Individual Final Project


ABSTRACT
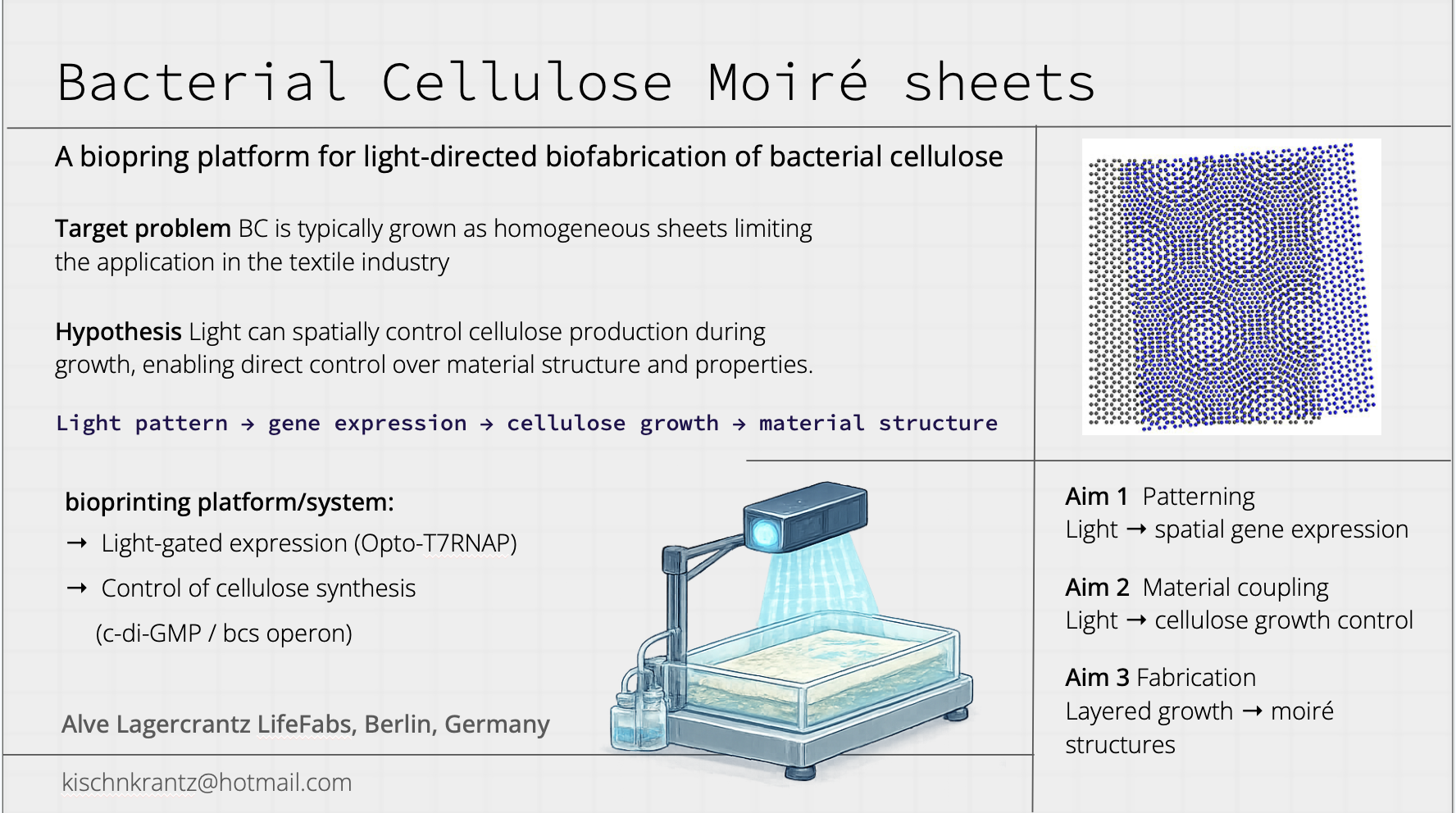
This project explores the use of optogenetic control to encode spatial structure into bacterial cellulose (BC) during growth. Recent advances in engineered Komagataeibacter rhaeticus have demonstrated that gene expression can be patterned within growing pellicles using projected light. However, current approaches primarily focus on pigment production, which is limited in practical textile applications and suffers from low contrast and background expression. This project addresses the need for new fabrication strategies in biomaterials that move beyond surface decoration toward material-level control, which is highly relevant for applications such as textile and bag production.
The overall objective is to develop an optogenetically controlled fabrication platform in which light is used as a programmable input to modulate BC growth and material properties. The central hypothesis is that coupling a light-responsive system to growth-related pathways will enable spatial control over cellulose deposition, resulting in patterned differences in material density, structure, and mechanical behavior that persist after drying.
To test this, the project will pursue several key aims.
- a light-responsive genetic system will be implemented in K. rhaeticus and validated using a reporter output.
- the system will be linked to growth- or material-related regulators to investigate whether projected light patterns can produce measurable structural differences in the pellicle.
- multiple patterned layers will be fabricated and combined to explore moiré-like effects arising from spatial interference between structures.
The project will use synthetic biology tools including modular Golden Gate assembly, transformation of K. rhaeticus, and optogenetic control systems. Patterned light exposure will be applied during pellicle growth, and resulting materials will be evaluated in terms of structure, optical appearance, and mechanical behavior. Together, this work aims to demonstrate a new paradigm for biofabrication in which light-directed biological growth is used to directly encode function and form into a material.
PROJECT AIMS
Aim 1: Experimental Aim
The first aim of my final project is to implement a light-responsive genetic system in Komagataeibacter rhaeticus and validate spatially controlled gene expression in growing bacterial cellulose pellicles by utilizing optogenetic constructs, modular cloning tools for Komagataeibacter, and projected light patterning.
This aim focuses on establishing the core synthetic biology platform needed for the project. The experimental goal is to confirm that projected light can act as a programmable input in K. rhaeticus and generate patterned output within the growing pellicle. As a first step, this system will be tested with a reporter output before being connected to material-regulating genes. Relevant resources include optogenetic systems, the Komagataeibacter Tool Kit (KTK) for modular Golden Gate assembly, and expression control elements from the Acetobacteraceae toolkit.
Aim 2: Development Aim
The second aim of my final project is to connect the validated light-responsive system to growth- or material-related regulators in K. rhaeticus in order to test whether projected light patterns can produce measurable structural differences in bacterial cellulose pellicles.
If Aim 1 is successful, this stage will move from reporter-based patterning to material-level control. This includes identifying and implementing genes or regulatory pathways involved in cellulose production, pellicle morphology, or local density. The goal is to evaluate whether light-controlled gene expression can generate spatial variation in structure, such as thickness, alignment, or surface topology within the BC material.
Aim 3: Visionary Aim
The third aim of my final project is to develop a fully programmable biofabrication platform in which light-directed growth of bacterial cellulose enables precise spatial control over material properties, structure, and function.
In this vision, projected light acts as a digital design interface for biological manufacturing, allowing bacterial cellulose to be grown with encoded patterns of density, texture, and mechanical behavior. This could enable the fabrication of multilayer materials with interference-based effects such as moiré, as well as tunable optical, tactile, and structural properties. If fully realized, this approach could transform bacterial cellulose from a passive biomaterial into an active, programmable manufacturing system for applications in textiles, fashion, and engineered living materials.
Background and Literature Context
Bacterial cellulose (BC) has increasingly been proposed as a sustainable alternative to petrochemically derived materials in fashion and textile contexts because it can be grown microbially, does not require agricultural land, and offers high purity, crystallinity, tensile strength, biocompatibility, and biodegradability. BC is produced as a dense nanofibrillar pellicle by Komagataeibacter strains and is already being explored across apparel, design, medicine, and electronics [1]. Due to its nanoscale structure, BC is typically fabricated as homogeneous leather-like sheets, although it has also been adapted to more traditional textile processes such as spinning and weaving.
However, the limitation of current BC-based textiles is not only material substitution but also manufacturing logic. Existing production systems rely on centralized, large-scale processes, while BC is inherently compatible with distributed, growth-based fabrication. This project argues that new materials require new modes of manufacturing.
This project addresses that limitation by exploring whether BC can be spatially programmed during growth rather than only modified after production. A key enabling development is the emergence of synthetic biology toolkits for Komagataeibacter rhaeticus. The Komagataeibacter Tool Kit (KTK) provides a modular Golden Gate cloning system for multigene assemblies, while the expanded Acetobacteraceae toolkit provides characterized promoters and regulatory parts for controlled gene expression [2,3]. Together, these systems establish Komagataeibacter as a programmable chassis for engineered living materials.
