Week 07 HW: Genetic circuits part ii
Assignment Part 1: Intracellular Artificial Neural Networks (IANNs)
What advantages do IANNs have over traditional genetic circuits, whose input/output behaviors are Boolean functions?**
Intracellular artificial neural networks (IANNs) have a major advantage over traditional Boolean genetic circuits because they can process graded, continuous signals rather than only treating inputs as ON/OFF states. In biological systems, many relevant signals such as metabolite concentration, RNA abundance, stress level etc are not naturally binary. Neural-network-like circuits are better suited to integrate these analog inputs and make decisions based on their combined strength. Rizik e.g 2022
A second advantage is that IANNs can implement more flexible and complex computations such as classification, soft majority decisions, analog-to-digital conversion, and multistage signal processing. Rizik (2022) show multilayer “perceptgene” circuits that compute a soft majority function, perform analog-to-digital conversion, and implement a ternary switch and argue that neuro-inspired circuit design can be more reliable, resource-efficient, and reconfigurable for different tasks.
A third advantage is better compatibility with biological noise and nonlinearity. The same paper reports that logarithmic-domain neuromorphic computing is more suitable than a linear-domain perceptron for their gene circuits, and that it is more robust to noise at low signal concentrations. This is important because intracellular environments are noisy and variable from cell to cell. In that sense, IANNs are often better matched to real biological computation than rigid Boolean logic alone.
Overall, Boolean circuits are useful when a strict yes/no rule is enough, but IANNs are more powerful when the task requires integrating multiple imperfect signals, weighting them differently, and producing a graded or thresholded response.
2. Describe a useful application for an IANN; include a detailed description of input/output behavior, as well as any limitations an IANN might face to achieve your goal.
One useful application for an IANN would be a cell-state classifier for targeted cancer detection or therapy. The idea would be to engineer a mammalian cell circuit that reads several intracellular biomarkers at once, such as microRNA levels, stress-response signals, or hypoxia-associated signals, and then decides whether the overall profile matches a diseased cell state. Synthetic biology has already been used to build multi-input circuits for identifying specific cancer cells, and broader synthetic signal-processing systems are being developed for diagnostics and therapies. Z. Xie 2011
In this application, the inputs would be several intracellular markers, for example: high miR-21, high miR-155, low activity from a tumor-suppressor-associated pathway, and a hypoxia-related signal. Instead of applying a strict Boolean rule such as “all markers must be present,” the IANN would assign different effective weights to each input. In the first layer, each biomarker would influence production of a regulatory RNA or protein. In a hidden layer, those intermediate signals would be combined into a weighted internal score. In the output layer, if the total score crosses a threshold, the circuit could activate an output such as GFP for detection, a therapeutic protein, or a kill-switch effector. This makes the system more tolerant of noisy or partially matching disease signatures.
The main limitation is that real cells have limited shared resources for transcription and translation. Synthetic genes compete for these resources, which can make otherwise separate modules interfere with one another and cause the actual circuit behavior to differ from the intended design. This is a serious issue for multilayer circuits because each additional node increases the load on the cell.
A second limitation is orthogonality and crosstalk. Endoribonuclease-based platforms are powerful because they are modular and composable, but not every regulator is perfectly orthogonal with every other one. The PERSIST platform Di Adreth 2022 showed that most endoRNases were orthogonal, but some pairs showed cross-reactivity and should be avoided. That means a practical IANN needs careful part selection and calibration.
A third limitation is that large intracellular neural circuits are still difficult to scale. The neuromorphic computing paper notes that these systems support only a limited number of distinct inputs, and multilayer gene circuits can also face issues such as slow dynamics, variability between cells, and tuning difficulties. So while the concept is powerful, achieving a reliable therapeutic IANN would require careful optimization and validation.
4. Diagram for an intracellular multilayer perceptron
Left / Layer 1: X1 = DNA encoding endoRNase 1 Middle / Hidden layer: DNA encoding endoRNase 2 with mRNA 2 containing target site for endoRNase 1 Right / Output layer: DNA encoding fluorescent protein with mRNA 3 containing target site for endoRNase 2
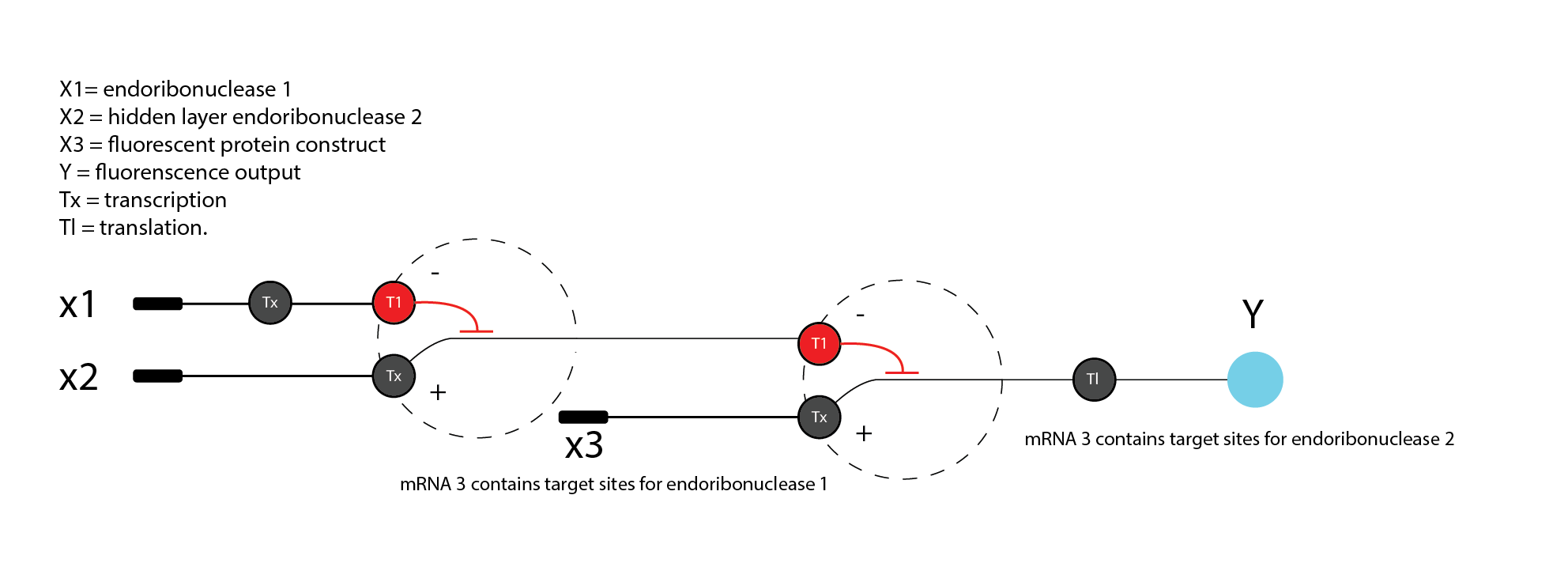

-X1 = DNA encoding endoribonuclease -Layer 1 output = endoribonuclease protein -X2 = DNA encoding fluorescent protein -Layer 2 = reporter transcript/protein regulated by the endoribonuclease from layer 1 -Y = fluorescence
References
HTGAA Spring 2026, Week 7 — Genetic Circuits Part II: Neuromorphic Circuits. Course assignment page listing the Part 1 questions and the multilayer perceptron drawing task.
