Week 03 — Opentrons: Automation Art + Post-Lab Questions
Part 1 — Automation Art (OT-2 “printing” a design)
This week I designed a microscope icon as “automation art” and converted it into a grid of XY dot coordinates that can be dispensed by the Opentrons OT-2 onto an agar plate.
1) Design → coordinate map
I started from the course Automation Art Interface, which makes it easy to draw a dot pattern on a circular “canvas.”
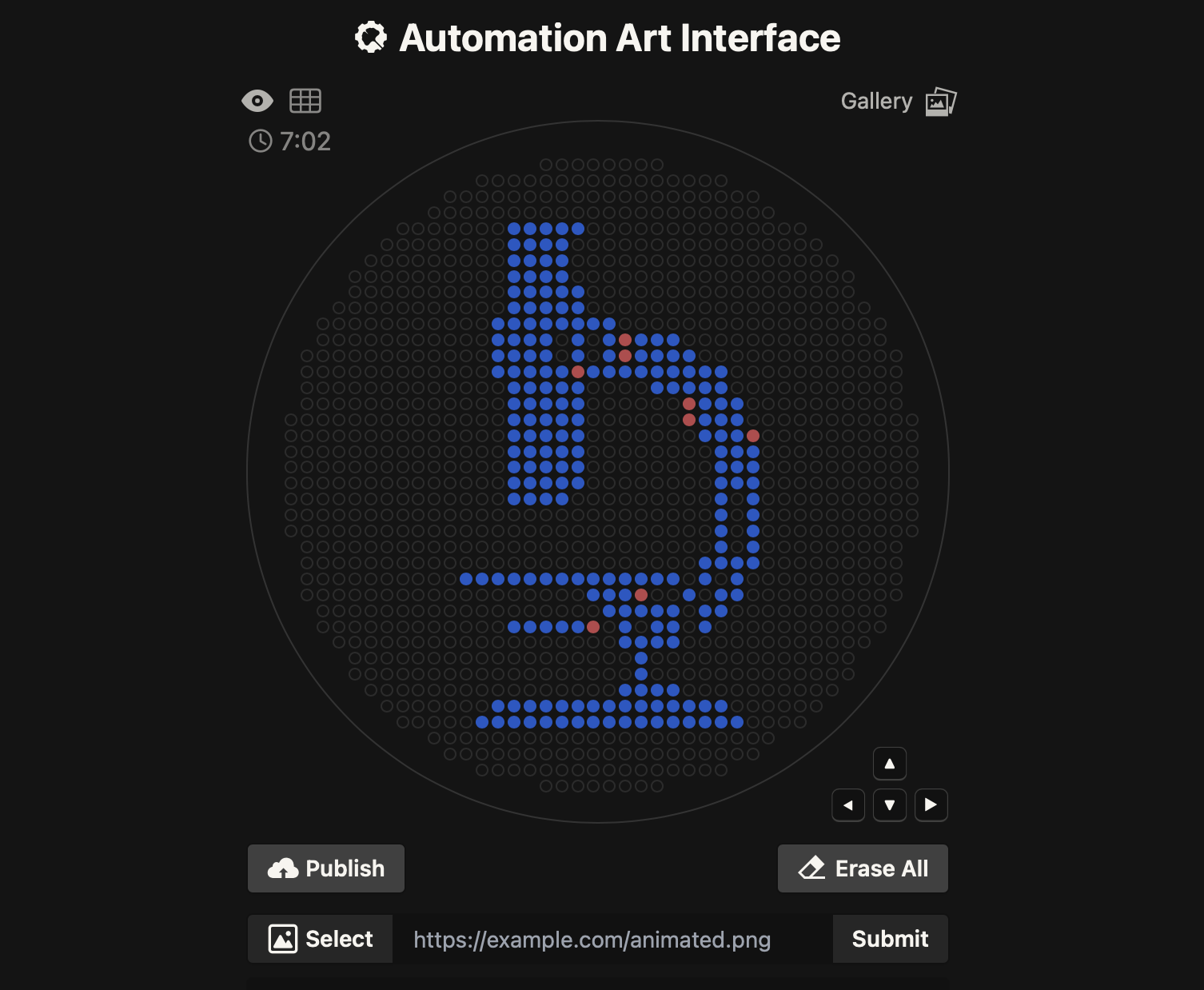

2) Convert the pattern into points + sanity-check in Python
To avoid trial-and-error on the robot, I used a Colab notebook to:
- convert pixels/dots → (x, y) coordinate lists
- preview the design as a scatter plot
- separate two colors (main shape vs highlights)
Colab notebook:
https://colab.research.google.com/drive/1tLENS2Rs0mxdN-pJp5QNfm1K6dfg9xsS?usp=sharing
The preview below shows the final point-map I used:
- Green = main “microscope” body
- Red = highlight/accent points (mScarlet)
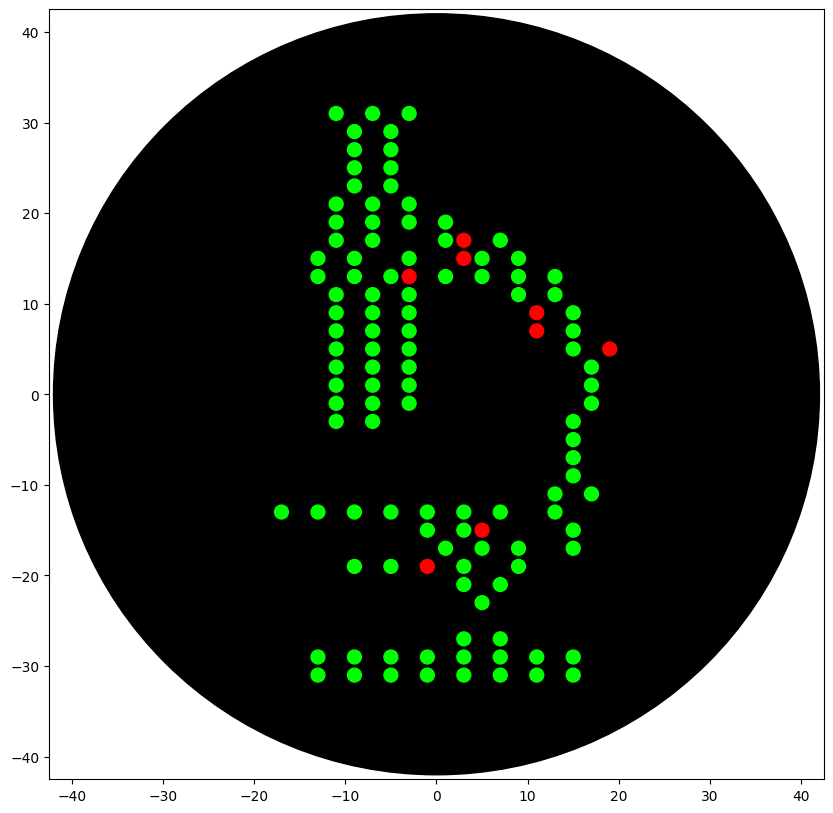

3) Implement in an OT-2 protocol
In my OT-2 protocol, the key idea is:
- store the design as coordinate lists (e.g.,
electra2_points,mscarlet_i_points) - aspirate enough volume for a “chunk” of dots (so we don’t aspirate for every single point)
- dispense each dot using a small helper that moves down to dispense and back up to detach the droplet cleanly
Snippet (from my protocol):