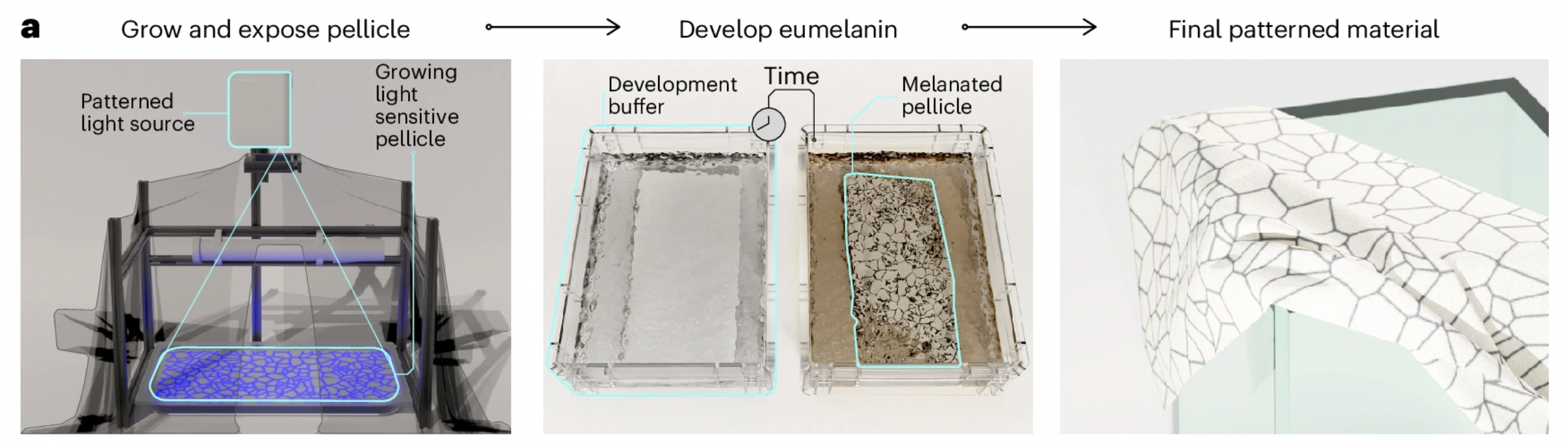

 [4] Walker, K. T. et al. (2025).
[4] Walker, K. T. et al. (2025).
The most direct precedent for this project is the recent Nature Biotechnology study demonstrating optogenetic patterning in BC-producing bacteria. In this work, K. rhaeticus was engineered to produce melanin through tyrosinase expression, and projected light was used to spatially control gene expression within the growing pellicle [4]. While this establishes light as a programmable input, the output remains limited to pigment, with issues such as background expression and low contrast. This highlights a key limitation: current systems enable patterned color, but not patterned material structure.
The conceptual foundation for light-based biological patterning originates from Levskaya et al., who demonstrated that projected light can be converted into spatial gene expression patterns in engineered bacteria [5]. This establishes light as a digital interface for biological fabrication.
Parallel work in BC engineering shows that cellulose production itself can be genetically controlled. Florea et al. demonstrated programmable cellulose production and functionalized materials in K. rhaeticus, suggesting that growth and material formation are accessible targets for engineering [6]. However, these approaches have not yet been combined with spatially resolved optogenetic control, leaving a gap in the ability to encode structure during growth.
The material concept of this project draws on moiré patterns as a structural principle. While moiré effects are widely studied in nanomaterials and optics, particularly in layered systems with offset periodic structures, they have not been explored in BC fabrication. BC’s nanoscale fibrillar network suggests that fine spatial patterning could influence not only visual appearance but also mechanical and structural behavior [7].
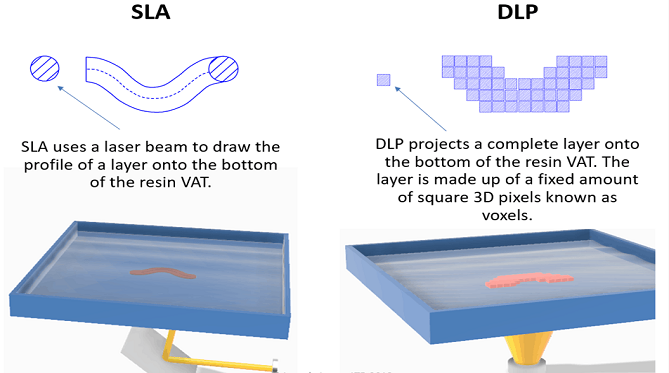
From a fabrication perspective, this project aligns with developments in additive manufacturing. Techniques such as DLP/SLA printing demonstrate how projected light can define spatial material formation, while architected materials research shows how geometry can control mechanical behavior. The concept of “Living Manufacture” proposes a microbial fabrication paradigm integrating digital design and biological growth [1].
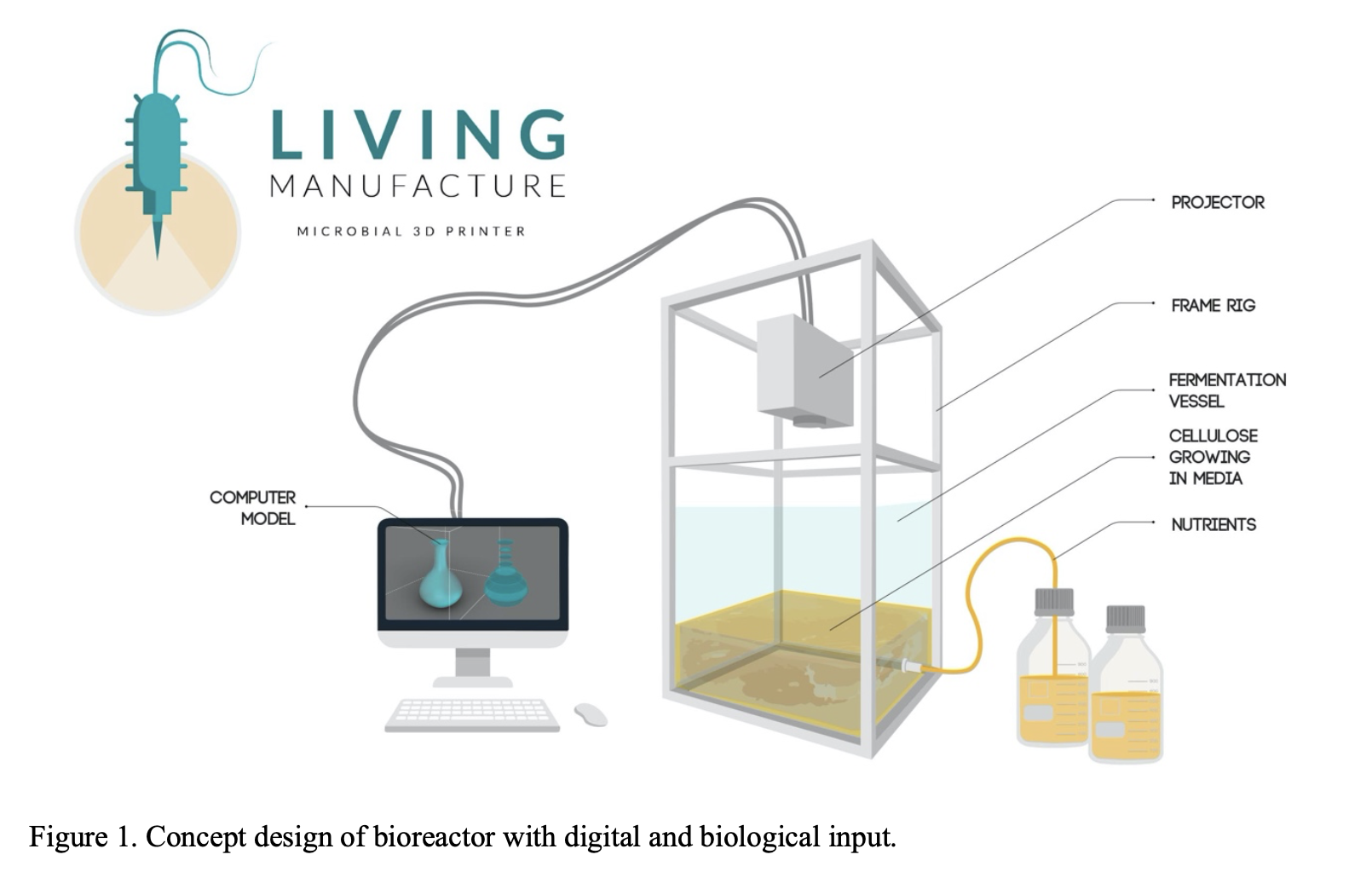
 Arnardottir, T. H. et al. Living Manufacture: Principles for a microbial 3D printer.
Arnardottir, T. H. et al. Living Manufacture: Principles for a microbial 3D printer.
