week-06-hw-genetic-circuits-part-i
Assignment: DNA Assembly
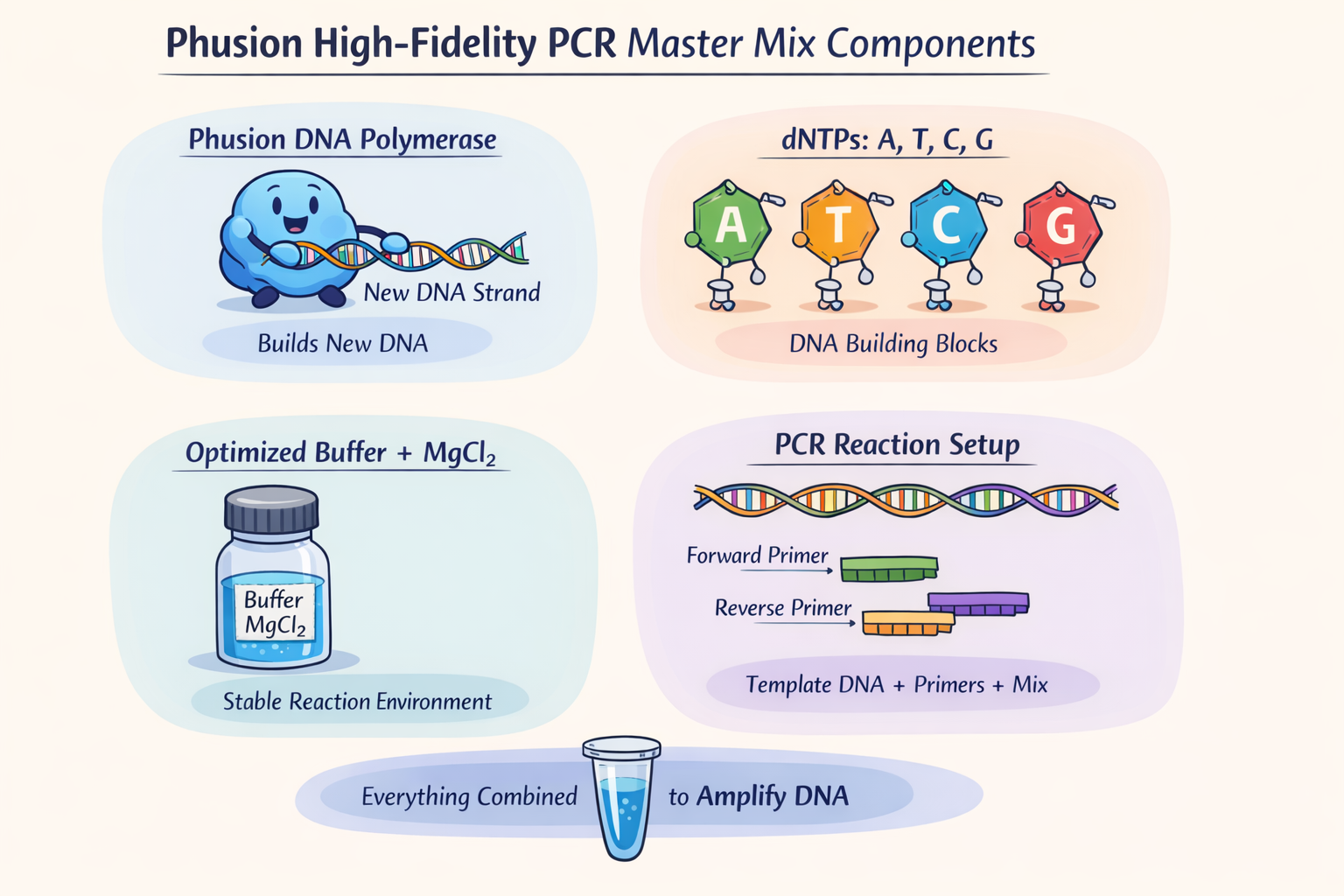
1. What are some components in the Phusion High-Fidelity PCR Master Mix and what is their purpose?
The Phusion High-Fidelity PCR Master Mix contains at least three key components: Phusion DNA polymerase, deoxynucleotides (dNTPs), and an optimized reaction buffer that includes MgCl₂. The polymerase is the enzyme that synthesizes new DNA strands during PCR, the dNTPs are the nucleotide building blocks incorporated into the new DNA, and the buffer/MgCl₂ provide the chemical environment and cofactor needed for efficient polymerase activity. According to the website of (New England)[https://www.neb.com/en/products/m0531-phusion-high-fidelity-pcr-master-mix-with-hf-buffer?srsltid=AfmBOorWPUiBMtKsQJJH0VLGPzLYHtMYELtt0wf7AQB0YZYF4nrTfFsz] the main benefit of rgw Master mix is high fidelity (50X comparing to Taq) and fast extension times.
2. What are some factors that determine primer annealing temperature during PCR?
The main factor is the melting temperature (Tm) of the primers. Tm depends on the primer’s sequence length, and base composition.
- Base composition: GC primers generally bind more strongly than AT-rich ones and therefor require higher Tm.
- sequence length: Longer sequences tend to bind better since their is more base pairs that can bind to eachother.
Good primer pairs should usually have Tms that are close to each other. The lab notes suggest a binding-region Tm around 52–58°C and within about 5°C of the partner primer, and annealing is chosen about 2–5°C below the lower primer Tm. Reaction conditions also matter; for example, additives such as DMSO can lower primer Tm, so the annealing temperature may need to be reduced. In our lab protocol the backbone PCR and insert PCR use different annealing temperatures (57°C vs 53°C) because the primer sets differ.