Part 2 — Post-Lab Questions (Opentrons paper + how it connects to my final project)
2.1 A published paper using Opentrons for a novel bio application
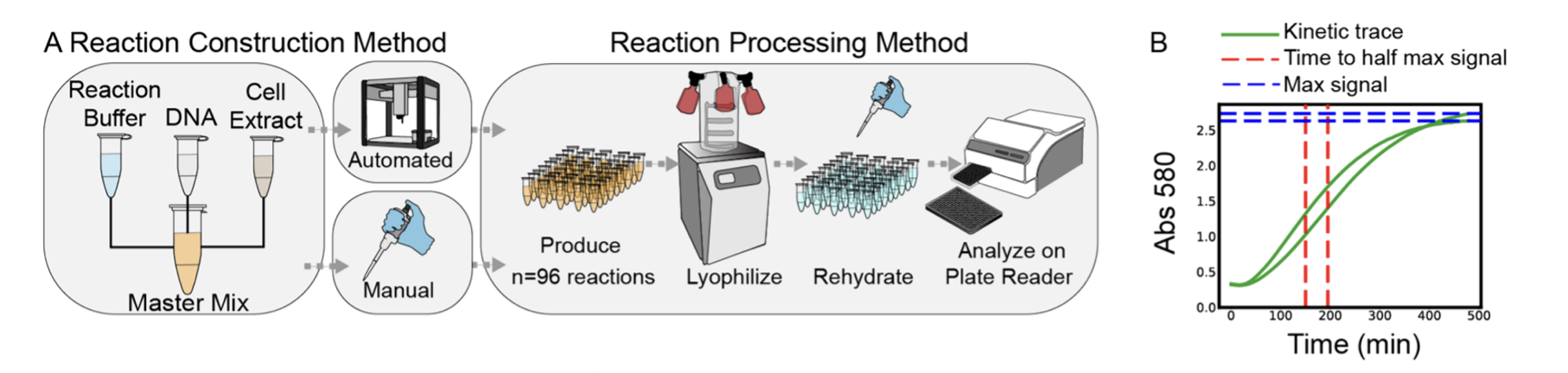
I chose Brown et al. (2025), “Semiautomated Production of Cell-Free Biosensors” (ACS Synthetic Biology) because it shows the OT-2 being used not just for “routine liquid handling,” but as a manufacturing platform for synthetic biology diagnostics.
In the paper, the authors use an Opentrons OT-2 to assemble large batches of cell-free biosensor reactions, then process them through a deployment-style pipeline: assemble → (optionally) lyophilize → rehydrate → measure output. They compare manual vs automated preparation and demonstrate reliable, scaled production (including a full 384-well plate format), which is exactly the kind of reproducibility you want when moving from “cool demo” to “repeatable product”.
2.2 How Opentrons could be “perfect” for producing a BC skincare sheet mask (pouch mask)
For my final project direction, I’m thinking of a skincare sheet mask, using bacterial cellulose (BC) as the carrier material. The OT-2 is a great fit because it turns a “handmade one-off” into a repeatable, batchable fabrication workflow.
Where OT-2 helps most