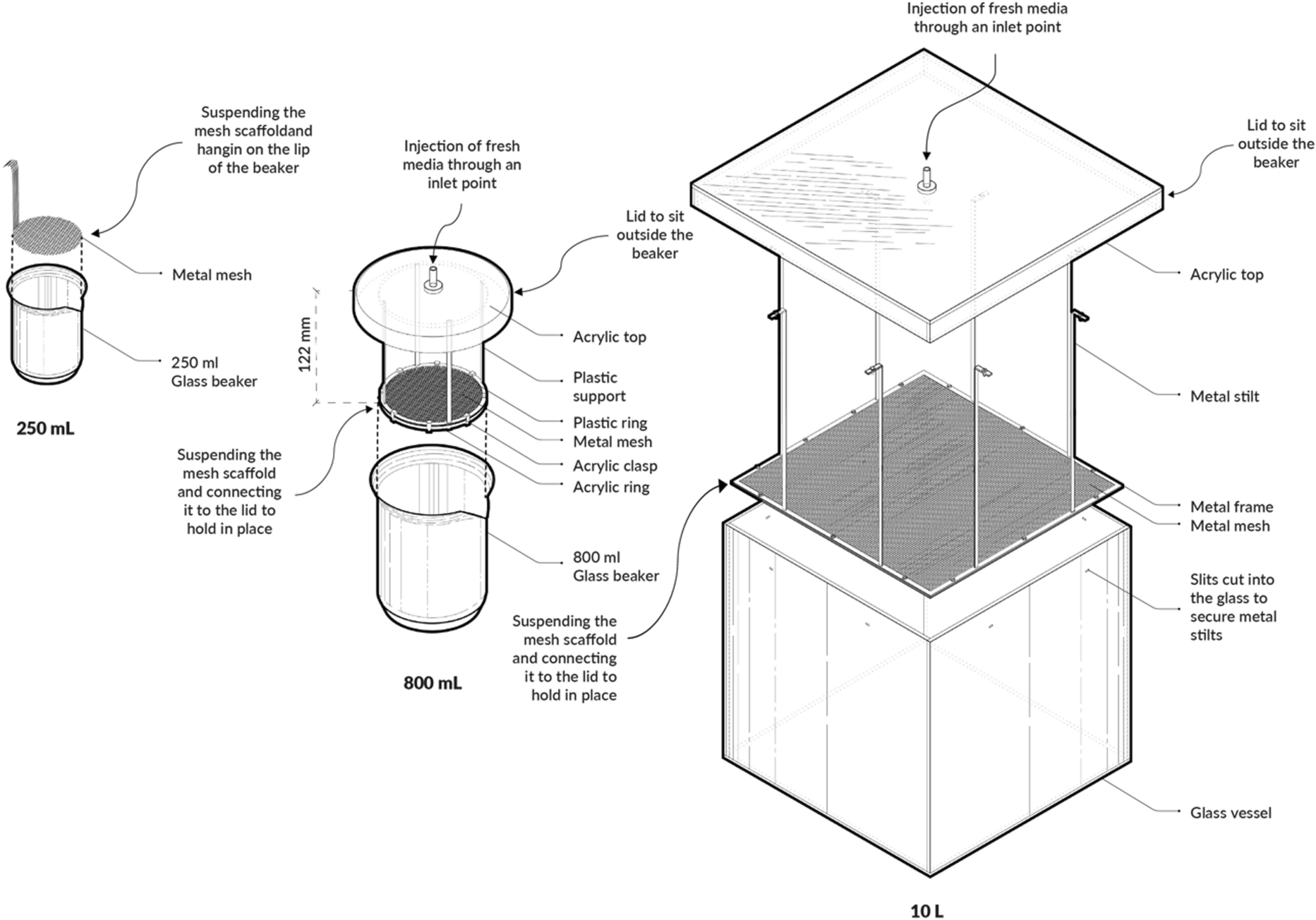
Recent advances in bioreactor design further support this approach. The mesh dispenser vessel (MDV) bioreactor enables top-fed growth, maintaining active cellulose production at the surface and improving layer control [8]. Similarly, the surface air-flow (SAF) bioreactor demonstrates that BC material properties can be tuned through process conditions and reactor geometry [9].
 [5] Levskaya, A. et al. (2005).
[5] Levskaya, A. et al. (2005).
Taken together, these studies show that while BC is a promising sustainable material and optogenetic control is feasible, there is currently no system that combines light-based spatial control with growth-driven material structuring. This project addresses this gap by developing a light-directed BC fabrication platform that encodes structure during growth, enabling layered interference effects such as moiré and advancing BC toward programmable biofabrication.
16/4
Discussion
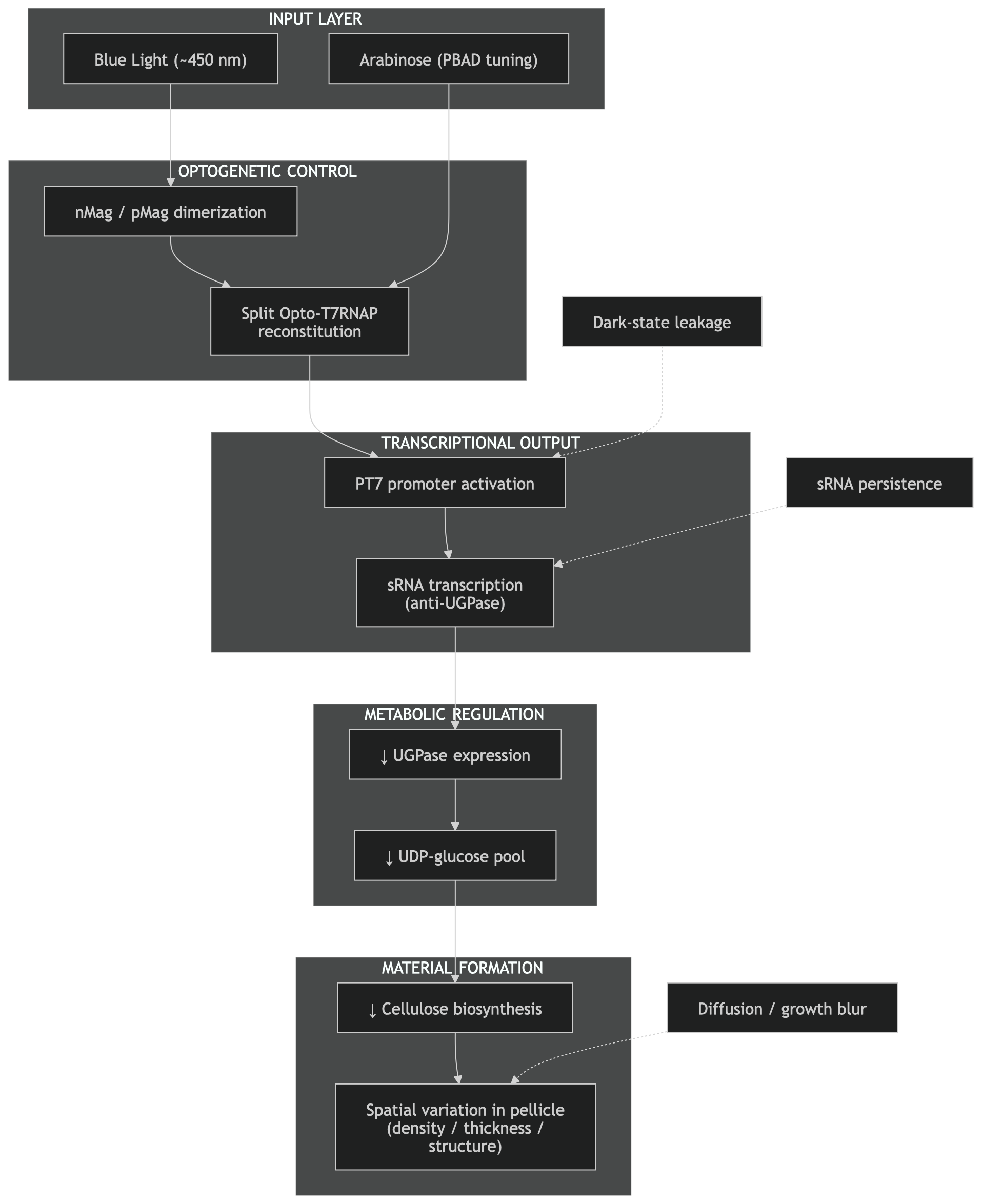
For this project, the optogenetic control system described in [4] is used as a programmable input layer, while the sRNA-based regulation of cellulose biosynthesis from [6] is used as the output layer controlling material formation. By combining these two systems, the aim is to translate spatial light information into spatial control over bacterial cellulose production.
Input
The optogenetic system implemented in [4] is based on a blue-light-sensitive split T7 RNA polymerase (Opto-T7RNAP), originally developed in E. coli and later adapted to Komagataeibacter rhaeticus. A key advantage of this system is that it relies on flavin adenine dinucleotide (FAD) as a chromophore, a metabolic cofactor present in most bacterial cells, including K. rhaeticus. This eliminates the need for additional genetic components for chromophore biosynthesis and makes the system relatively straightforward to transfer across organisms.
Functionally, the T7 RNA polymerase is divided into two inactive fragments (N-terminal and C-terminal), each fused to light-sensitive dimerization domains (nMag and pMag). In the absence of blue light, these fragments remain separate and inactive. Upon illumination at ~450 nm, nMag and pMag heterodimerize, bringing the two halves of the polymerase together and restoring its transcriptional activity. This creates a reversible, light-gated transcription system, where gene expression can be dynamically switched on and off depending on light exposure.
A central feature of this system is the use of the T7 promoter (PT7), which is orthogonal (not interfering with) to the host transcription machinery. Only the reconstituted T7RNAP can recognize and activate PT7, enabling tight and specific control of downstream gene expression. In the original paper, this system was used to drive expression of reporter genes such as GFP or mCherry, as well as functional outputs like tyrosinase (Tyr1) for melanin production.