https://2026a.htgaa.org/2026a/course-pages/weeks/week-07/index.htmlRizik, L. et al. (2022). Synthetic neuromorphic computing in living cells. Nature Communications, 13, 5602.
https://www.nature.com/articles/s41467-022-33288-8DiAndreth, B. et al. (2022). The PERSIST platform provides programmable RNA regulation using CRISPR endoRNases. Nature Communications, 13, 2582.
https://www.nature.com/articles/s41467-022-30172-3Gao, Y., Wang, L., and Wang, B. (2023). Customizing cellular signal processing by synthetic multi-level regulatory circuits. Nature Communications, 14, 8415.
https://www.nature.com/articles/s41467-023-44256-1Frei, T. et al. (2020). Characterization and mitigation of gene expression burden in mammalian cells. Nature Communications, 11, 4641.
https://www.nature.com/articles/s41467-020-18392-xXie, Z. et al. (2011). Multi-input RNAi-based logic circuit for identification of specific cancer cells. Science, 333(6047), 1307–1311.
https://www.science.org/doi/10.1126/scisignal.4189ec246Assignment Part 2: Fungal Materials
Fungi are eukaryotic organisms, meaning they belong to the same broad domain as animals and plants, but they form their own biological kingdom. This group includes yeasts, molds, and mushrooms. Unlike bacteria and archaea, fungi have complex cells with a nucleus. Their unique growth behavior, especially through filamentous networks called mycelium, has made them highly interesting for biomaterial research.
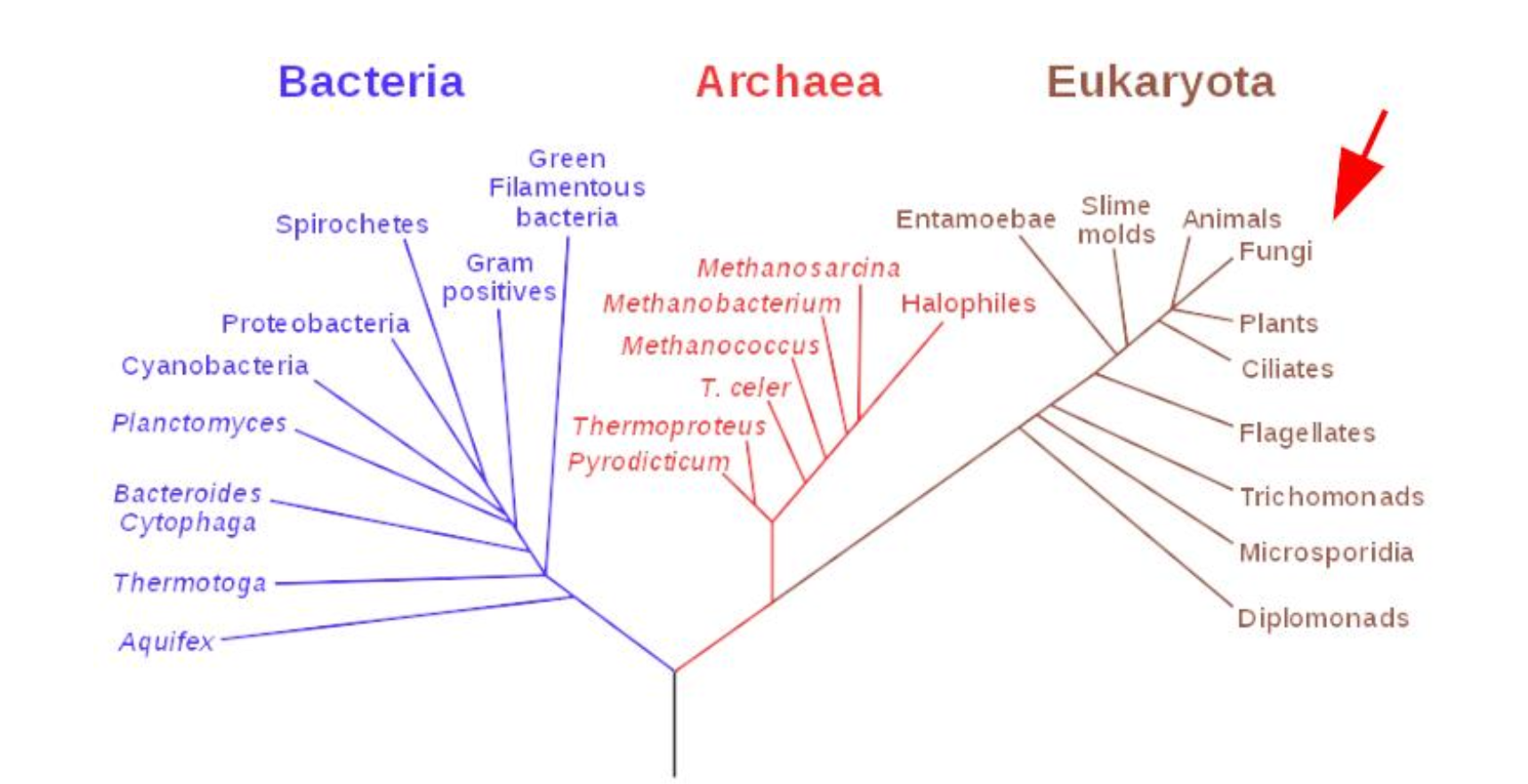

Mycology is the branch of biology concerned with the study of fungi and their many roles and applications, including:
- pathogenic activity
- drug discovery
- ecology
- bioremediation
- biomaterials
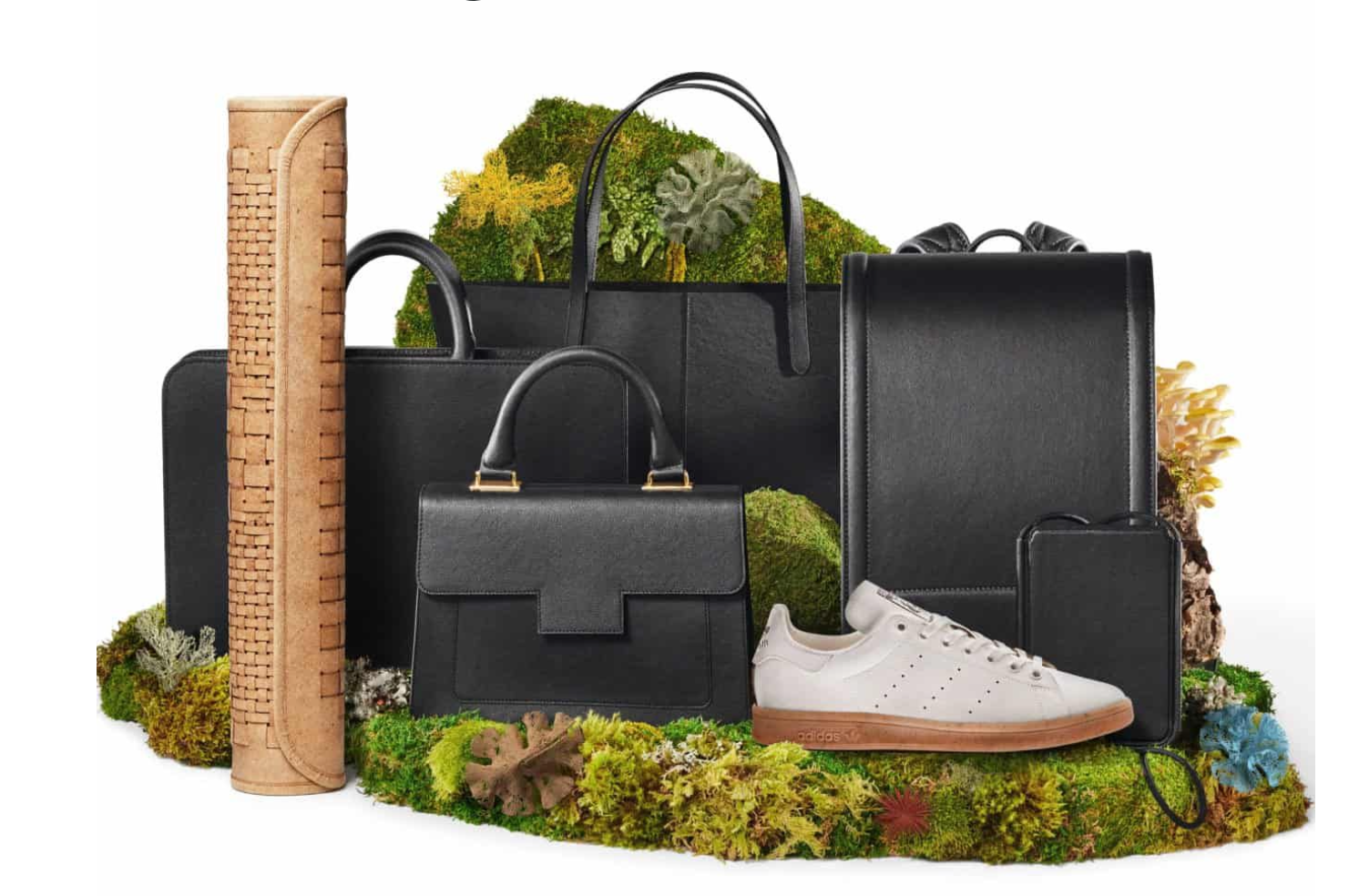
In the context of material design, the most important part of the fungus is often the mycelium, the root-like vegetative network that grows through a substrate. In recent years, mycelium has been widely explored as a biomaterial for packaging, construction, insulation, acoustic panels, and leather-like alternatives for fashion.
A major reason for this interest is that fungal materials can be grown on cheap and abundant feedstocks, such as sawdust, straw, wood chips, or other agricultural waste. They are also attractive because they are generally lightweight, biodegradable, and relatively fast to cultivate compared with many conventional manufacturing processes.
The material chart below suggests that mycelium composites often behave more like foams or lightweight natural materials than like dense polymers, ceramics, or metals. This makes them especially promising where low weight, cushioning, insulation, or biodegradability are more important than very high structural strength.
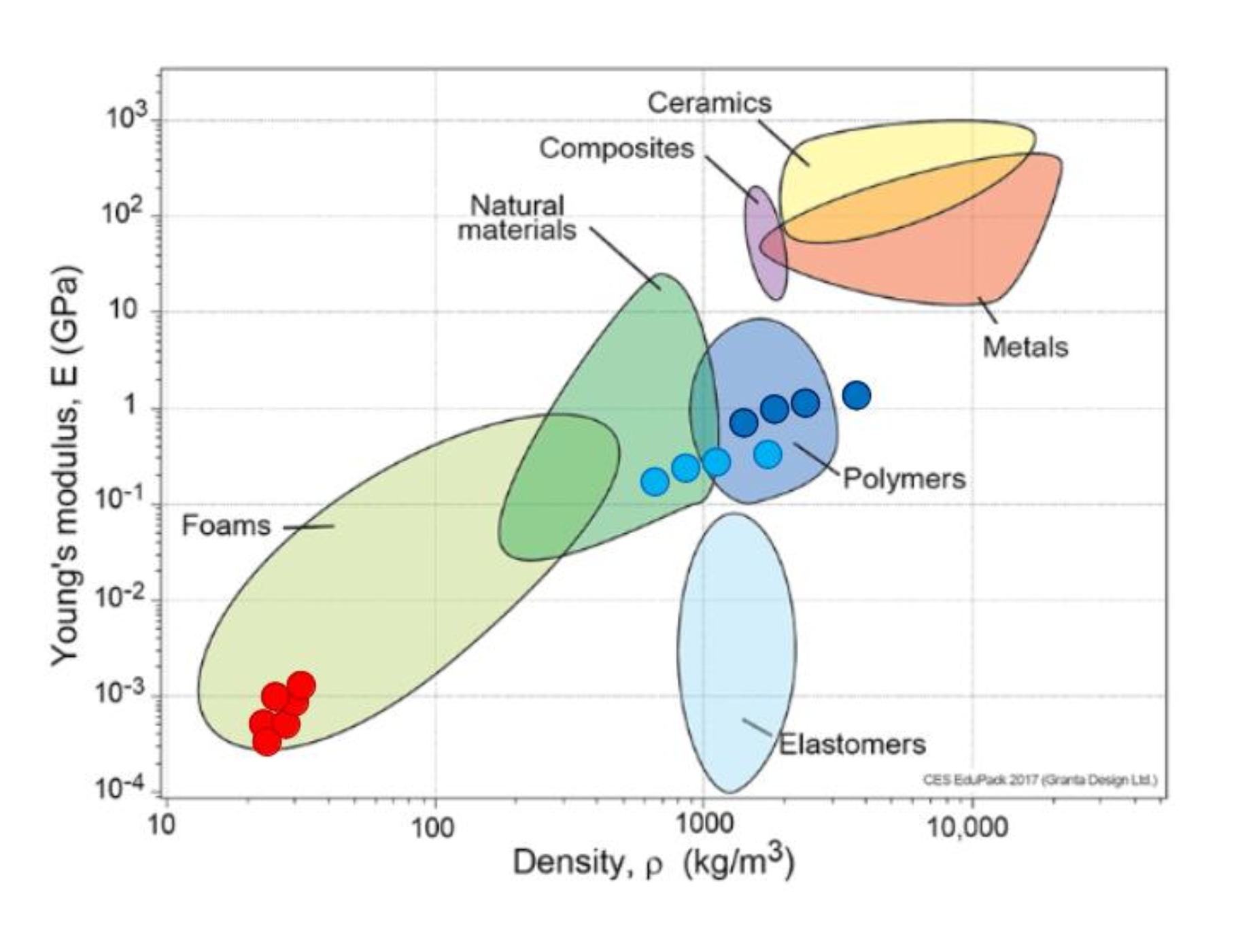

red dots are mycelium
Examples of existing fungal materials
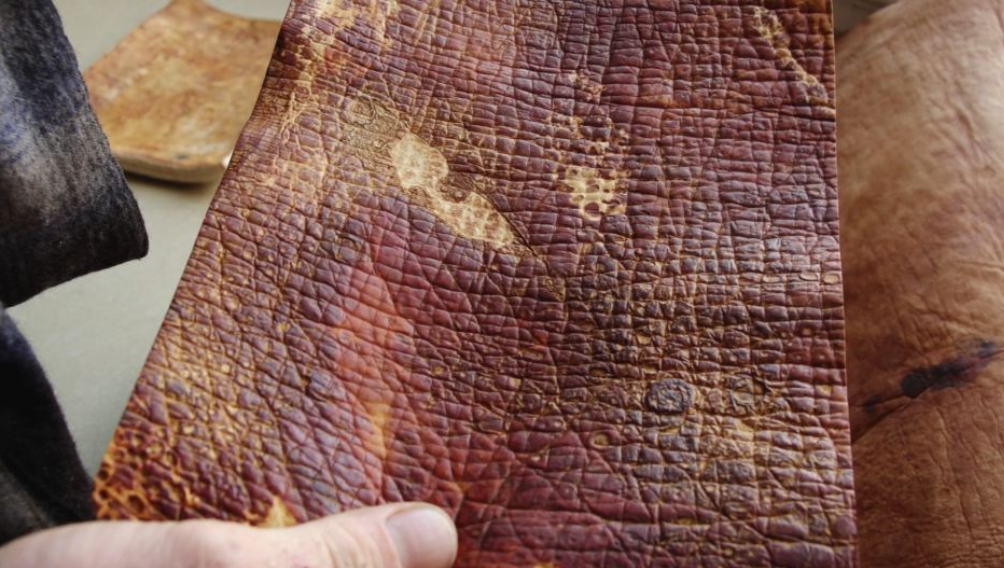
1. Mycelium leather-like materials
One of the best-known applications of fungal materials is in the fashion industry, where mycelium is used to create leather-like sheets and surfaces. Companies such as Bolt Threads and their material Mylo helped popularize this category by presenting fungal alternatives for bags, shoes, and accessories.
These materials are interesting because they can be developed either from liquid-grown fungal biomass or from solid-substrate growth systems, depending on the intended texture and manufacturing process.
Advantages over traditional leather:
- animal-free
- potentially lower environmental impact
- can be grown rather than fully extracted from animals
- texture, thickness, and surface finish can be tuned
- can fit circular and bio-based design strategies
Disadvantages:
- often still require coatings or backing layers for durability
- may not yet match the longevity of high-quality animal leather
- industrial scaling and consistency are still developing
- some products are expensive compared with conventional synthetic leather or mass-market leather
Compared with synthetic “vegan leather” made from plastics, fungal leather alternatives may also offer a more bio-based route, although in practice some current products still include polymer coatings, so they are not always fully biodegradable.
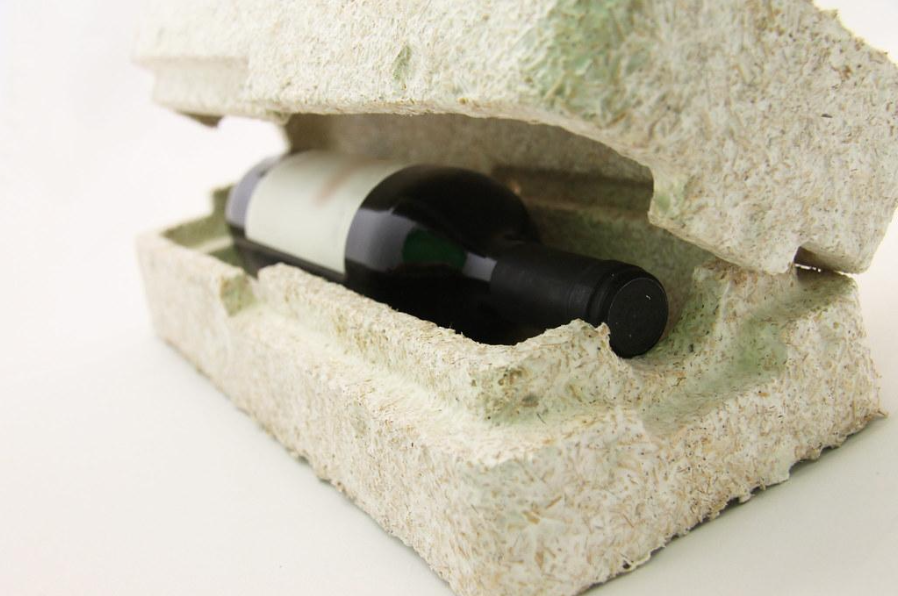
2. Mycelium packaging
Another important example is mycelium packaging, especially developed by companies such as Ecovative. In this case, mycelium is grown through agricultural waste to form protective packaging shapes that can replace expanded polystyrene or other petrochemical foams.
Uses:
- protective packaging for bottles, electronics, and fragile goods
- molded cushioning forms
- compostable alternatives to foam packaging
Advantages over conventional foam packaging:
- biodegradable and compostable
- grown from low-cost waste streams
- lower dependence on fossil-based plastics
- good shock absorption and lightweight performance
Disadvantages:
- more sensitive to moisture than plastic foams
- less suitable for very long-term wet storage
- can be bulkier or less standardized than industrial plastic packaging
- production speed and storage conditions may be more demanding than mass-produced plastic
3. Acoustic and interior panels
Mycelium is also being used for acoustic panels, tiles, and interior surfaces. Companies such as Mogu have developed products that use fungal composites for sound absorption and architectural finishes.
These materials work well because their internal porous structure can help absorb sound, while their low density can also contribute to thermal insulation.
Advantages over conventional acoustic materials:
- bio-based and renewable
- visually distinctive and suitable for interior design
- lightweight
- can provide acoustic and thermal benefits at the same time
Disadvantages:
- usually better suited to indoor than exposed outdoor use
- may require treatment for moisture resistance and durability
- performance can vary depending on substrate, density, and fabrication process
- still less common and less standardized than mineral wool, foam, or gypsum-based systems
4. Architectural and construction experiments
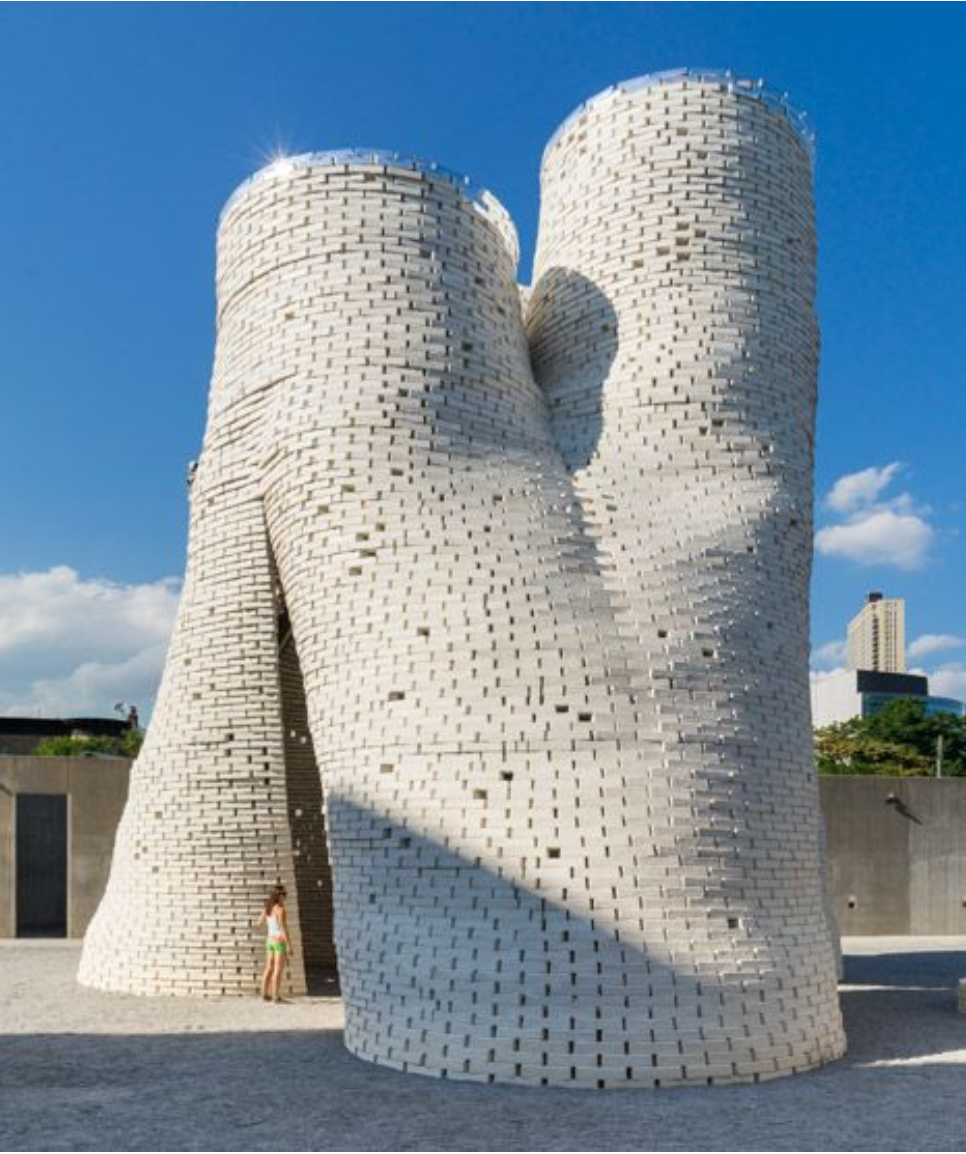
Mycelium has also been used in architecture, especially in experimental pavilions and temporary installations. One famous example is the Hy-Fi pavilion, which demonstrated the potential of mycelium-grown bricks for lightweight, low-carbon construction.
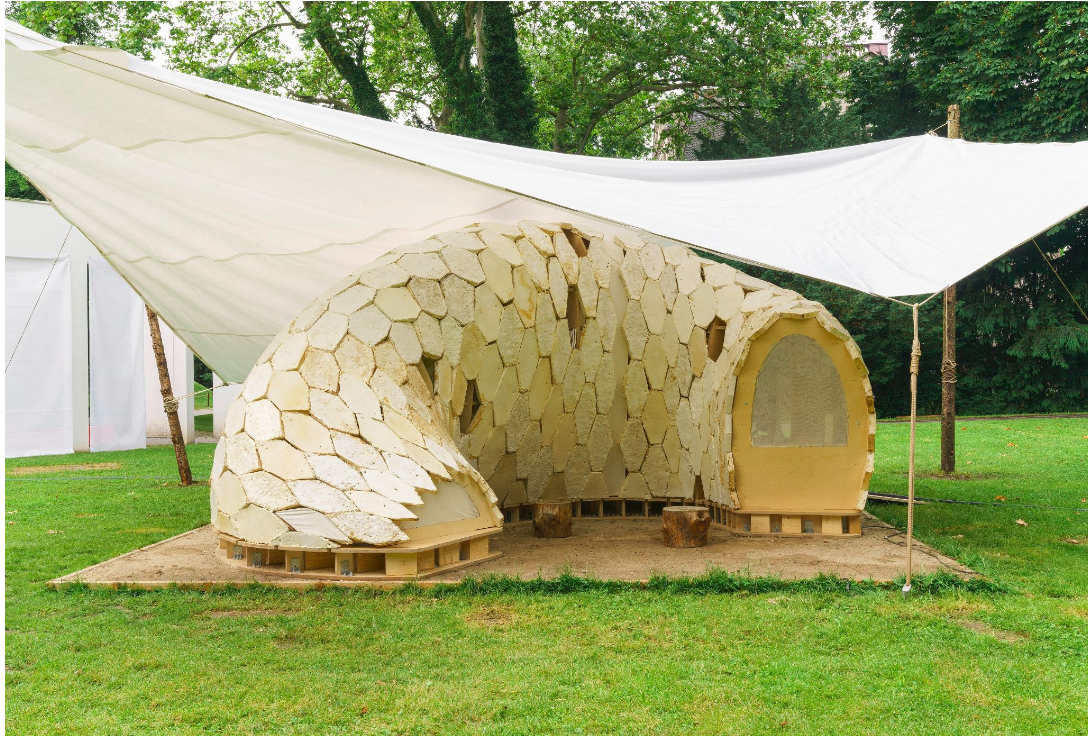
We have also seen exhibition pavilions such as MY-CO SPACE, which use mycelium-based building elements in semi-protected environments.
These projects show that fungal materials can be used not only for products, but also for spatial design and architectural expression.
Advantages over traditional building materials:
- low weight
- grown from renewable waste-based feedstocks
- low embodied energy compared with many fired or petrochemical materials
- biodegradable and visually unique
- suitable for temporary structures, exhibitions, and circular design experiments
Disadvantages:
- limited durability in outdoor conditions without protection
- vulnerable to moisture, weathering, and biological degradation
- lower mechanical strength than brick, concrete, or many engineered panels
- building regulations and long-term structural reliability remain challenges
Conclusion
Fungal materials are a rapidly growing area of biomaterial research and design. Existing examples already include packaging, leather-like materials, acoustic panels, and architectural installations. Their main advantages are that they are lightweight, bio-based, biodegradable, and can be grown on cheap waste substrates. However, compared with traditional materials, they still face important limitations in durability, water resistance, standardization, and structural performance.
For these reasons, fungal materials are especially promising in applications where low weight, sustainability, compostability, and material experimentation are more important than maximum strength or long-term outdoor durability. Rather than replacing all conventional materials, they are currently most valuable as specialized alternatives in design, packaging, interiors, and temporary architecture.
What might you want to genetically engineer fungi to do and why? What are the advantages of doing synthetic biology in fungi as opposed to bacteria?
Compared with bacteria and yeasts, the synthetic biology infrastructure for filamentous fungi is still less mature. One recent review states that, relative to bacteria and yeasts, synthetic biology in filamentous fungi is “rather underdeveloped,” especially in mushroom-forming species, and links this to factors such as slower growth, lower-throughput transformation, unwanted enzyme secretion, and limited plasmid tools [1]. At the same time, this gap is beginning to close. Recent work has developed a modular synthetic biology toolkit for filamentous fungi that includes natural and synthetic promoters, terminators, fluorescent reporters, selection markers, transcriptional regulatory domains, and components for CRISPR-based technologies [2]. This means fungi are no longer only interesting as natural material producers, but are increasingly becoming engineerable biological chassis.
Fungi also offer some major advantages over bacteria for biomaterial-based synthetic biology. Unlike most bacteria, filamentous fungi naturally grow as multicellular, spatially distributed mycelial networks that branch and intermesh across large areas [1]. These networks are well suited for the development of macroscopic living materials that can sense, respond, and potentially compute across space. In addition, filamentous fungi secrete enzymes that degrade lignocellulosic biomass, allowing them to grow on cheap and abundant waste feedstocks [1]. This makes them especially attractive for biomaterials and biomanufacturing, because the fungus can function both as the material itself and as the engineered sensing or production chassis.
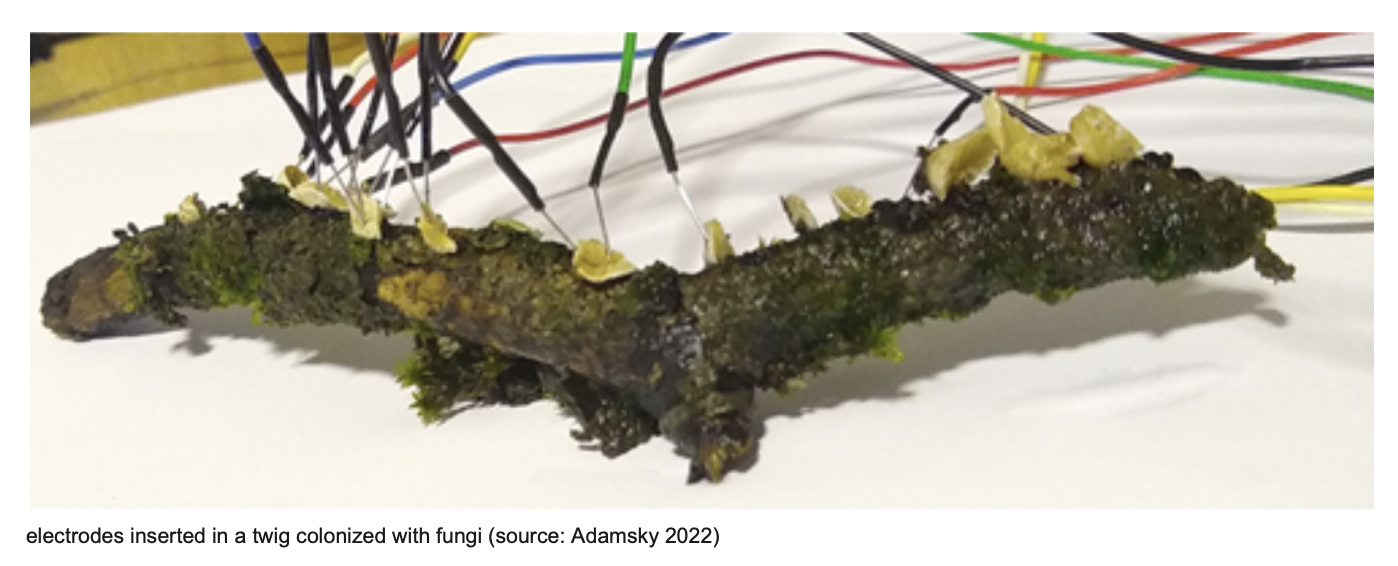
An especially interesting direction is the idea of fungi as living sensory-computational materials. Research by Adamatzky and others suggests that mycelial networks behave as electrically active distributed systems. Fungal colonies generate measurable extracellular voltage spikes, and these spike trains vary in duration, amplitude, and temporal patterning [3]. In related work, Adamatzky and colleagues argue that mycelium exhibits neuron-like spiking behaviour and a wide range of non-linear electrical properties, and they show that electrical signals in Aspergillus niger colonies can in principle be used to implement logical gates and circuits [4]. In that study, they also used an A. niger strain expressing green fluorescent protein (GFP) from the glucoamylase (glaA) promoter [4]. Although this line of research is not always synthetic biology in the strict sense, it provides a compelling conceptual basis for future engineered fungal systems.
It would therefore not be far-fetched to imagine genetically engineering fungi to detect vibration, touch, humidity changes, or electrical activity, and to convert these signals into readable outputs such as fluorescence, color change, altered growth patterns, or production of a specific metabolite. Such systems could be useful for self-monitoring building materials, environmental sensing, smart packaging, or living interfaces. This idea is strengthened by recent evidence that fungi may also respond to sound: Robinson et al. found that acoustic stimulation increased fungal biomass and enhanced Trichoderma harzianum conidia activity [5]. Synthetic biology could extend these native electrical and environmental response behaviours into programmable sensing and response systems.
Overall, bacteria remain easier and faster to engineer in many contexts, but fungi offer a different set of advantages. Their value lies not mainly in engineering simplicity, but in their eukaryotic biology, secretion capacity, growth on low-cost substrates, and ability to form large living material networks [1]. For applications in which the organism itself is meant to become part of a responsive, structural, or computational material, fungi may offer possibilities that bacteria cannot provide as easily.
References
[1] Jo, C., Zhang, J., Tam, J. M., Church, G. M., Khalil, A. S., Segrè, D., & Tang, T.-C. (2023). Unlocking the magic in mycelium: Using synthetic biology to optimize filamentous fungi for biomanufacturing and sustainability. Materials Today Bio, 19, 100560. https://pmc.ncbi.nlm.nih.gov/articles/PMC9900623/
[2] Mózsik, L., Pohl, C., Meyer, V., Bovenberg, R. A. L., Nygård, Y., & Driessen, A. J. M. (2021). Modular Synthetic Biology Toolkit for Filamentous Fungi. ACS Synthetic Biology, 10(11). https://pubs.acs.org/doi/10.1021/acssynbio.1c00260
[3] Adamatzky, A. (2022). Language of fungi derived from their electrical spiking activity. Royal Society Open Science, 9(4), 211926. https://doi.org/10.1098/rsos.211926
[4] Adamatzky, A., Ayres, P., Beasley, A. E., Roberts, N., & Wösten, H. A. B. (2022). Logics in Fungal Mycelium Networks. Logica Universalis, 16(4), 655–669. https://doi.org/10.1007/s11787-022-00318-4
[5] Robinson, J. M., Annells, A., Cando-Dumancela, C., & Breed, M. F. (2024). Sonic restoration: Acoustic stimulation enhances plant growth-promoting fungi activity. Biology Letters, 20(10), 20240295. https://doi.org/10.1098/rsbl.2024.0295
Final Project
I know this part of the homework is not really required for our node but I will use part of the template to try to evaluate some of the ideas that I have. Idea 1 is now canceled and I have narrowed down my research to 2 different ideas
IDEA 1 — BC Face Mask as a cell-free biosensing textile
IDEA 2 — “Water-resistant BC leather” via in-growth synbio functionalization
IDEA 3 — Light-input → color-output BC “bio-print” for moiré effects (E. coli + BC co-culture)
1. Your abstract should briefly address the following elements:
The signafiance: both projects are adressing two separate problems with bacterial cellulose usecases in the textile industry, but they both lead to a clear patch towards a more sustainable fashion industry and have a clear industrial importance concidering the environmental impact of fashion. The Broad Objective: for both projects would be to find sustainable ways to produce textile using bacterial cellulose.
SECTION 3: BACKGROUND
Background and Literature Context Provide background research that explains the current state of knowledge and identifies the gap in knowledge or capability that your project addresses.
IDEA 2 — “Water-resistant BC leather” via in-growth synbio functionalization
These two papers are useful for my final project because they address different parts of the same material problem: how to reduce the strong water absorption of bacterial cellulose by attaching a hydrophobic function to the cellulose surface. The first paper provides a practical fusion-protein strategy. It shows that a class I hydrophobin, HGFI, can be fused to a cellulose-binding domain (CBD), which improves its soluble expression in E. coli and allows the fusion protein to bind directly to bacterial cellulose. This is important for my project because it demonstrates that a CBM/CBD–hydrophobin fusion is a realistic way to deliver a hydrophobic function onto a cellulose material. [1]
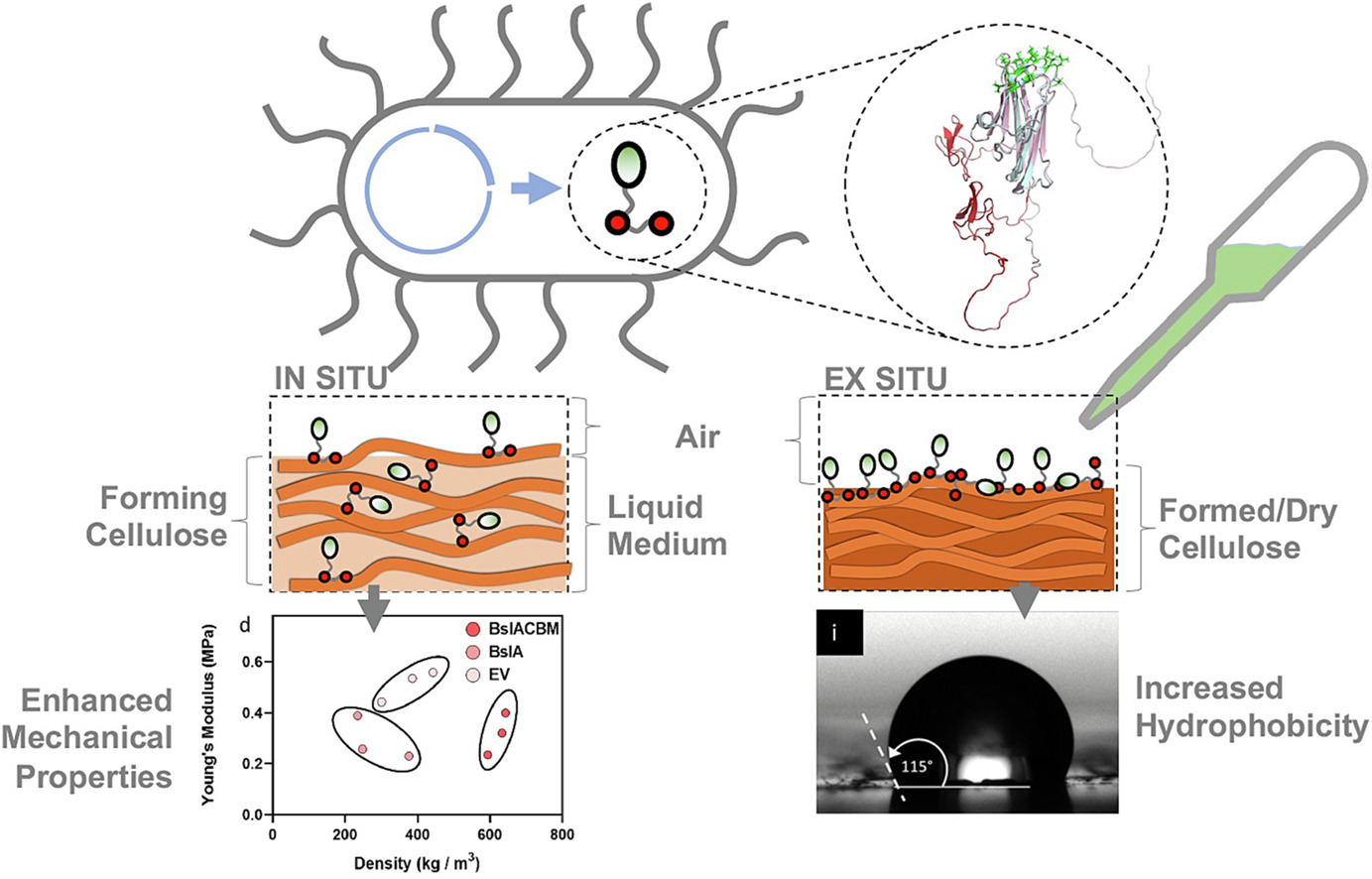

The second paper is useful in a different way. Rather than focusing on hydrophobin production, it identifies a new cellulose-binding module, CBM104, which binds very selectively to native crystalline cellulose I and does so with much higher adsorption efficiency than the more common fungal CBM1. The authors also suggest that CBM104 binds to hydrophilic regions of cellulose microfibrils, while CBM1 recognizes hydrophobic surfaces. This matters for my project because it suggests that the cellulose-binding part of the fusion is not just a generic anchor: choosing a different CBM could change how strongly and where the hydrophobic protein attaches to bacterial cellulose. [2]
It might be possible to speculate that CBM104 could be used as targeted “glue” attaching on the specific part of (hydrophilic regions) that is interesting for me.
Together, these papers suggest a clear strategy for addressing the BC water-absorption problem. The first paper offers a practical method for building and expressing a hydrophobin–CBM fusion, while the second paper suggests a way to improve that strategy by selecting a more specific cellulose-binding domain. For my project, this means I could design a hydrophobin-based bio-finish for bacterial cellulose and compare a standard CBD/CBM with CBM104 to test whether more selective binding to native crystalline cellulose improves water resistance and overall material performance. The research gap is that the first paper does not test water-resistant BC finishing directly, and the second paper does not test a hydrophobin fusion at all, so my project would combine these two ideas into a new BC-finishing approach. [2]
Next research stept:
Paper 1 — Hydrophobin–CBM fusion (HGFI–CBD)
- Design fusion construct (Hydrophobin + linker + CBM)
- Order DNA (e.g. Twist)
- Clone into expression vector (e.g. pET28a)
- Transform into E. coli (Top10 → BL21(DE3))
- Protein expression (IPTG induction)
- Cell lysis (sonication)
- Collect cell-free extract (CFE) or purify protein
- Grow BC pellicles (Komagataeibacter in HS)
- Ex situ: coat BC with fusion protein
- or In situ: add CFE during BC growth
- Dry / process pellicle
- Test hydrophobicity (contact angle)
Paper 2 — CBM104 discovery / application direction
- Select CBM (CBM104 vs CBM1)
- Design fusion construct (Hydrophobin + linker + CBM104)
- Order DNA (Twist)
- Clone into expression vector
- Transform into E. coli
- Protein expression
- Cell lysis → collect CFE / purify
- Grow BC pellicles
- Apply fusion protein to BC (ex situ or in situ)
- Compare binding / performance vs CBM1
- Test material properties (hydrophobicity, absorption)
Future research needed:
- Hydrophobic protein alternatives What other hydrophobin could be interesting?
- Class I vs Class II hydrophobins
- BslA (bacterial hydrophobin-like protein) ← already proven for BC
- Amphiphilic peptides (shorter, easier expression)
- Elastin-like polypeptides (ELPs) (tunable hydrophobicity)
- Lipid-binding proteins / oleosins
- Designed peptides (ProteinMPNN / simple repeats)
- CBM selection strategy
Compare:
- Standard CBM1 / CBD
- CBM104 (paper 2)
- Possibly bacterial CBMs vs fungal CBMs
What to research:
- Binding strength
- Binding location (hydrophilic vs hydrophobic cellulose faces)
- Expression system: E. coli vs Komagataeibacter (VERY IMPORTANT)
Option A — E. coli (current paper approach) pro:
- Easy
- High expression
- Fast con:
- Not integrated into material
- Post-processing step
Option B — Komagataeibacter (KIK / KTK system)
I havent found any researh on this but should maybe be possible to use KTK (Komagataeibacter Tool Kit) cloning system to clone the system straight into Komagataeibacter
- In situ vs ex situ functionalization Compare: Ex situ coating (CFE / purified protein) In situ addition (add protein during growth) Fully engineered BC producer (genetic insertion) Research:
- diffusion into pellicle
- stability of protein during growth
- whether proteins get trapped vs surface-localized
- Material performance metrics
- Water contact angle (WCA)
- Water uptake % (swelling ratio)
- Mechanical properties (tensile strength, flexibility)
- Durability after drying
References
[1] Puspitasari, N., & Lee, C.-K. (2021). Class I hydrophobin fusion with cellulose binding domain for its soluble expression and facile purification. International Journal of Biological Macromolecules, 193, 38–43.
article
[2] Kojima, Y. et al. (2025). A cellulose-binding domain specific for native crystalline cellulose in lytic polysaccharide monooxygenase from the brown-rot fungus Gloeophyllum trabeum. Carbohydrate Polymers, 347, 122651.
article
[3] Gilmour, K. et al. (2025). *Environmentally conscious hydrophobic spray coatings on bacterial cellulose for sustainable and reusable textiles_ [article](https://www.sciencedirect.com/science/article/pii/S0959652625011254#bib15
IDEA 3 — Light-input → color-output BC “bio-print” for moiré effects (E. coli + BC co-culture)
The most important paper is the 2025 Nature Biotechnology study on self-pigmenting bacterial cellulose. It shows that Komagataeibacter rhaeticus can be engineered to produce black bacterial cellulose through tyrosinase expression, and that this pigmentation can be combined with optogenetic control to pattern gene expression in the growing pellicle. This is directly relevant to my project because it proves that BC can be colored from within the growth process itself, rather than only by post-dyeing, and that light can be used as a programmable input for spatial patterning. At the same time, the paper also shows the current limitation: patterned eumelanin still has high background pigmentation and limited contrast, so accurate visual patterning remains a research gap [1].
A second reference is the paper by Levskaya et al., which is one of the foundational demonstrations of bacterial optogenetics. Although it was done in E. coli rather than Komagataeibacter, it established the key idea that a projected light pattern can be converted into a two-dimensional biological image. For my project, this paper is useful as conceptual background: it shows that light can function as a precise design input for pattern formation, which supports the idea of using projected light to “bio-print” patterns into a growing cellulose material [2].
To make this feasible in Komagataeibacter, the genetic toolkit papers are also important. The KTK paper shows that K. rhaeticus can be engineered using a modular Golden Gate cloning system for multigene constructs, while the expanded Acetobacteraceae toolkit provides characterized promoters, RBSs, terminators, and reporter systems for fine control of gene expression in cellulose-producing bacteria. Together these papers show that Komagataeibacter is not only a BC producer, but also a realistic synthetic biology chassis for building more complex circuits such as light-responsive melanin production [3][4].
The more recent Trends in Biotechnology paper by Zhou et al. is useful mainly as a future-direction reference. It shows that colored BC can also be produced through a co-culture strategy using pigment-producing E. coli and BC-producing K. xylinus, achieving seven different colors. This paper might be less relevant as the immediate experimental route, because it is more complex and requires co-culture with E. coli. However, it is valuable because it shows that melanin-based black BC is only one starting point, and that in the future a light-programmed BC system could potentially be extended toward a broader color palette [5].
Together, these papers suggest a direction for my final project. The Nature Biotechnology paper provides the direct experimental basis for light-programmed melanin patterning in bacterial cellulose, Levskaya provides the conceptual foundation for using projected light as a spatial control system, and the KTK / Acetobacteraceae toolkit papers show that Komagataeibacter can realistically be engineered as the host. The research gap is not simply whether BC can be colored, because that has already been shown, but whether higher-fidelity, lower-background, spatially programmable patterning can be achieved in BC, and whether such patterned pellicles can be used to create multilayer optical effects such as moiré.
Potential process
- Design output system (mCherry for prototyping, tyr1 for melanin as final output)
- optogenetic switch construct (light-control system (Opto-T7RNAP))
- Order DNA parts (Twist)
- Assemble constructs KTK / Golden Gate where compatible other cloning strategy if needed for optogenetic parts
- Transform into E. coli for plasmid build/propagation
- Transform engineered plasmids into K. rhaeticus
- Validate reporter expression in liquid culture
- Test and optimize light response with mCherry first
- Grow thin BC pellicles with engineered K. rhaeticus
- Project patterned light during pellicle growth
- Image / quantify pattern quality with mCherry
- Swap reporter to PT7-tyr1
- Grow BC pellicles under patterned light
- Transfer pellicles to melanin development buffer and develop visible eumelanin pattern
- Compare pattern quality
- Grow two separately patterned thin pellicles, overlay them to test moiré effects
Next research stept:
- which Opto-T7RNAP variant is most suitable
- blue-light requirements
- dynamic range
Output choice: mCherry vs tyr1
- Pattern fidelity in BC
- diffusion / blur during growth
- how pellicle thickness affects resolution
- how long you can expose before patterns spread whether thin pellicles give better contrast
- Reactor / growth geometry
- whether to grow each layer separately
- Komagataeibacter toolkit options
- KTK for modular multigene assembly in K. rhaeticus
- promoter / RBS / terminator choices from the expanded Acetobacteraceae toolkit whether you need one plasmid or two antibiotic markers and compatibility
- Development chemistry for melanin
- melanin development buffer composition effect of pH
- effect of tyrosine and copper
- whether development can be made faster or cleaner
- Future color expansion
- whether melanin should stay the final target
- whether the Zhou co-culture platform is better as a future direction for broader color range
- whether one-color high-fidelity patterning is stronger than many colors with weak control
References
[1] Walker, K. T. et al. (2025). Self-pigmenting textiles grown from cellulose-producing bacteria.
Article
[2] Levskaya, A. et al. (2005). Synthetic biology: engineering Escherichia coli to see light.
Article
[3] Goosens, V. J. et al. (2021). Komagataeibacter Tool Kit (KTK): A Modular Cloning System for Multigene Constructs and Programmed Protein Secretion from Cellulose Producing Bacteria.
PDF
[4] Teh, M. Y. et al. (2019). An Expanded Synthetic Biology Toolkit for Gene Expression Control in Acetobacteraceae.
Article
[5] Zhou, H., Lin, P., Jeong, K. J., & Lee, S. Y. (2026). One-pot production of colored bacterial cellulose.
Article