- Standardized loading of serum / actives: dispense precise volumes of humectants (e.g., glycerol), buffers, preservatives (if used), fragrance-free additives, etc. into pouches or soaking trays so every mask gets the same dose.
- Patterned deposition (“pixel printing”) onto BC: print micro-spots or zones of different formulations (e.g., soothing zone vs brightening zone) or a visible “QC pattern” to confirm even loading.
- Built-in controls + QC: include calibration spots or a reference color patch on each sheet (so each mask is self-verifiable in documentation/photos).
How this connects to the Brown et al. OT-2 paper Brown et al. use the OT-2 as a manufacturing platform for cell-free biosensor reactions (assemble → process → rehydrate → readout). My mask workflow is conceptually similar, just with a different substrate:
- assemble formulations (or cell-free mixes for R&D prototypes)
- deposit onto/into BC in a controlled way
- package / dry / store
- rehydrate on use (when the sheet mask is applied)
What I would document as “automation value”
- Repeatability across a batch (mass gain of BC after dosing, or volume dispensed per pouch)
- Uniformity (image-based check of a printed pattern across masks)
- Optional: a simple visual indicator that activates upon rehydration (e.g., a time/usage indicator patch for R&D proof-of-concept)
This makes the OT-2 useful not only for lab experiments, but for building a small-scale manufacturing pipeline for BC skincare sheet masks.