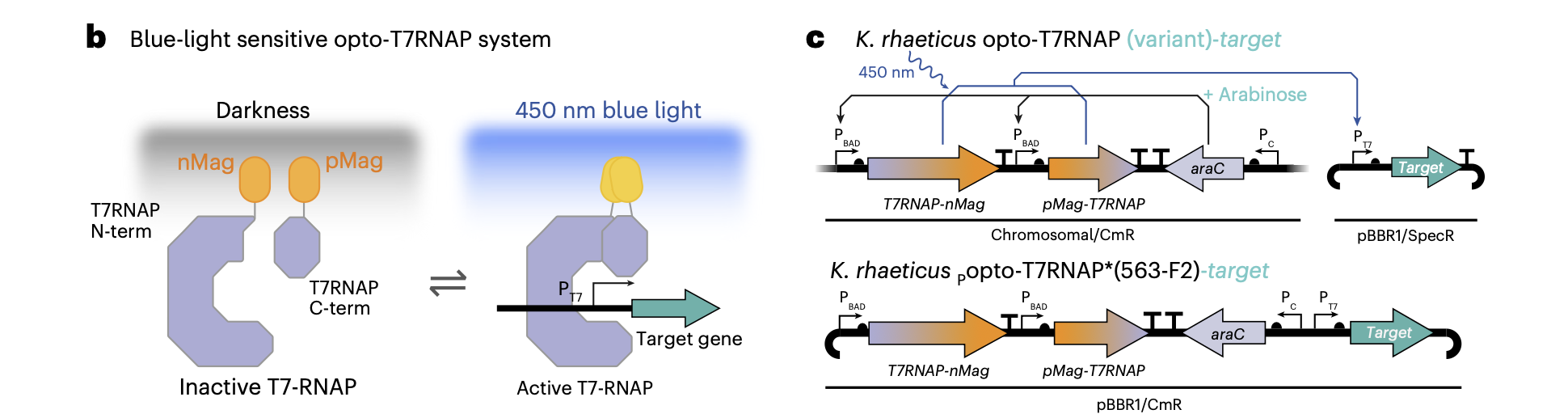

Walker, K. T. et al. (2025)
Importantly, the system operates with two layers of control. First, arabinose induction regulates expression of the Opto-T7RNAP components via the pBAD promoter. This determines how much of the split polymerase is present in the cell. Second, blue light controls the activity of the polymerase by inducing dimerization (nMag and pMag). In this sense, arabinose does not directly control the output gene, but rather tunes the dynamic range of the system. As shown in [4], too high expression levels lead to background activity in the dark (leaky expression), while too low levels reduce overall signal output. Therefore, careful tuning of arabinose concentration is essential to achieve a high contrast between light and dark states.
Footnote:
A dimerization domain is a part of a protein that makes it bind to another protein (or itself)—so that two proteins come together and form a dimer.
- “di-” = two
- “-mer” = unit
Output
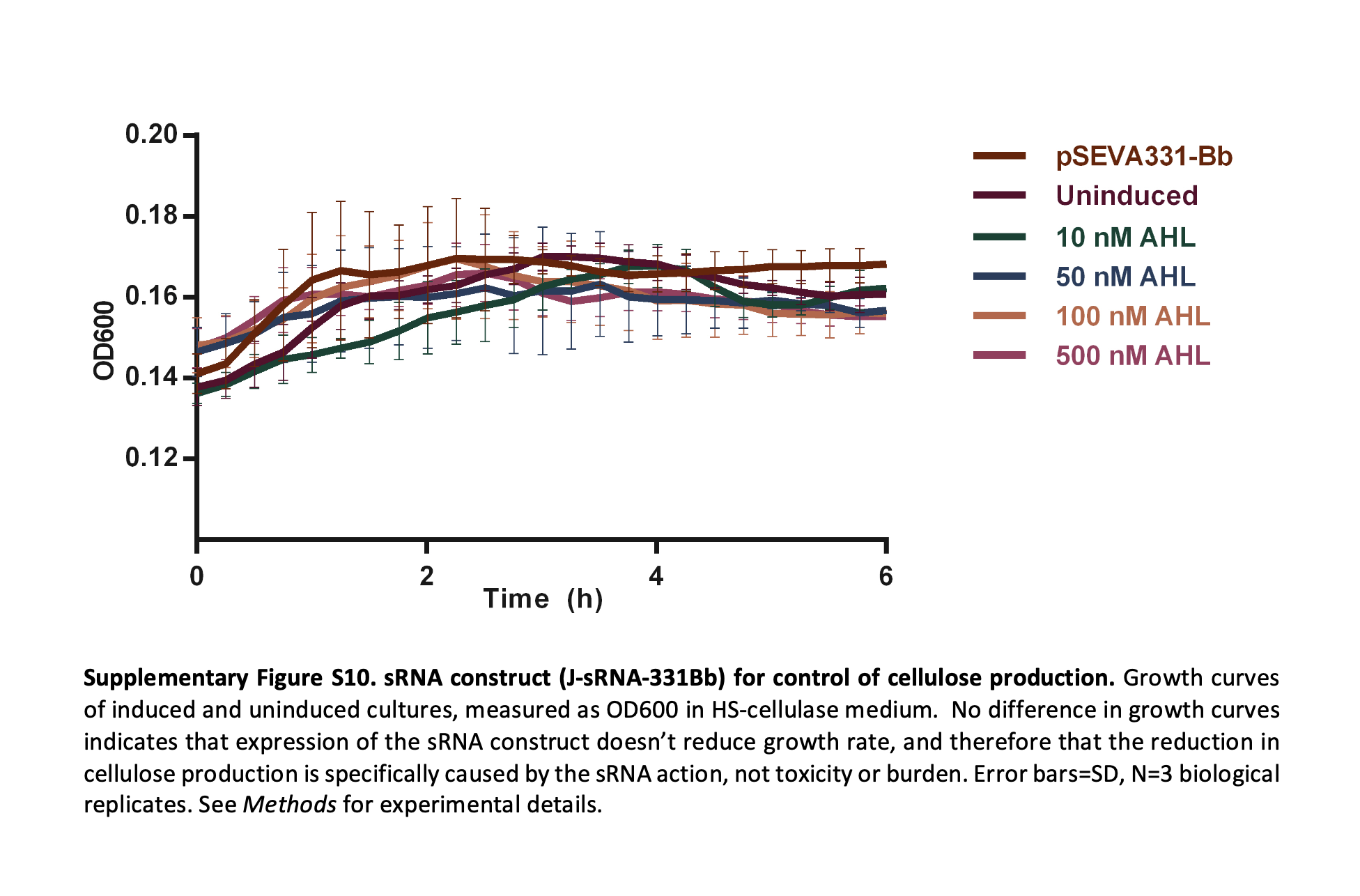
For the output layer, the system described in [6] provides a mechanism to directly modulate bacterial cellulose production. In this work, small regulatory RNA (sRNA) is used to target and downregulate UGPase, an enzyme responsible for producing UDP-glucose, the key precursor for cellulose biosynthesis. By reducing UGPase expression, the metabolic flux toward cellulose production is decreased, allowing control over the thickness and density of the resulting material.
A key insight from [6] is that this intervention does not significantly affect bacterial growth, as shown by unchanged OD600 growth curves across different induction conditions. This decoupling of growth and material production is particularly important for this project, as it enables modulation of material properties without destabilizing the system or reducing viability.
In the original implementation, sRNA expression is controlled via a quorum sensing system based on acyl-homoserine lactone (AHL). In this system, AHL binds to the transcriptional regulator LuxR, which in turn activates the pLux promoter to drive sRNA expression. However, for the purposes of this project, this chemical input can be replaced with an optogenetic one by placing the sRNA construct directly under control of the T7 promoter.
This leads to a simplified and more direct architecture:
- From: AHL → LuxR → pLux →sRNA → ↓ UGPase → ↓ cellulose
- To: Light → Opto-T7RNAP → PT7 → sRNA → ↓ UGPase → ↓ cellulose
In this configuration, spatial light patterns directly regulate sRNA expression, which in turn modulates cellulose production locally within the growing pellicle. This avoids the need for intermediate regulatory layers such as LuxR/pLux, reducing system complexity and potential sources of noise or delay.
However, the results from [4] also highlight important limitations that are directly relevant to this project. When using enzymatic outputs such as tyrosinase, the system exhibited significant background activity, leading to melanin production even in the absence of light. This reduced the contrast and resolution of spatial patterning. Similar challenges are likely to arise when controlling cellulose production via sRNA, where leaky expression could lead to global suppression rather than localized modulation.
Achieving high spatial resolution therefore depends on minimizing background expression and maximizing the dynamic range between light and dark states. This can be addressed through several strategies, including tuning arabinose concentrations, optimizing promoter strength, reducing plasmid copy number (e.g., through chromosomal integration), or introducing degradation mechanisms for the sRNA to reduce persistence.