3. There are two methods from this class that create linear fragments of DNA: PCR, and restriction enzyme digests. Compare and contrast these two methods, both in terms of protocol as well as when one may be preferable to use over the other.
PCR creates a linear DNA fragment by using primers and a DNA polymerase to amplify a chosen region through repeated cycles of denaturation, annealing, and extension. Its biggest advantage is flexibility: it can amplify almost any desired region and can also add useful sequence features through the primers, such as mutations, overlaps for Gibson assembly, or restriction sites. That makes PCR preferable when a fragment must be engineered, when no convenient restriction sites exist, or when only a small defined region should be copied.
A restriction digest, by contrast, creates linear DNA by cutting at specific recognition sequences with restriction enzymes. This is often simpler and very reliable when the needed sites are already present in the plasmid or multiple cloning site, and it is especially useful for subcloning, plasmid linearization, or diagnostic digests. Its limitation is that it depends on sequence context: the enzyme sites must be present where you need them and absent where you do not want cuts. So in practice, restriction digestion is often preferable when the construct already has good enzyme sites, while PCR is preferable when you need more freedom in fragment boundaries or sequence design.
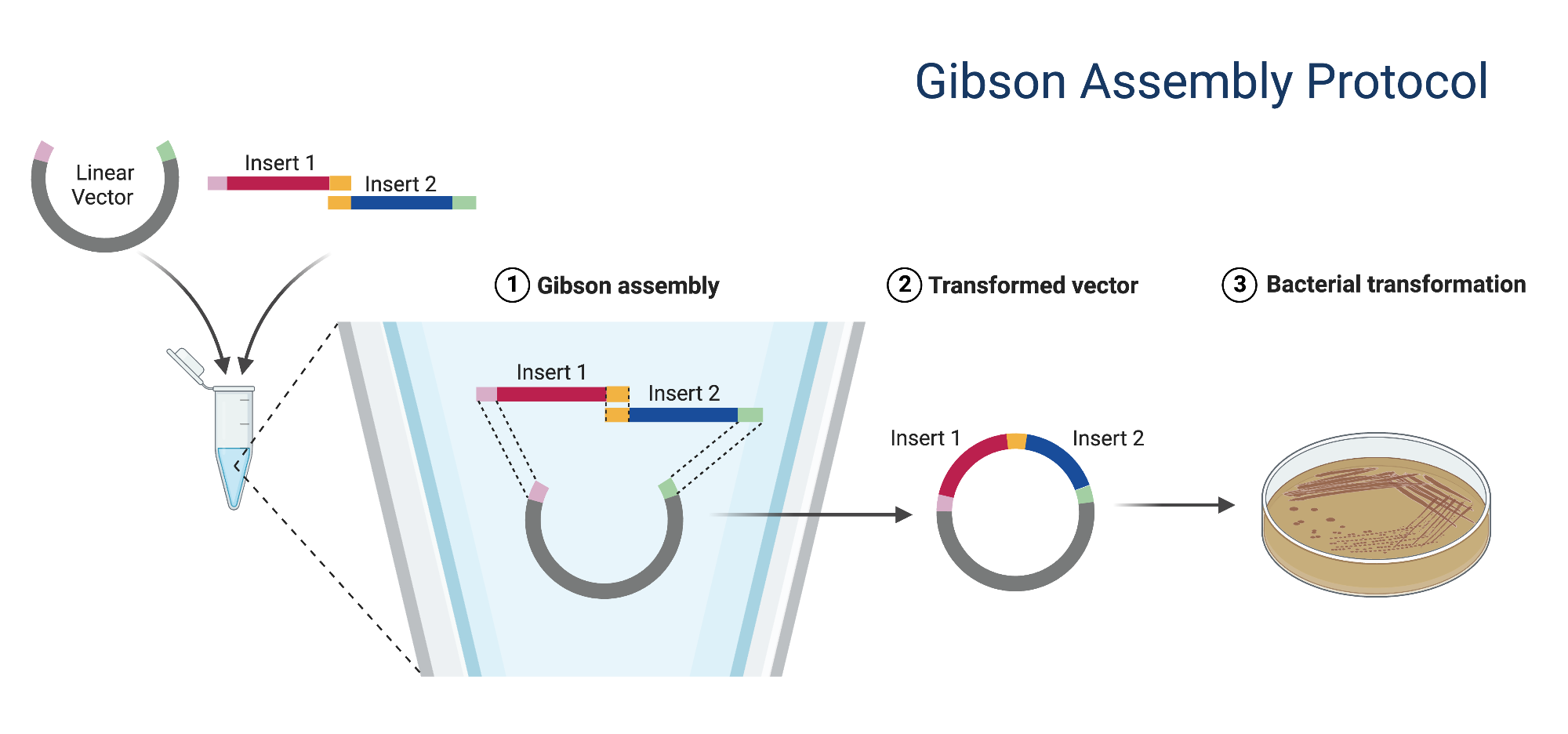