Reference
- Brown, D. M. et al. (2025). Semiautomated Production of Cell-Free Biosensors. ACS Synthetic Biology. DOI: 10.1021/acssynbio.4c00703
Links (for citation / screenshots):
Final Project Ideas
Idea 1 — OT-2 “manufactured” BC skincare sheet masks (pouch masks)
Concept: Use the Opentrons OT-2 as a small-scale manufacturing tool to reproducibly load / pattern skincare formulations onto bacterial cellulose (BC) sheet masks that come in a sealed pouch and sit on skin for ~1–2 hours.
Problem: BC have excelant water holding capacity however handmade BC sheet masks are hard to standardize (dose, uniformity, repeatability across a batch).
Hypothesis: Automation + coordinate-based dispensing can turn BC sheet masks into a consistent, documented “biofabrication pipeline.” bacteria can be engineered to “read” your skin health and express it in simple color cues.
embed a cell-free color indicator patch as a “time / health/ hydration indicator.
Approach (R&D workflow):
- Grow/harvest BC sheets → press to target thickness → load into a deck jig/holder.
- OT-2 dispenses exact volumes of serum/actives into:
- (A) the pouch (soak method), and/or
- (B) directly onto the BC in patterns/zones (“forehead zone”, “cheek zone”, etc.).
MVP demo: 6–12 masks with identical dosing; photo + mass-gain and uniformity checks.
What to measure: repeatability (dispensed volume, BC mass gain), uniformity (image analysis), user-facing consistency (feel, tack, wetness over time).
Idea 2 — Water-resistant BC “leather” via in-growth synbio
Concept: Reduce BC water uptake during growth by programming the system to deposit a cellulose-bound amphiphilic layer (e.g., a hydrophobin–cellulose binding domain fusion) that self-assembles on/within the BC network.
Problem: When using BC as leather substitude (material production) one of the main problems is that it absorbs a lot of water + swells; tradtionally the solution have been different post-coatings different oils or waxes however they tend to not be very long lasting.
Hypothesis: A cellulose-binding, self-assembling protein layer produced during growth period can reduce wetting and wicking without heavy post-treatment.
Approach:
- Engineer a production strain or a modular functionalization step to present hydrophobin–CBD/CBM at the BC interface.
- Compare conditions:
- control BC
- BC + in-process hydrophobin–CBD functionalization
- BC + conventional post-coat (baseline comparison)
MVP demo: small “bag panel” swatch set + simple rain/soak tests.
What to measure: water uptake %, wicking height, thickness change after wetting, flex/crack after dry–wet cycles.
Stretch goal: combine with in-growth pigment or optogenetic patterning for functional + aesthetic “self-finished” BC.
Idea 3 — Light-input → color-output BC bio-print for moiré effects (BC + engineered E. coli)
This project is based on week01 homework


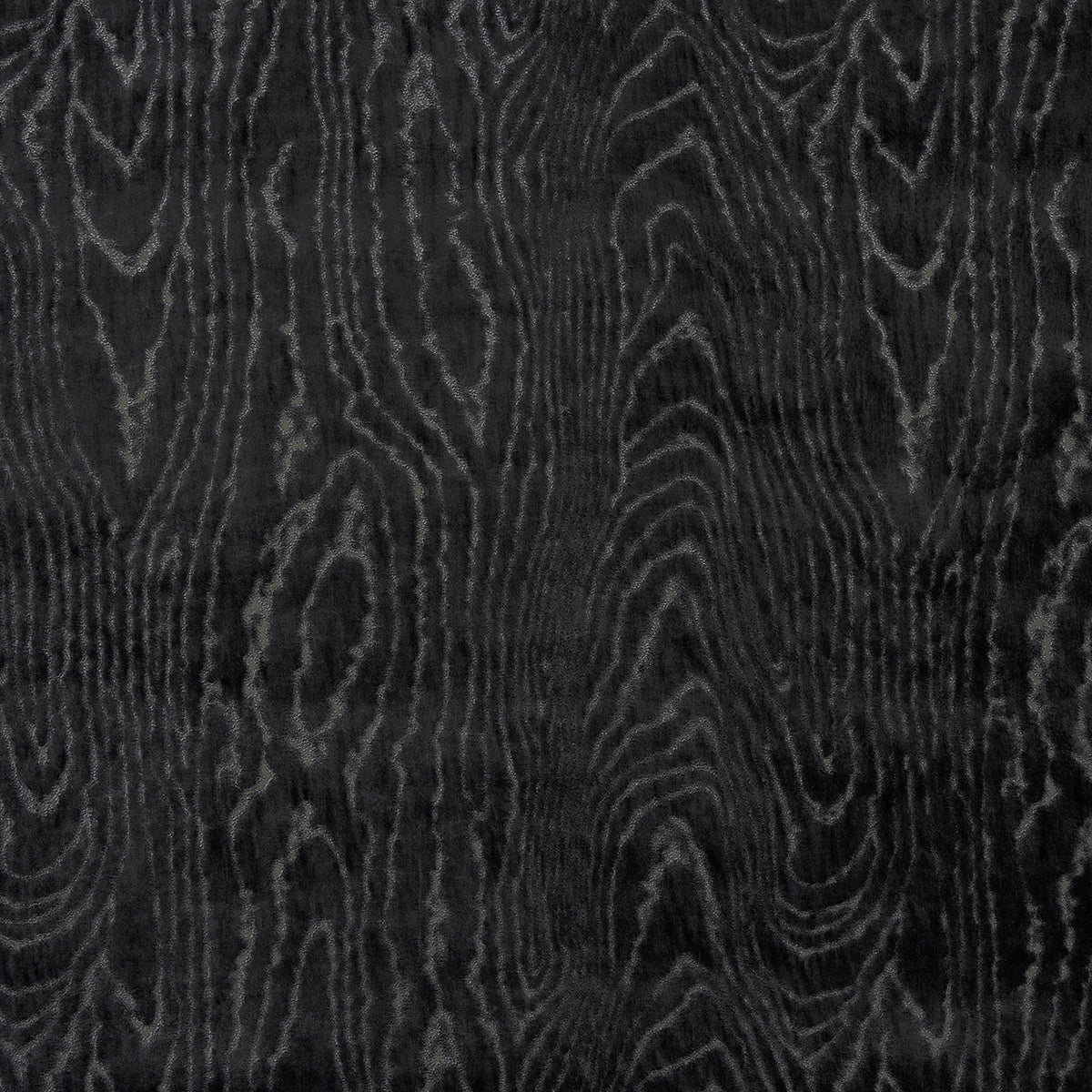
Concept: A co-culture “living printer”: Komagataeibacter grows the BC sheet while engineered E. coli produces pigments under light control, enabling projected patterns. Two patterned layers with slightly different line frequencies create moiré interference when stacked.
- Problem: Dyeing BC is slow/uneven; patterning usually requires post-processing.
- Hypothesis: Optogenetics enables spatial control: light patterns → localized gene expression → localized color on/within a growing material.
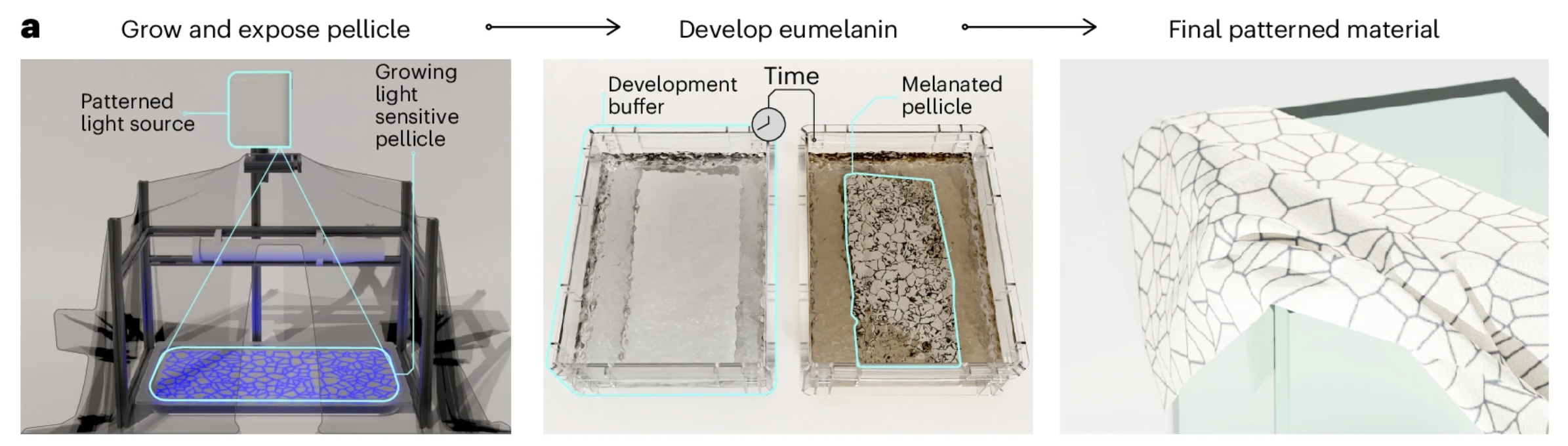


- Approach (research plan):
- Build/borrow a light-gated expression system in E. coli (red/green/blue input).
- Drive a visible output (pigment pathway or chromoprotein).
- Pattern with projector/photomask onto a co-culture or onto E. coli deposited on BC.
- Grow/prepare two sheets with slightly offset gratings → overlay for moiré visuals.
- MVP demo: one light-patterned colored sheet + photo documentation of resolution/contrast.
- What to measure: pattern sharpness (edge blur), color contrast, stability after drying, moiré strength with layer overlay.
- Stretch goal: multi-color “logic-like” prints (different wavelengths → different pigments).


Reff. project 1:
- Brown, D. M. et al. Semiautomated Production of Cell-Free Biosensors. ACS Synthetic Biology (2025). https://pubmed.ncbi.nlm.nih.gov/40073441/
- Amnuaikit, T. et al. (2011). Effects of a cellulose mask synthesized by a bacterium on facial skin characteristics and user satisfaction. https://pmc.ncbi.nlm.nih.gov/articles/PMC3417877/
- Nguyen, P.Q. et al. (2021). Wearable materials with embedded synthetic biology sensors for biomolecule detection. Nature Biotechnology. https://www.nature.com/articles/s41587-021-00950-3
- Pardee, K. et al. (2014). Paper-Based Synthetic Gene Networks. Cell. https://pubmed.ncbi.nlm.nih.gov/25417167/
- Ba, F. et al. Chromoproteins: visible tools for advancing synthetic biology. https://pubmed.ncbi.nlm.nih.gov/41309430/
Reff. project 2:
- Puspitasari, N. Class I hydrophobin fusion with cellulose binding domain… (PDF thesis/report, 2021). https://repositori.ukwms.ac.id/id/eprint/31910/1/1-Class_I_hydrophobin_fusion_with_%28Nathania%29.pdf
Reff. project 3:
- Walker, K. T., Li, I. S., Keane, J., Goosens, V. J., Song, W., Lee, K.-Y., & Ellis, T. (2025). Nature Biotechnology, 43, 345–354. https://doi.org/10.1038/s41587-024-02194-3
- Zhou, H., Lin, P., Jeong, K. J., & Lee, S. Y. (2025). Trends in Biotechnology. https://doi.org/10.1016/j.tibtech.2025.09.019
- Levskaya, A. et al. Synthetic biology: engineering Escherichia coli to see light. Nature (2005). https://pubmed.ncbi.nlm.nih.gov/16306980/