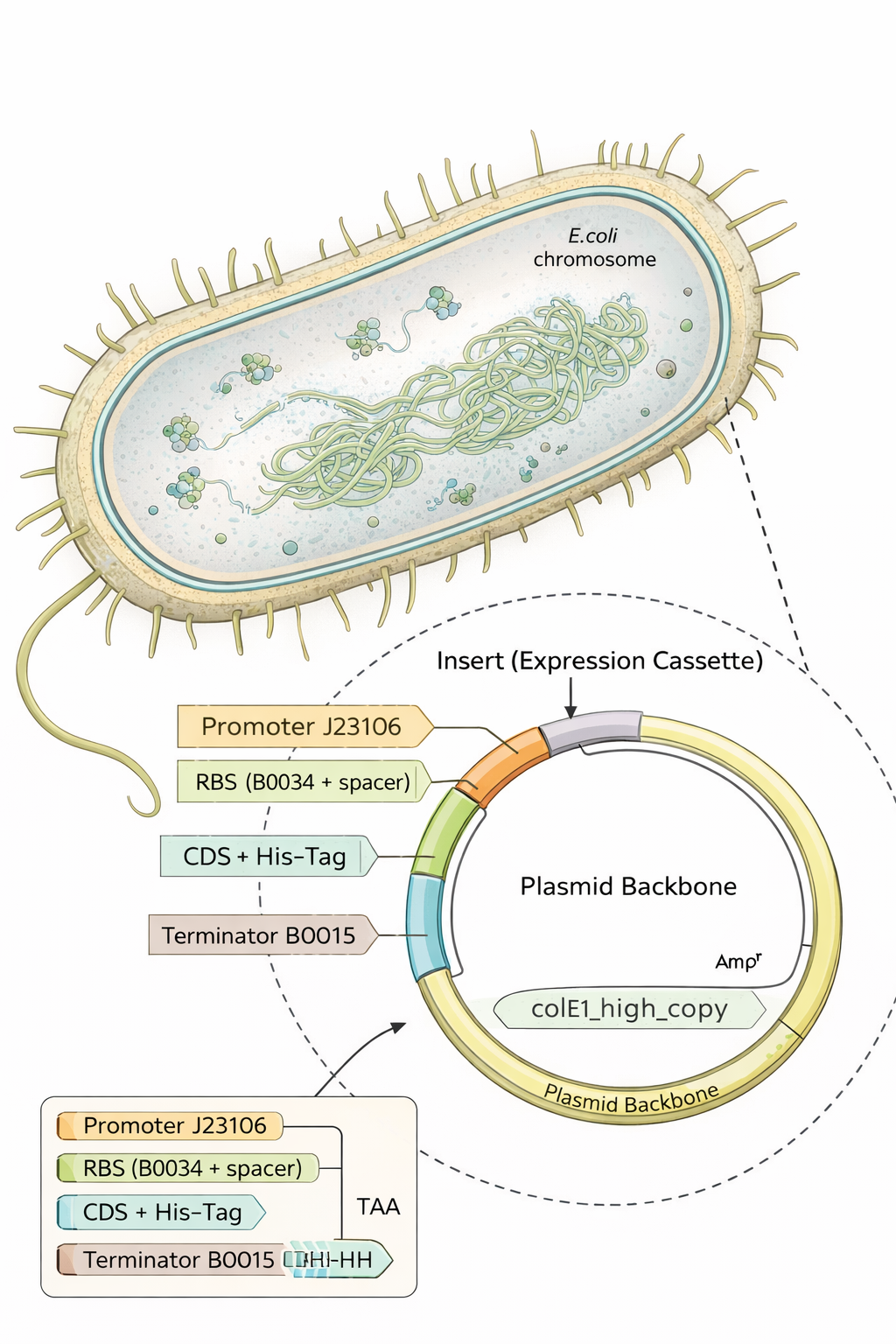

Part 5 — DNA Read / Write / Edit (pigment-colored SCOBY / bacterial cellulose sheets)
This builds directly on my Week 1 project idea (“Programmable colors for bacterial cellulose production”):
https://pages.htgaa.org/2026a/alve-lagercrantz/homework/week-01-hw-principles-and-practices/index.html
5.1 DNA Read (sequencing)
(ii) What sequencing technology would you use and why?
Because SCOBY is a mix of different types of DNA (bacteria, yeast etc) I would use Oxford Nanopore long-read sequencing with shotgun metagenomic DNA from the SCOBY. One run can tell me both who is present (community composition) and help reconstruct full plasmids/inserts, which matters for checking stability during long fermentations.
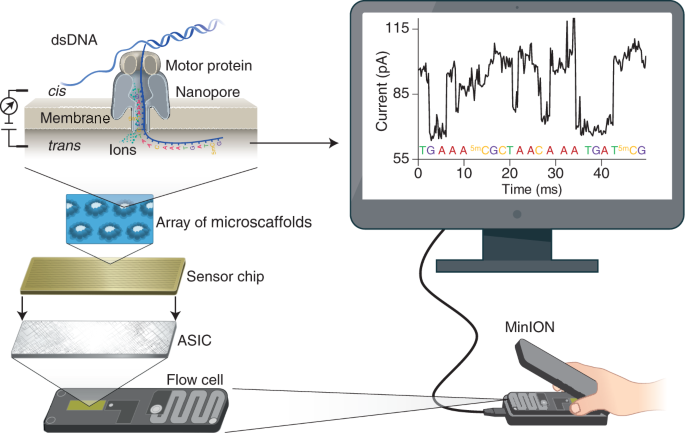

- Generation: Third-generation (single-molecule, long-read sequencing).
- Input: Total genomic DNA extracted from the SCOBY (mixed community DNA).
- Essential prep steps: Extract DNA carefully (aim for high molecular weight) → optionally size-select / gently shear if needed → ligate Nanopore adapters (or use rapid prep) → load on flow cell.
- How bases are decoded (base calling): DNA passing through a nanopore changes the ionic current; a basecaller converts the signal into A/C/G/T sequences.
- Output: FASTQ (reads + quality scores) (often plus raw signal files) → downstream: taxonomic profiling + assembly to recover plasmids/contigs and verify constructs.
5.2 DNA Write (synthesis)
The Part 4 cassette I built is an E. coli expression-style design (promoter/RBS/terminator suited for E. coli). To make color, I can keep the same cassette architecture but swap the coding sequence to a pigment gene (or pathway). For SCOBY/BC specifically, there are two realistic “write” directions:
In-situ pigmentation inside the cellulose producer
Engineer a cellulose-producing Komagataeibacter strain to biosynthesize pigment while it grows the pellicle. A strong example is melanin via tyrosinase expression, which yields dark, robust coloration in BC.1Co-culture / division-of-labor pigmentation
Keep the cellulose producer focused on making BC, and pair it with a second microbe engineered to produce pigments (broad palette). A published example uses E. coli strains producing violacein derivatives and carotenoids alongside Komagataeibacter xylinus to generate multiple BC colors.2
Important design note: If the target host is Komagataeibacter (not E. coli), the regulatory parts (promoters/RBS/terminators, plasmid backbone) must be chosen for that host; otherwise the pigment genes may not express even if the coding sequence is correct.
Material/safety note (relevant for textiles/skin contact):
- Some pigments (e.g., violacein) are bioactive, so “write” decisions should also consider leaching, irritation risk, and safe handling/disposal pathways. 3
5.3 DNA Edit (genome editing)
For stable, repeatable colored BC (especially over long growth periods), genome editing can be attractive because it can:
- reduce dependence on plasmid maintenance,
- improve stability across generations,
- enable more predictable performance in a mixed or semi-open fermentation context.
Conceptually, “edit” could mean integrating a pigment function into the cellulose-producer genome, or tuning regulatory control (e.g., linking pigment production to growth phase or light-patterning concepts used in engineered living materials).
Bonus — a bacterial-cellulose (BC) face mask that changes color via cell-free pigment expression
BC is already a compelling cosmetic substrate because it holds a lot of water, conforms well to skin, and has been tested as a moisturizing sheet mask material. In one evaluation, a single application of a bacterial-cellulose mask increased facial skin moisture more than a moist towel control.4
Instead of putting living engineered cells on the face, a safer “synthetic biology” route is to embed freeze-dried cell-free gene expression (TX-TL) into the BC sheet as small patterned “sensor dots.” These cell-free circuits stay inactive when dry, then turn on when the mask hydrates during wear; outputs can be colorimetric (visible) or optical.5
Because freeze-dried cell-free circuits activate upon rehydration, a conventional pre-hydrated sheet mask would trigger prematurely during storage. A practical design might be a dry-stored BC mask (or a separate paper sensor tab) that is activated only at time of use by releasing fluid.
How it could work:
- Input (skin/sweat biomarker): pH (skin barrier/irritation proxy), lactate (sweat/metabolic proxy).
- Sensing layer (cell-free circuit): a biomarker-responsive regulatory element controls whether a reporter is expressed.6
- Output (visible color): express a chromoprotein (strong color under normal light) so the mask visibly shifts color in specific zones without any instrument; chromoproteins are attractive for “naked-eye” readouts.7

Why this is interesting for BC masks:
- The mask provides hydration + intimate contact, which can reactivate freeze-dried cell-free systems.
- Patterning multiple “dots” enables a simple visual map (e.g., pH zones at cheeks vs T-zone), turning the mask into a wearable readout rather than just a carrier.
[^^1][^3]
References (footnotes)
Walker, K. T. et al. Self-pigmenting textiles grown from cellulose-producing bacteria with engineered tyrosinase expression. Nature Biotechnology (2025, published online 2024). https://doi.org/10.1038/s41587-024-02194-3 ↩︎
Zhou, H. et al. One-pot production of colored bacterial cellulose. Trends in Biotechnology (2025). https://doi.org/10.1016/j.tibtech.2025.09.019 ↩︎
WEEK 1 HW: PRINCIPLES AND PRACTICES https://pages.htgaa.org/2026a/alve-lagercrantz/homework/week-01-hw-principles-and-practices/index.html ↩︎
Amnuaikit, T. et al. (2011). Effects of a cellulose mask synthesized by a bacterium on facial skin characteristics and user satisfaction. https://pmc.ncbi.nlm.nih.gov/articles/PMC3417877/ ↩︎
Nguyen, P.Q. et al. (2021). Wearable materials with embedded synthetic biology sensors for biomolecule detection. Nature Biotechnology. https://www.nature.com/articles/s41587-021-00950-3 ↩︎
Ba, F. et al. Chromoproteins: visible tools for advancing synthetic biology. https://pubmed.ncbi.nlm.nih.gov/41309430/ ↩︎
Pardee, K. et al. (2014). Paper-Based Synthetic Gene Networks. Cell. https://pubmed.ncbi.nlm.nih.gov/25417167/ ↩︎