Week 2 HW: DNA Read, Write, and Edit
Prelecture Homework:


In preparation for Week 2’s lecture on “DNA Read, Write, and Edit," please review these materials:
- Lecture 2 slides as posted below.
- The associated papers that are referenced in those slides.
- In addition, answer these questions in each faculty member’s section:
Homework Questions from Professor Jacobson:
- Nature’s machinery for copying DNA is called polymerase. What is the error rate of polymerase? How does this compare to the length of the human genome. How does biology deal with that discrepancy?
The biological machinery of copying DNA (polymerase) has an error rate of approximately 1 mistake per 10⁶ bases during replication when proofreading is active. (slide 8). This error is a variation based on the error rate, from 103 to 108. Compared to the length of the human genome, which is about 3.2 billion base pairs (≈3.2 × 10⁹ bp). This means that even with this high fidelity, thousands of errors could theoretically occur each time a genome is copied. (slide 10).
Biology addresses this discrepancy through multiple layers of error correction, including:
- Post-replication mismatch repair systems (such as MutS-based repair). (slide 14)
- Polymerase proofreading via 3′–5′ exonuclease activity.
- Additional cellular DNA repair pathways.
These mechanisms dramatically reduce the effective mutation rate, allowing organisms to maintain genomic stability despite the enormous size of their genomes.
- How many different ways are there to code (DNA nucleotide code) for an average human protein? In practice what are some of the reasons that all of these different codes don’t work to code for the protein of interest?
An average human protein requires approximately 1036 base pairs of DNA. (slide 6). It’s because the genetic code is degenerate. The majority of amino acids are encoded by multiple codons, which theoretically encode the same protein. However, in practice, not all of these sequences work well. Some reasons are:
- Codon predominance: cells prefer certain codons over others, affecting translation efficiency. (slide 34)
- GC content: extreme GC or AT richness can cause instability or poor expression. (slide 39)
- Secondary DNA/RNA structures: some sequences fold in ways that interfere with transcription or translation.
These constraints mean that although many DNA sequences could encode the same protein, only a small subset is biologically practical and manufacturable.
Homework Questions from Dr. LeProust:
What’s the most commonly used method for oligo synthesis currently? The most widely used method today is solid-phase phosphoramidite chemical synthesis, which was originally developed by Caruthers. (slide 10-11). In this approach, nucleotides are added one by one on a solid support through repeated cycles of coupling, capping, oxidation, and deprotection. This is the standard chemistry behind modern automated DNA synthesizers and high-throughput platforms, as reviewed on slides.
Why is it difficult to make oligos longer than 200nt via direct synthesis? Because each nucleotide addition is imperfect. Even with very high coupling efficiencies, small errors accumulate with every cycle in PCR. As length increases, the fraction of full-length, error-free molecules drops sharply. You also get more truncated products and substitutions, making purification harder and lowering overall yield. Practically, this limits reliable direct synthesis to ~150–200 nucleotides. (slides 36-39)
Why can’t you make a 2000bp gene via direct oligo synthesis? Because of the numbers of steps, if there is a 2000bp gene, the synthesis will take around 2000 steps. And at that scale, it’s probably to appear more chemical errors, full-length products become extremely rare, and the purity of the product will collapse. (slides 25-29).
To avoid synthesizing long genes directly, the standard strategy is: Use shorter bp (60-200nt) → assemble them enzymatically (PCR or gene assembly) → verify the final gene.
In result, the assembly reduces error and makes long genes.
Homework Question from George Church: [Using Google & Prof. Church’s slide #4] What are the 10 essential amino acids in all animals and how does this affect your view of the “Lysine Contingency”?
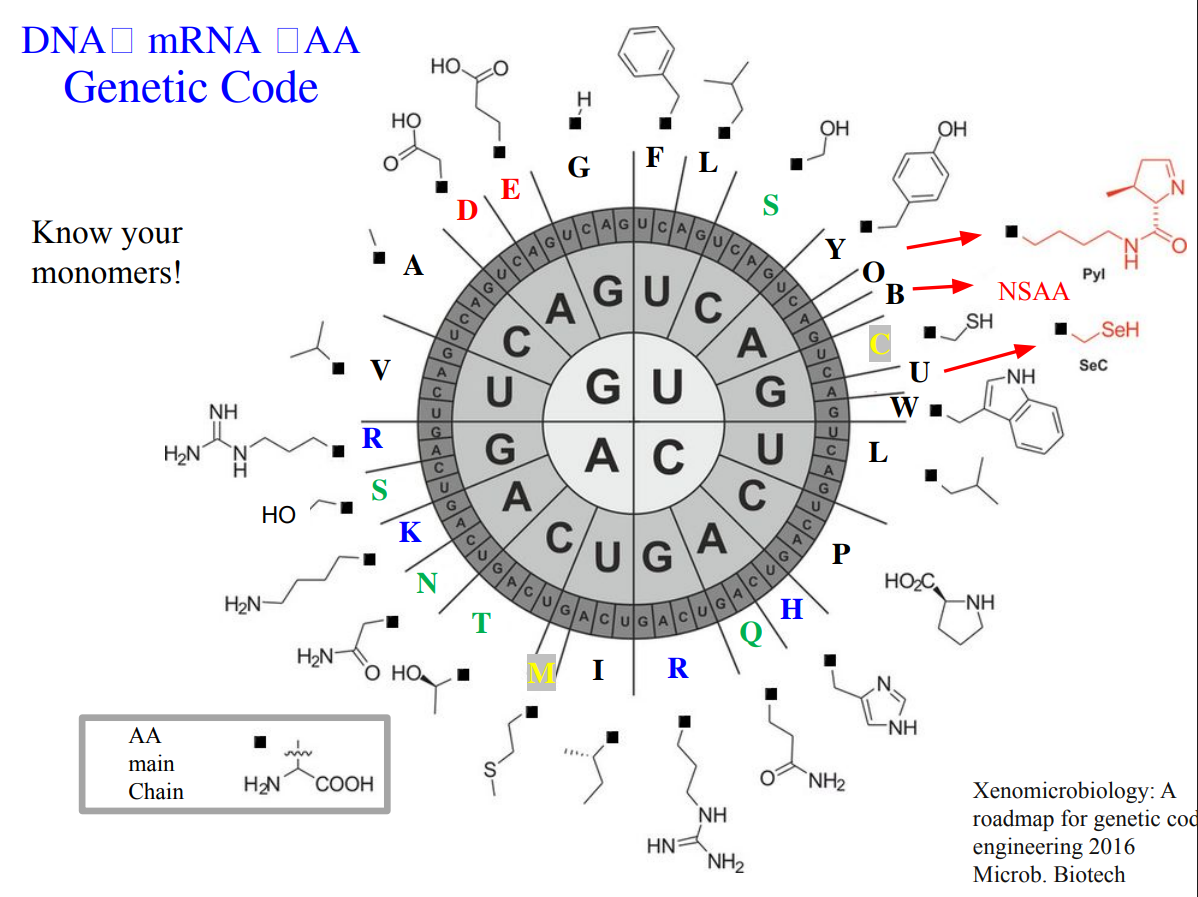

Essential aminoacid in animals
- Isoleucine, 2. Leucine, 3. Lysine, 4. Histidine, 5. Methionine, 6. Threonine, 7. Valine, 8. Arginine, 9. Tryptophan, 10. Phenylalanine Those amino acids are considered essential because animals cannot synthesize them de novo and must obtain them from dietary sources.
Usually, lysine is limited in plant-based diets and many agricultural feeds. So, lysine contingency highlights how biological systems, including humans and livestock, depend heavily on external lysine availability for protein synthesis, growth, and health. Because lysine cannot be synthesized by animals, entire food chains rely on microorganisms and plants capable of producing it.
In conclusion of 3 prelecture activities, those changed my view of genetic coding as not only an informational system but also an ecological dependency network. As well, to understand the limitations and how technology advances for creating solutions and continue researching.
Thanks for reading. For more information, there is my Notion webpage with the homework Notion prelecture week 2