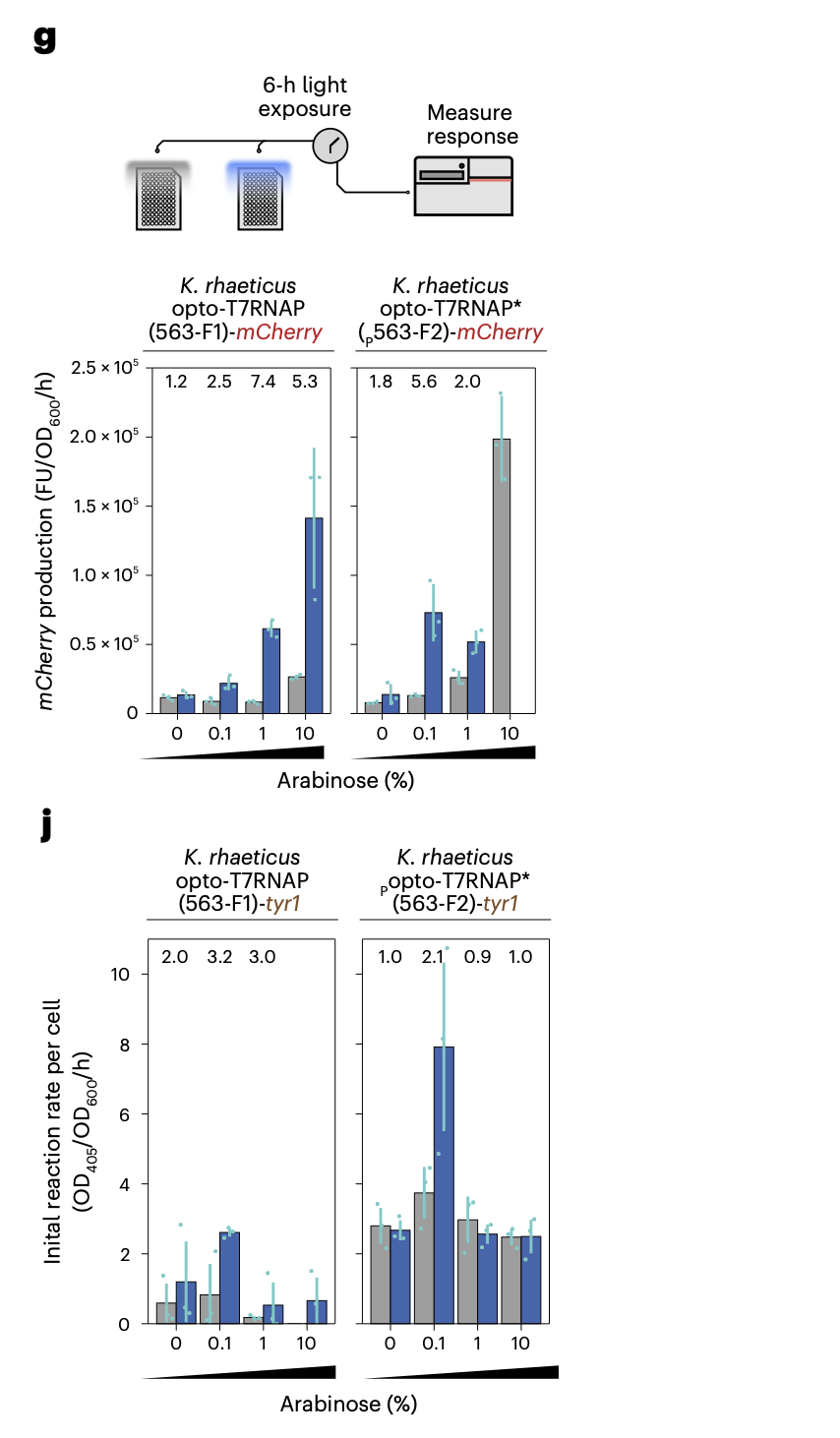

Characterization of optogenetics constructs with mCherry target gene under differing arabinose percentage (wt/vol) concentration. Bars (blue, exposed and gray, unexposed) show mean increase in red fluorescence after 6 h normalized by OD600. Error bars show s.d. of three biological replicates. Fold difference between exposed and unexposed cells is shown above, except in instances of poor growth j, Optogenetic construct characterization with tyr1 under differing arabinose induction. The bars (blue,exposed and gray, unexposed) show the mean and s.d. of three biological replicates of initial (0–100 min) reaction rate of eumelanin production measured at OD405, normalized to the number of initial cells at OD600 at time point 0. source Walker, K. T. et al. (2025).
Note the opto-T7RNAP system demonstrates tunable, light-responsive gene expression in K. rhaeticus. However, downstream functional outputs (e.g. Tyr1 activity) show nonlinear dependence on induction conditions (see pic above), highlighting a key challenge in translating gene-level control into predictable material outcomes such as bacterial cellulose structure.
Additional constraints arise from the physical properties of the system. In [4], the spatial resolution of patterned gene expression was reported to be on the order of ~0.8 mm, and significant exposure times (up to 40–80 hours) were required to generate visible outputs. These temporal and spatial limitations are particularly relevant for bacterial cellulose, which grows as a continuous pellicle at the air–liquid interface. Over time, diffusion of signaling molecules, enzymes, or metabolic intermediates may further blur spatial boundaries, reducing pattern fidelity.
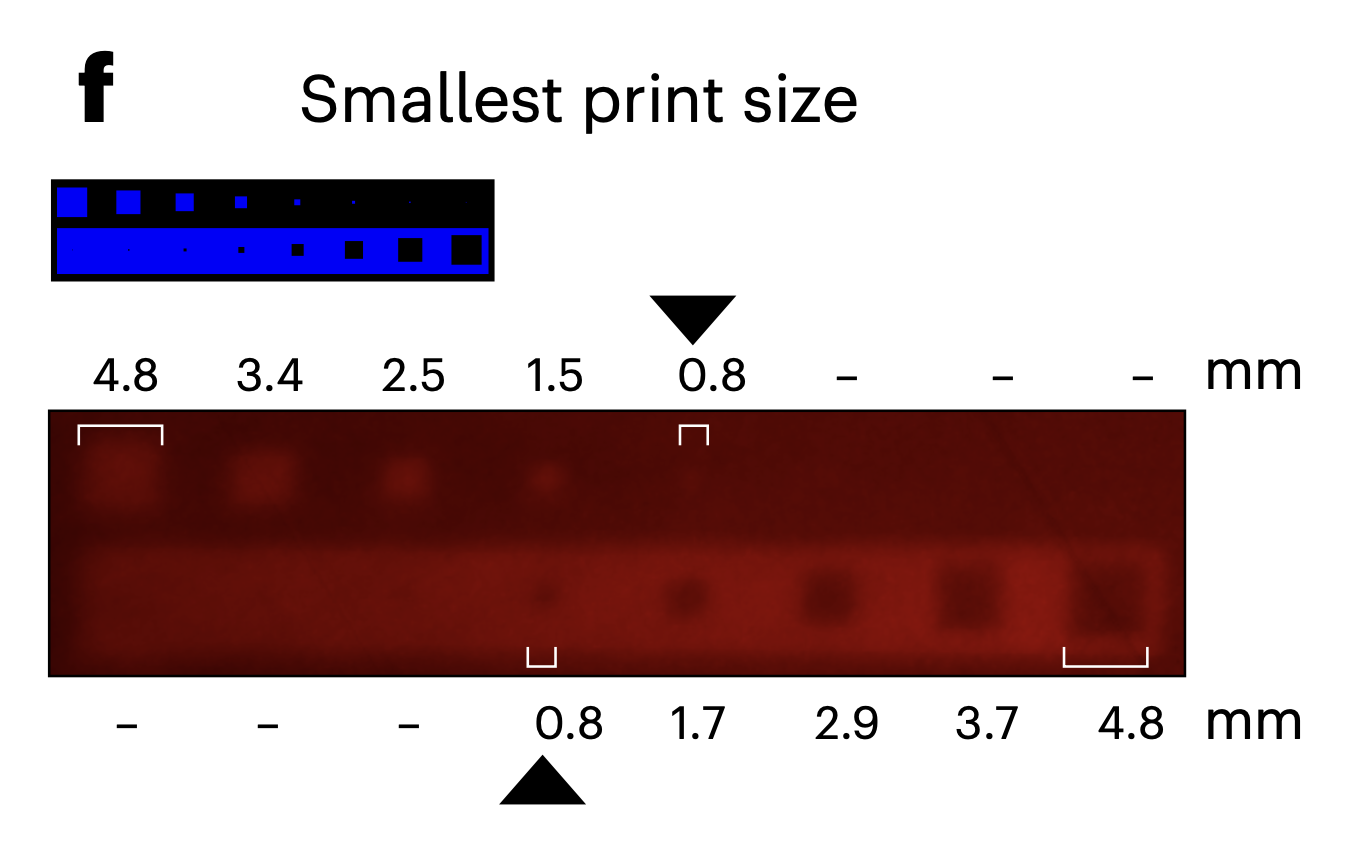

Smallest projected mark on the exposed pellicle. Walker, K. T. et al. (2025)
Taken together, these findings suggest that while optogenetic control of cellulose production is feasible, achieving precise and high-resolution material patterning will require careful optimization at both the genetic and process level. In particular, balancing expression strength, minimizing background activity, and accounting for diffusion and growth dynamics will be critical for translating light patterns into stable material structures.
18/4
Concidering the logistic and the cost related to try to develop this project in vitro I have reformulated my 3 aims:
Aim 1: Experimental Aim (In Silico)
To validate, in silico, a light-responsive genetic architecture for spatial control of bacterial cellulose production in Komagataeibacter rhaeticus by combining the Opto-T7RNAP system with an sRNA module targeting UGPase. This aim focuses on establishing the core synthetic biology logic of the project and testing whether projected light patterns could, in principle, be translated into localized suppression of cellulose production. The validation for this aim will consist of DNA construct design, circuit architecture, and computational analysis of system behavior under different conditions of light exposure, arabinose tuning, leakage, and spatial blur.
Aim 2: Development Aim (in vitro)
The second aim of my final project is to test how the proposed circuit would behave in a growing pellicle. This includes testing, dark-state leakage, sRNA persistence, and pattern diffusion affect the expected spatial resolution and material contrast of the system. The goal is to determine which design parameters are most critical for achieving patterned structural output before wet-lab implementation.
Aim 3: Visionary Aim (in vitro)
The third aim of my final project is to construct a programmable biofabrication platform in which projected light acts as a digital design interface for growing bacterial cellulose materials with encoded structure, density, and interference-based effects such as moiré.
Aim 1 success criteria
- one final circuit architecture
- one DNA design package
- one working simulation notebook
- at least 3 simulated pattern cases
- one quantitative interpretation of leakage / blur limits
- one future wet-lab roadmap
Aim 1 validation
- Design the DNA architecture
- Build the construct logic for
Light → Opto-T7RNAP → PT7 → sRNA → ↓ UGPase → ↓ cellulose
- Specify parts, regulatory logic, and construct organization.
- Software: Benchling, Twist, GenBank / NCBI
- Run computational analysis
- Simulate projected light input versus predicted output.
- Based on data from paper [4] and [6] Test how leakage and blur affect pattern contrast.
- Software: Python, Notebooks, Grasshopper?
- Interpret the result biologically
- Show whether the system is likely to support spatial control in principle.
- Identify the parameter window where the design works or fails.
Experimental Plan (in this order)
- finalizing the biological architecture of the proposed system, combining the blue-light-responsive Opto-T7RNAP input layer from Walker et al. with the sRNA-mediated UGPase repression strategy from Florea et al. in order to create the circuit logic: Light → Opto-T7RNAP → PT7 → sRNA → ↓ UGPase → ↓ cellulose.
- Define the core DNA components required for the system, including the split Opto-T7RNAP modules, the PBAD/arabinose tuning layer, the PT7-controlled sRNA cassette, and the target sequence directed against UGPase mRNA.
- Design the proposed constructs in Benchling
- Translate the biological circuit into a computational model (Notebook) that links projected light intensity to predicted transcriptional activation.
- The first layer of the model will represent projected light as a two-dimensional spatial input map, using simple test patterns such as stripes, gradients, dots, and offset line fields relevant to the moiré concept.
- The second layer will model the optogenetic response of Opto-T7RNAP, including both basal dark-state activity and light-induced activity, the literature shows that the system can become leaky depending on arabinose concentration and target gene choice.
- Arabinose will be included as a tuning parameter that changes the total level of Opto-T7RNAP expression and therefore influences both output strength and background activity.
- The third layer of the model will represent sRNA accumulation and persistence over time, using a simplified response function rather than a full kinetic model.
- The fourth layer will convert predicted sRNA levels into relative UGPase repression and then into a predicted cellulose suppression map, representing the expected material-level output of the system.
- Include a spatial blur or diffusion to represent the effects of time integration, biological averaging, and possible spreading of the signal during pellicle growth.
- Test the system under multiple scenarios, including low-, moderate, and high-leak conditions, to determine at which point spatial patterning becomes unreliable.
- I will compare different projected feature sizes in the model to estimate what resolution might be realistically retained once leakage and blur are included.
- I will use these simulations to identify a plausible operating window for the system, including the kinds of patterns most likely to survive and the types of patterns most likely to collapse into uniform output.
- Finally, I will interpret the results as a feasibility analysis for future wet-lab implementation in Komagataeibacter rhaeticus, including a clear decision tree for what should be experimentally tested first if the project is continued after HTGAA.
SECTION 5: RESULTS & QUANTITATIVE EXPECTATIONS
I will validate the core circuit logic of the project: whether a light-responsive Opto-T7RNAP input system coupled to a PT7-controlled sRNA targeting UGPase could plausibly generate spatially differentiated cellulose production in Komagataeibacter rhaeticus.
- Detailed protocol of how you validated it
- Design the proposed DNA architecture in Benchling.
- Define circuit components and regulatory connections.
- Build a simplified computational model linking light input to predicted cellulose suppression.
- Simulate multiple projected patterns and parameter regimes.
- Quantify contrast, feature retention, and failure conditions under increasing leakage and blur.
- Compare outputs across scenarios and interpret which parameter ranges are viable.
3. What synthetic biology techniques did you use?
- DNA construct design
- Genetic circuit design
- Literature-based chassis and part selection
- Computational modeling of gene regulation and spatial output
- Quantitative analysis of predicted circuit behavior
4. Present data and analysis
- Construct maps
- Parameter table
- Simulated 2D pattern outputs
- Plot of leakage versus contrast
- Plot of blur versus retained feature size
- Comparison of optimistic versus failure-mode scenarios
5. Challenges, limitations, and alternative strategies
- The model does not predict exact biological kinetics.
- Leakage in the real organism may differ from the simulated assumptions.
- Diffusion and pellicle growth dynamics are simplified.

19/4
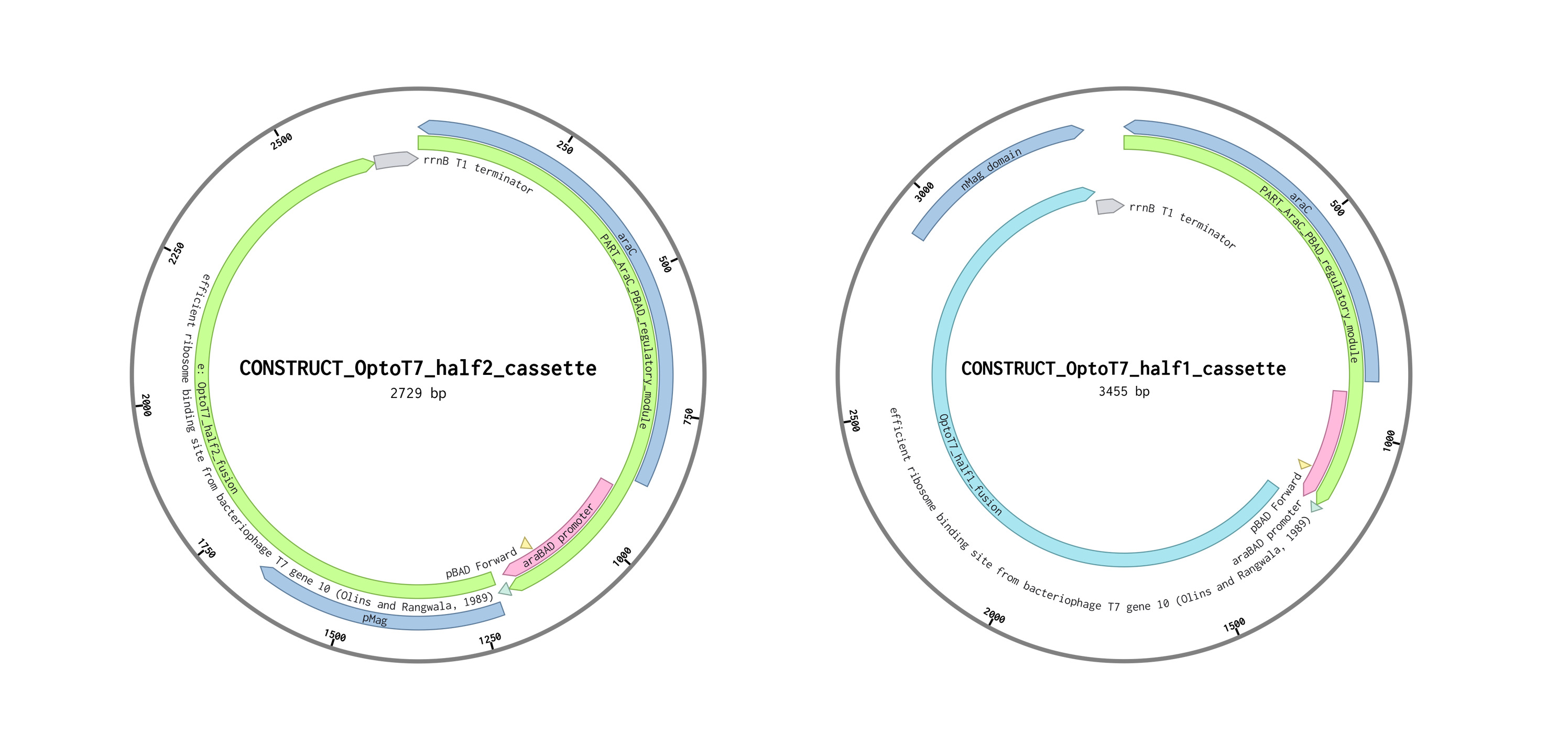
Opto-T7RNAP Cassette Design (Half1 & Half2)
This work focused on constructing two expression cassettes for the split Opto-T7RNAP system in Komagataeibacter rhaeticus, enabling light-controlled gene expression.