4. How can you ensure that the DNA sequences that you have digested and PCR-ed will be appropriate for Gibson cloning?
For Gibson assembly, the most important requirement is that adjacent DNA fragments have matching homologous overlaps (similar in position). In the lab, the primer design guidelines specify about 20–22 bp overlaps.


Beyond design, you should verify the fragments experimentally. In this protocol, that means using DpnI (a restriction enzyme that cuts methylated DNA at the sequence GATC) to remove methylated parental plasmid template after PCR, purifying the PCR products, checking DNA concentration, and running a diagnostic gel to confirm that the backbone and insert have the expected sizes. For the assembly itself, the lab recommends an approximately 2:1 insert:vector molar ratio, which also helps improve successful Gibson cloning. It is also possible to confirm the whole assembly in silico in Benchling before doing the wet lab step, to make sure the overlap sequences are exact and nothing missing.
About (Gibsom assembley)[https://www.youtube.com/watch?v=tlVbf5fXhp4]
5. How does the plasmid DNA enter the E. coli cells during transformation?

Plasmid DNA enters the E. coli cells by heat-shock transformation of chemically competent cells. The cells are kept on ice with the DNA, then briefly exposed to 42°C, which causes the membrane to become transiently permeable. The lab handout explains this as the membrane “opening up,” after which the plasmid enters the cells by diffusion. The cells are then allowed to recover in SOC medium for about an hour so they can repair their membranes and begin expressing the antibiotic-resistance marker before they are plated on selective agar.
SOC medium is a growth medium for bacteria
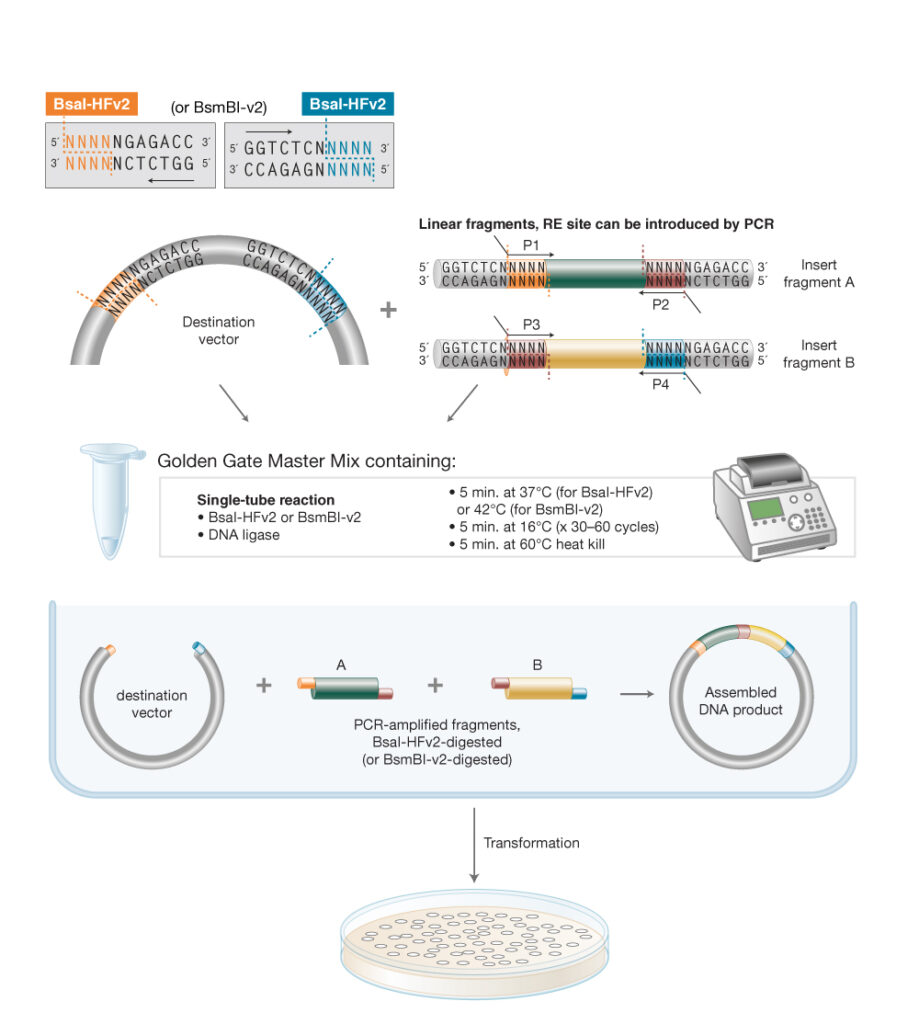
6. Describe another assembly method in detail: Golden Gate Assembly
Golden Gate Assembly is a DNA assembly method that uses a Type IIS restriction enzyme such as BsaI or BsmBI together with T4 DNA ligase in a single reaction. Unlike standard restriction enzymes, Type IIS enzymes cut outside their recognition sequence, so the researcher can design custom overhangs that determine exactly which fragments join to each other. Because the recognition sites are placed so they are removed during assembly, the final product is usually scarless and cannot be re-cut in the same way, which allows digestion and ligation to happen in the same tube. This makes Golden Gate especially useful for assembling multiple fragments in a defined order, such as promoter–RBS–CDS–terminator constructs in synthetic biology. A major design requirement is that the parts must not contain unwanted internal sites for the Type IIS enzyme being used; if they do, the sequence must be “domesticated” first. Compared with Gibson, Golden Gate is excellent for modular, repeatable multi-part assembly, while Gibson is often more convenient when overlaps are easier to design than restriction-site architecture.
Simple diagram of Golden Gate Assembly: