Week 3 HW: Lab Automation
Week 3: Lab Automation


Part 1: Phyton Code & Agar Design


Documentation:
- For the first part of the Lab Automation assignment, I worked with Opentrons Python code using Google Colab. During this process, I used ChatGPT primarily as a debugging and learning aid. It helps me resolve execution errors, install missing packages (via pip), and understand how to structure the notebook so the design can be visualized correctly.
- Because the shared notebook relies on Opentrons hardware-specific functions (such as load_labware), the code was adapted to allow local visualization without a physical robot. My draft version originally included labware definitions intended for real laboratory execution, but these were temporarily removed to enable Plotly-based visualization.
- If you are interested in reading my code, please enter the following link: https://colab.research.google.com/drive/18Pb0JAgtB5Sv8v3VHhfop3mpF-nUiMp8?usp=drive_link
- The agar design was inspired by the ducks from Spirited Away (Studio Ghibli), based on my own drawing, combined with online references.
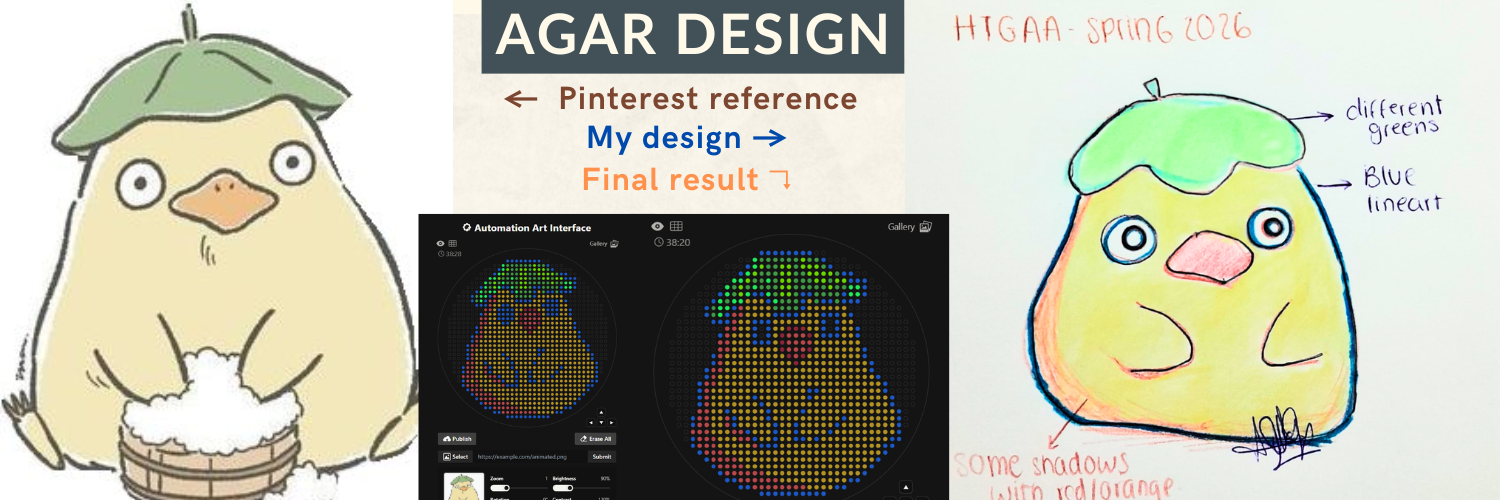

The final pixel-art layout was generated using the Opentrons Art Generator and can be viewed here: https://opentrons-art.rcdonovan.com/?id=5s7w0mpt758a7af
Code Building Pipeline:
To make the workflow clearer, the notebook was divided into three logical blocks:
flowchart TD
A[OpentronsMock Definition] --> B[Main Protocol Code]
B --> C[Visualization with Plotly]
- Block 1: Defines the virtual Opentrons environment and data recording
- Block 2: Executes the dispensing logic and color mapping
- Block 3: Displays the final agar pixel-art model
1. Opentrons Mock Definition: This block defines a mock version of the Opentrons protocol (OpentronsMock). Its purpose is to simulate robot behavior and record dispensing coordinates, enabling visualization without physical hardware. This block also sets up Plotly for graphical rendering.
2. Main Protocol Code: This is the core of the script, where:
- Color sources are assigned
- Coordinate points are paired with each fluorescent protein
- The virtual pipette iterates through each point set
- Dispensing actions are simulated
For visualization purposes, hardware-specific commands (such as load_labware) were removed in this version. The original draft protocol made for real robot execution is documented separately in “draft” inside the code.
3. Visualization: This final block executes the protocol and renders the design using Plotly. Here, all recorded coordinates are plotted, allowing inspection of:
- Spatial accuracy
- Color placement
- Overall agar pattern
This step is essential to verify that the design prints correctly before transferring it to a real Opentrons workflow. As well, the final result of the visualization is in the next image:
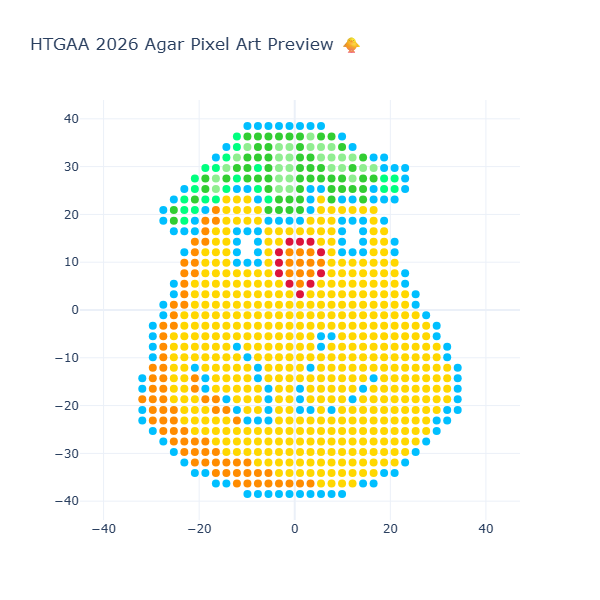

Part 2: Post-Lab Questions
Part 2.1: Research Paper automation application:
Scientific paper: “Technical upgrade of an open-source liquid handler to support bacterial colony screening” Available in: https://pmc.ncbi.nlm.nih.gov/articles/PMC10315574/
General view: This paper presents COPICK, a technical modification of the open-source Opentrons OT-2 liquid handling robot to automate bacterial colony screening. Colony picking is traditionally a labor-intensive bottleneck in genetic engineering workflows, especially when screening large numbers of variants generated by high-throughput DNA assembly. While commercial colony pickers exist, their high cost limits accessibility for smaller laboratories. COPICK addresses this limitation by integrating image acquisition and artificial intelligence into an affordable OT-2 platform.
The system combines a mounted USB camera with a Detectron2-based panoptic segmentation model to identify bacterial colonies directly from Petri dish images. The inference engine processes raw images, performs pixel- and object-level classification, and maps detected colony coordinates into the physical space of the robot. The OT-2 pipette then autonomously selects colonies based on user-defined criteria such as size, color, or fluorescence intensity. This integration enables on-board automated colony selection without the need for expensive commercial equipment.
Findings:
- Benchmark experiments performed with E. coli and P. putida demonstrated reliable performance across different screening scenarios (raw picking, color-based selection, and fluorescence-based cherry picking).
- COPICK achieved a raw performance of 73% over total screened colonies, increasing to 82% when considering only pickable colonies.
- The system showed high sensitivity (92%) and acceptable precision (78%), validating its potential as a cost-effective automation tool.
- Even if the classification errors existed in the model, the study suggests that performance could further improve using next-generation segmentation models such as SAM.
Why is it a novel application? I found this paper interesting, with a novel application for biology. First, COPICK reduces human bias and variability in colony selection by replacing manual visual inspection with algorithm-based inference. Also, the integration of AI-driven image segmentation with robotic actuation creates a reproducible, scalable workflow for microbial screening. And this approach democratizes high-throughput synthetic biology by making automated colony picking accessible to smaller laboratories, expanding the reach of biofoundry-style workflows.
Figures:
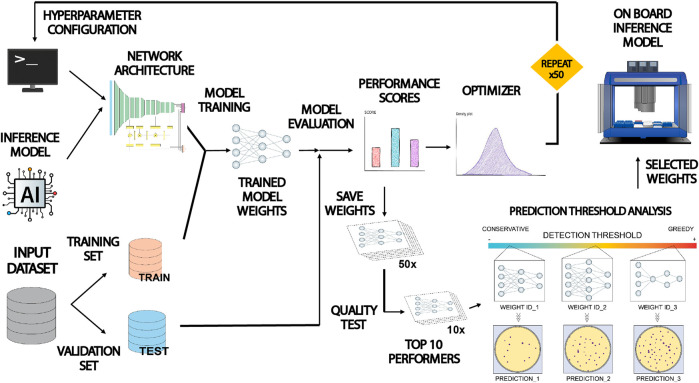
 Figure 3 from (Del Olmo Lianes et al., 2023), It shows the workflow diagram of the paper
Figure 3 from (Del Olmo Lianes et al., 2023), It shows the workflow diagram of the paper
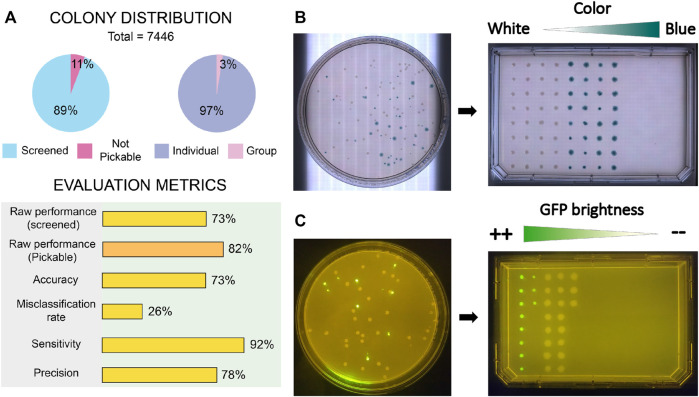
 Figure 7 (Del Olmo Lianes et al., 2023) shows the results, including the performance metrics that validate the assays.
Figure 7 (Del Olmo Lianes et al., 2023) shows the results, including the performance metrics that validate the assays.
Part 2.2: Application of Automation in Final Project:
Idea: Automated Screening of Lactate Biosensor Constructs using Cell-Free Systems
This idea comes from my W1 homework, where I propose to create a waterproof lactate biosensor tattoo for competition swimmers. I want to automate the screening of genetic lactate biosensor variants using cell-free protein synthesis (CFPS) in a 96-well plate. This will help with the optimization before proving it in vivo. Automation will be used to:
- Screen multiple lactate-responsive genetic constructs
- Test different lactate concentrations
- Quantify fluorescence output
- Select the best-performing biosensor variants
Flowchart
flowchart TD
A[Automated Workflow] --> B[Dispense CFPS master mix into 96-well plate]
B --> C[Add biosensor DNA variants]
C --> D[Apply lactate gradient 0–20 mM]
D --> E[Incubate at 37 °C]
E --> F[Measure fluorescence]
F --> G[Analyze response curves]The goal is to identify the most sensitive and dynamic lactate-responsive construct.
Possible pseudocode
Disclaimer: this mini pseudocode was created with IA’s help– ChatGPT 5.2
This idea was inspired by: (Jia et al., 2013); (Ghaffari et al., 2021); (Schmiedeknecht et al., 2022)
Part 3: Slides for final project:
My ideas for the project are: Main Idea: waterproof lactate biosensor tattoo for competition swimmers
I propose developing a semi-permanent, waterproof biosensor tattoo that detects lactate levels in athletes during pool training. The system would rely on engineered biological circuits that respond to lactate and trigger a visible fluorescent or colorimetric signal, functioning as a traffic-light-style, semi-quantitative indicator of physiological stress.
The idea is connected to course topics such as genetic circuit design and fluorescent protein signaling. Lactate would act as the biological input, while the output would be a color change generated by chromoproteins or fluorescent reporters, similar to the chromophore and genetic circuit.
Second idea: based on the toehold switch in biosensors: mRNA of biofilm formation on kitchen elements, the idea is to create a biosensor that detects biofilm formation in kitchen surfaces or utensils before it matures, like a pH paper or a device
Third idea: Creating a Biopatch of Metformin, where the delivery of metformin is better, also targeting Type 2 diabetes patients and patients with gastrointestinal intolerance to oral metformin
Link for final project slides: Final project slides ideas look for: 2026-a-ana-gomez | or Biopunk (updated!)


Weekly reflection:
This week was especially enjoyable because I got to design agar art in silico, which felt like a creative way to engage with lab automation concepts. While looking for a research paper, I was reminded of a researcher whose work uses algorithms from a different biological angle (using math algorithms to scan spheres that are attached to cells and visualize where the cancer cells are), and that made me realize how many areas of biology could benefit from automation in the future. I also noticed that my project ideas have been changing as I learn more about the course topics, which feels like part of the learning process itself. Overall, this week helped me reflect on how my interests are evolving, and it motivated me to keep exploring new perspectives and projects as I continue in the course.
Thank you for reading my weekly assignment! If you are interested in reading my Notion website, please enter the following link: https://www.notion.so/Assignment-Week-3-31125717f670808db22fc0687c7f7b19?source=copy_link
References and resources:
Part 2: Del Olmo Lianes, I., Yubero, P., Gómez-Luengo, Á., Nogales, J., & Espeso, D. R. (2023). Technical upgrade of an open-source liquid handler to support bacterial colony screening. Frontiers in bioengineering and biotechnology, 11, 1202836. https://doi.org/10.3389/fbioe.2023.1202836
Ghaffari, R., Yang, D. S., Kim, J., Mansour, A., Wright, J. A., Jr, Model, J. B., Wright, D. E., Rogers, J. A., & Ray, T. R. (2021). State of Sweat: Emerging Wearable Systems for Real-Time, Noninvasive Sweat Sensing and Analytics. ACS sensors, 6(8), 2787–2801. https://doi.org/10.1021/acssensors.1c01133
Jia, W., Bandodkar, A. J., Valdés-Ramírez, G., Windmiller, J. R., Yang, Z., Ramírez, J., Chan, G., & Wang, J. (2013). Electrochemical Tattoo Biosensors for Real-Time Noninvasive Lactate Monitoring in Human Perspiration. Analytical Chemistry, 85(14), 6553-6560. https://doi.org/10.1021/ac401573r
Schmiedeknecht, K., Kaufmann, A., Bauer, S., & Solis, F. V. (2022). L-lactate as an indicator for cellular metabolic status: An easy and cost-effective colorimetric L-lactate assay. PLoS ONE, 17(7), e0271818. https://doi.org/10.1371/journal.pone.0271818
Additional paper
Peñaherrera-Pazmiño, A. B., Isa-Jara, R. F., Hincapié-Arias, E., Gómez, S., Belgorosky, D., Agüero, E. I., Tellado, M., Eiján, A. M., Lerner, B., & Pérez, M. (2024). AQSA—Algorithm for Automatic Quantification of Spheres Derived from Cancer Cells in Microfluidic Devices. Journal of Imaging, 10(11), 295. https://doi.org/10.3390/jimaging10110295