Each cassette follows the same modular architecture: AraC + pBAD regulatory module → RBS → OptoT7 fusion protein → rrnB T1 terminator
“Each OptoT7 cassette includes the AraC–pBAD regulatory module, enabling arabinose-inducible expression of the split OptoT7 RNAP halves.”
Sequence sourcing
The AraC/pBAD module was extracted as a complete regulatory unit from a validated plasmid (as described in [4]), rather than rebuilt from individual parts, ensuring correct promoter regulation and arabinose responsiveness. The RBS is a strong ribosome binding site derived from the T7 gene 10 sequence, chosen for efficient translation. The rrnB T1 terminator was selected as a reliable bacterial terminator to ensure proper transcriptional termination.
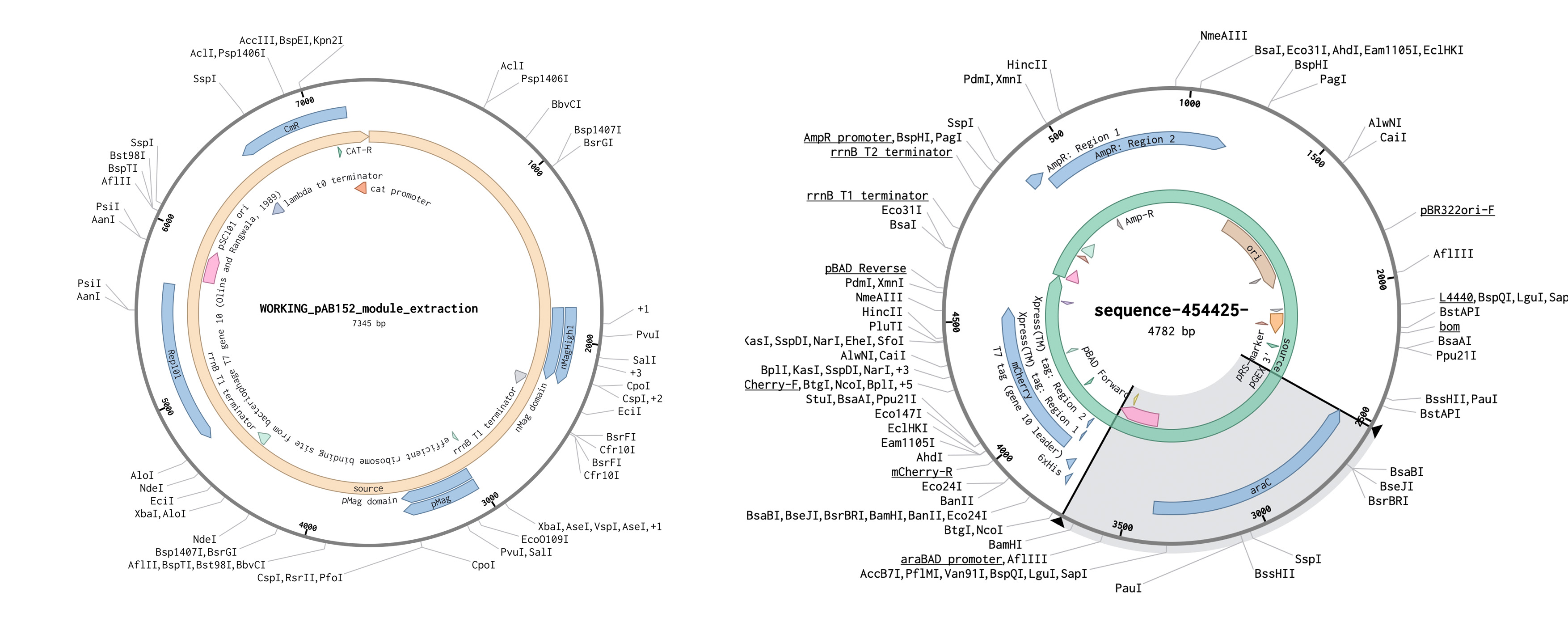
 The two plasmids used for the construct
The two plasmids used for the construct
Source T7RNAP fragment fused to nMagHigh1 and pMag: https://www.addgene.org/54630/ (same as article [4]) Source pBAD: https://www.addgene.org/54630/ picked because it is a common plasmid
The Opto-T7RNAP split system was implemented as two separate constructs:
- Half1: T7RNAP fragment fused to nMagHigh1
- Half2: T7RNAP fragment fused to pMag
Design strategy
Rather than assembling parts from scratch, the approach was to:
- Extract coherent functional modules from existing, validated constructs preserve native regulatory architecture and minimize risk of misassembly or unexpected behavior.
- Both cassettes were finalized as linear, synthesis-ready DNA sequences, with verified structure, correct reading frames, and consistent regulatory logic.
the two cassetes can be found here:
- https://benchling.com/s/seq-IW5krI1Tp60w9H4YLG3P?m=slm-faHhPi0XNjdWFQRu9eTS
- https://benchling.com/s/seq-ozdvCTDIXOeobU2qpBWa?m=slm-TK6zIsI9BJnzYeyE8O1l
20/4
Output cassette design rationale
The output layer of this project is adapted from the Florea et al. UGPase silencing construct (J-sRNA-331Bb / BBa_K1321328), which was originally designed to suppress cellulose production in Komagataeibacter rhaeticus using an AHL-inducible sRNA system. In the original construct, LuxR is expressed constitutively, and AHL induction activates expression of both a UGPase-targeting synthetic sRNA and E. coli Hfq. The sRNA contains a 5′ sequence complementary to the UGPase mRNA RBS region and a 3′ Hfq-binding region, allowing Hfq-mediated translational repression of UGPase and thereby reducing cellulose production.
Note *Hfq is an RNA-binding protein that:
- stabilizes small RNAs (sRNAs)
- brings sRNA + target mRNA together
- helps them base-pair efficiently*
For this project, the output mechanism is kept as close as possible to the Florea design, but the input logic is rewired. Instead of using LuxR/pLux/AHL, the output will be placed under Opto-T7RNAP control. This means the final adapted output architecture is:
- PT7 → UGPase-targeting sRNA → terminator
- PT7 → E. coli Hfq → terminator
This preserves the original silencing logic while integrating it into a light-responsive genetic system intended for spatial control of cellulose production.
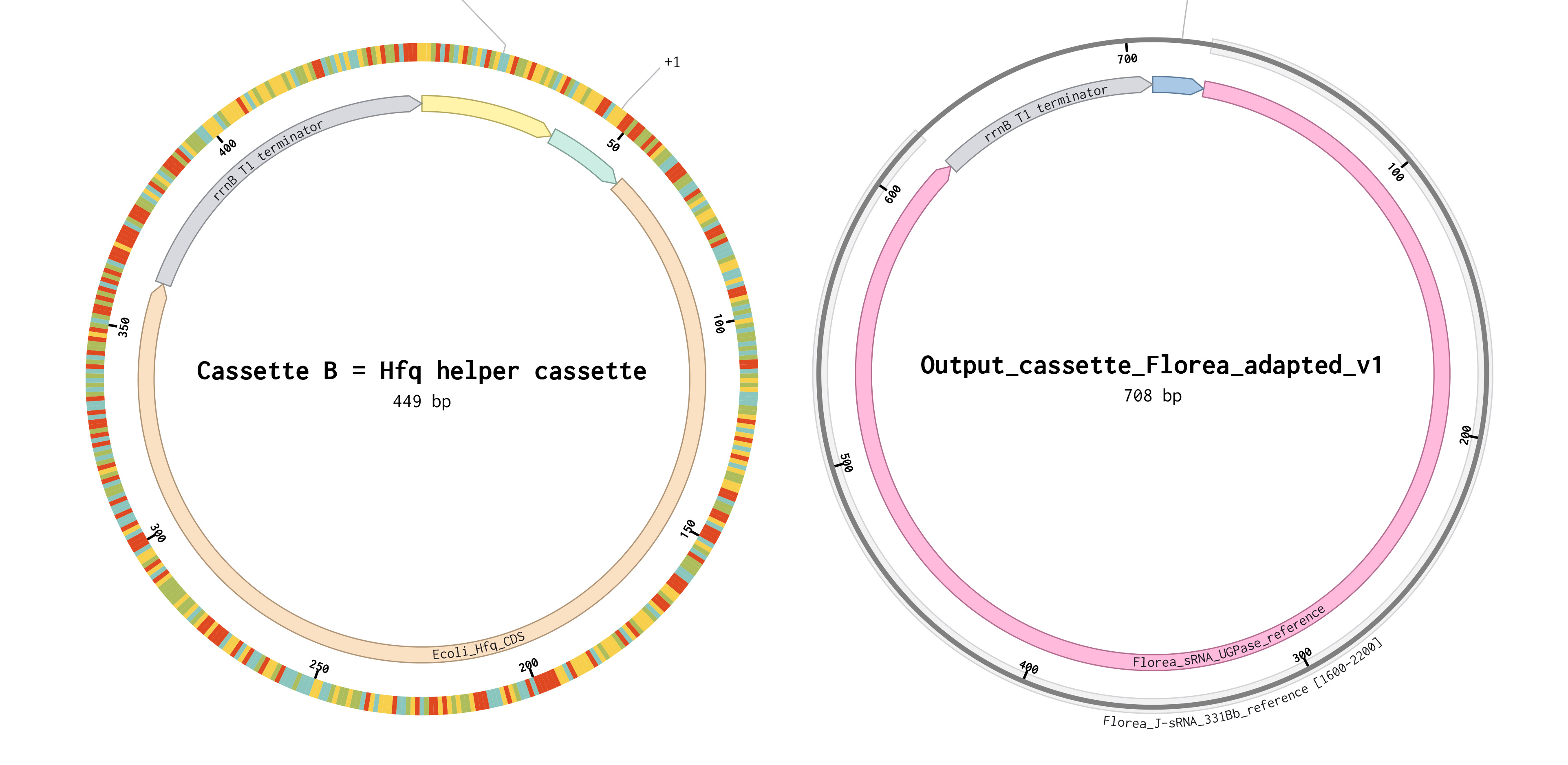
 Constructs are shown as circular maps for visualization purposes; all sequences were designed and ordered as linear DNA fragments.
Constructs are shown as circular maps for visualization purposes; all sequences were designed and ordered as linear DNA fragments.
Annotation note
Because the Registry / toolkit sequence is housed in the pSEVA331-Bb (a low-copy, stable plasmid) context and not presented in the paper as a fully base-resolved minimal feature map, the exact minimal nucleotide boundaries of the synthetic sRNA are not directly obvious from the paper figures alone. Therefore, this project reuses the published functional module logic from Florea rather than claiming a fully independently rederived minimal sRNA core from the schematic
So:
- minimal sRNA boundaries were not independently remapped base-by-base
- Hfq helper cassette was added as a separate constitutive module
- design is in silico and not yet experimentally validated
Note:
SEVA = Standard European Vector Architecture
Modular plasmid system designed for synthetic biology
Cassete B= Hfq helper cassete: https://benchling.com/s/seq-rCaVP0yYtscWeJatsMno?m=slm-2WFD8BIFwapoau6hUpCz
Output_sRNA_cassette: https://benchling.com/s/seq-84ojC9Fag5k6a36PuKDG?m=slm-M3qlgER1ZUeCJPtLXprw
DNA Synthesis Strategy
All four genetic constructs designed in this project were evaluated for synthesis feasibility using Twist Bioscience as linear double-stranded DNA fragments. The constructs were submitted as individual sequences in FASTA format without plasmid backbones, corresponding to modular expression cassettes.
The following fragments were included:
- OptoT7_half1 AraC/pBAD input cassette
- OptoT7_half2 AraC/pBAD input cassette
- PT7-driven sRNA output cassette targeting UGPase
- Constitutive Hfq helper cassette
All sequences were provided in 5′→3′ orientation and structured according to standard genetic architecture (promoter → functional element → terminator), with the exception of the sRNA cassette, which is a non-coding regulatory construct.
Note: A non-coding regulatory construct is a DNA module that produces RNA which regulates other genes, rather than being translated into a protein.
Synthesis Outcome
All four constructs passed automated synthesis screening and were accepted under standard synthesis conditions. The total estimated cost for synthesis was approximately $613.83, corresponding to four gene fragments ranging from 461 bp to 3455 bp in length. Minor warnings were observed for the sRNA cassette due to the absence of a conventional open reading frame (ORF), as the sequence encodes a functional regulatory RNA rather than a protein. These warnings did not affect synthesis feasibility and are expected for non-coding constructs.
Sequence Design Constraints Commercial DNA synthesis imposes several constraints on sequence composition, which were considered during construct design:
- GC Content Sequences must typically fall within a moderate GC range (~25–65%) to ensure efficient synthesis and amplification. All constructs in this project fall within acceptable GC ranges:
- sRNA cassette: ~43%
- Hfq cassette: ~47%
- OptoT7 constructs: ~50–52%
Repetitive Elements Long or highly identical repeats can lead to recombination or synthesis failure. The OptoT7 constructs contain a shared AraC/pBAD regulatory module (~1.2 kb), which introduces partial sequence redundancy between fragments. While this did not prevent synthesis acceptance, such repetition could pose challenges in downstream cloning or genomic integration and would typically be minimized in large-scale implementations.
Secondary Structure Strong secondary structures (e.g., hairpins in GC-rich regions) can interfere with oligo assembly. No significant secondary structure issues were flagged during synthesis screening, suggesting that all sequences are within acceptable structural complexity limits.
Homopolymer Runs Long stretches of identical nucleotides (e.g., AAAAAA) can reduce synthesis fidelity. No problematic homopolymer regions were identified in any of the constructs.
Coding vs Non-Coding Sequences Automated synthesis platforms often assume protein-coding sequences and may flag constructs lacking clear open reading frames. The PT7-sRNA output cassette triggered such warnings, as it is intentionally non-coding. This highlights a limitation of automated synthesis validation tools when applied to regulatory RNA systems.
Design Considerations and Trade-offs
Several design decisions influenced synthesis feasibility: Inclusion of AraC regulatory module The OptoT7 constructs include the full AraC/pBAD system rather than a minimal promoter. This increases construct size but ensures functional arabinose responsiveness. Separate Hfq expression cassette Hfq was implemented as an independent module to support sRNA function, increasing modularity at the cost of an additional fragment. Use of reference sRNA sequence The sRNA cassette was derived from a published system and not fully minimized, prioritizing functional reliability over sequence optimization.
Limitations and Future Optimization
While all constructs passed synthesis screening, several optimizations could improve robustness in future iterations: Reducing redundancy between OptoT7 constructs to minimize recombination risk Codon optimization of coding sequences for Komagataeibacter rhaeticus Minimization of non-essential regulatory regions Experimental validation of promoter activity and sRNA efficiency in the target organism
Conclusion
The successful synthesis validation of all four constructs demonstrates that the proposed optogenetic system is compatible with current commercial DNA synthesis capabilities. This supports the feasibility of translating the in silico design into a physical genetic system for experimental implementation.
24/4
Simulation of Pattern
Initial simulations were performed using periodic stripe patterns of varying feature sizes. The model shows that pattern fidelity strongly depends on feature size.
As shown in the figure :
- Small features (5 px ≈ 0.2 mm) disappear completely
- Intermediate features (20 px ≈ 0.8 mm) remain partially visible
- Large features (50 px ≈ 2.0 mm) are clearly preserved
Simulation of spatial pattern propagation for different feature sizes. Small features (~0.2 mm) are lost due to biological blur, while larger features persist.
Simulation of spatial pattern plotted agains real world data Walker, K. T. et al. (2025).
This demonstrates that the system exhibits a minimum resolvable feature size, below which spatial information is lost.
Effect of Biological Parameters
To understand what controls this resolution limit, a parameter sweep was performed across:
- Dark-state leakage (background gene expression)
- Biological blur (representing diffusion, growth, and material integration)
The resulting phase diagram shows the predicted minimum visible feature size (mm) across parameter space.
Key observations: Increasing leakage reduces contrast between patterned regions Increasing blur reduces spatial resolution through mixing The combination of both creates a sharp transition between:
- a regime where patterns are preserved
- a regime where patterns collapse
Predicted resolution limit as a function of dark-state leakage and biological blur. The red contour indicates the experimentally observed ~0.8 mm resolution regime.
Comparison to Experimental Work A reference contour (~0.8 mm) was overlaid based on previously reported optogenetic patterning in K. rhaeticus (tyrosinase-based pigment system).
The model predicts that this resolution lies along a continuum of parameter combinations, rather than representing a fixed system property.
Code: Stripe resolution simulation
Code: Resolution phase diagram with paper reference
Next planned experiements:
- Delay model The system is not instant: light → Opto-T7 activation → sRNA production → UGPase repression → cellulose change
- simulate a time delay between light pattern and material response.
- bacterial cellulose grows over days/weeks, while gene expression happens on minutes/hours.
- Layer / moiré simulation spatial structure, simulate sequential layers:
- layer 1: pattern A
- layer 2: shifted pattern B
- layer 3: rotated pattern C
- UGPase knockdown threshold model cellulose output depends on remaining UGPase activity. Example:
- 100% UGPase → normal cellulose
- 70% → slightly reduced
- 40% → visible reduction
- 10% → severe growth defect / too much stress
References
[1] Arnardottir, T. H. et al. Living Manufacture: Principles for a microbial 3D printer. download
[2] Goosens, V. J. et al. (2021).
https://doi.org/10.1021/acssynbio.1c00358
[3] Teh, M. Y. et al. (2019).
https://doi.org/10.1021/acssynbio.8b00168
[4] Walker, K. T. et al. (2025).
https://www.nature.com/articles/s41587-024-02194-3
[5] Levskaya, A. et al. (2005).
https://doi.org/10.1038/nature04405
[6] Florea, M. et al. (2016).
https://doi.org/10.1073/pnas.1522985113
[7] Zhou, H. et al. (2026).
https://doi.org/10.1016/j.tibtech.2025.09.019
[8] Loh, J. et al. (2025).
https://link.springer.com/article/10.1007/s10570-024-06367-w
[9] Cielecka, I. et al. (2020).
https://www.mdpi.com/2076-3417/10/11/3850
Full reff list can be found here: