week-07-hw-genetic-circuits-part-II
Week 7

Part 1: Intracellular Artificial Neural Networks
1. Advantages of IANNs vs traditional genetic circuits
Traditional genetic circuits usually behave like Boolean logic systems (ON/OFF), meaning they respond in discrete states (e.g., gene expressed or not). In contrast, IANNs offer several key advantages:
| Criteria | Description |
|---|---|
| Graded responses instead of binary outputs | IANNs can process inputs in a continuous manner (like real neural networks). This allows more nuanced control of gene expression |
| Integration of multiple inputs simultaneously | Instead of simple AND/OR logic, IANNs can weigh inputs differently (e.g., X1 contributes more than X2) |
| Higher computational complexity | They can approximate nonlinear functions and make more sophisticated “decisions” inside cells |
| Scalability | Multilayer architectures allow hierarchical information processing, similar to deep learning |
| Better noise tolerance | Weighted systems can be more robust to biological variability compared to strict Boolean thresholds |
(Nilsson et al., 2022; Müller et al., 2025)
2. Application of an IANN
For example, it could be a “Smart infection-detection system,” where the goal is to engineer a cell that detects early-stage infection and produces a therapeutic or reporter signal.
How it would work?
Inputs:
- X1: Presence of bacterial quorum sensing molecules (e.g., AHLs)
- X2: Host inflammation marker (e.g., ROS levels)
- X3: pH changes (acidic microenvironment)
Processing (IANN behavior):
- Each input is weighted differently
- The network integrates signals:
- High AHL + moderate ROS → strong activation
- Low AHL + high ROS → weak activation
- Uses a threshold function to decide output intensity
Output:
Expression of:
- Fluorescent protein (diagnostic)
- OR antimicrobial peptide (therapeutic)
There are some limitations in the application process, for example:
- Noise in gene expression
- Difficult tuning of weights (promoter strength, RBS, degradation rates)
- Crosstalk between biological components
- Metabolic burden on the host cell
- Limited dynamic range compared to electronic systems
3. Multilayer perceptron (conceptual diagram)
Before explaining the Multilayer perceptron (as a conceptual diagram), it’s important to understand how it works for a single-layer perceptron.
Single-layer perceptron:
The diagram represents an intracellular single-layer perceptron where:
- Input X1 encodes the Csy4 endoribonuclease, which acts as a negative regulator by cleaving target mRNA.
- Input X2 encodes a fluorescent protein, whose expression is regulated at the RNA level by Csy4.
Csy4 functions as a biological weight, modulating the effective expression of the output gene. The final fluorescence output depends on the balance between transcription of the fluorescent protein and post-transcriptional repression by Csy4.
This system mimics a perceptron where:
flowchart TD
A[Single-layer perceptron] --> B(1: X1 contributes a negative weight)
B --> C(2: X2 contributes a positive signal) --> c[3: The output is a graded fluorescence response]Multi-layer perceptron:
A multilayer intracellular perceptron can be constructed by cascading regulatory layers:
- In the first layer, inputs (X1 and X2) produce different endoribonucleases (e.g., Csy4 variants) that regulate the expression of a second-layer regulator.
- The hidden layer output is another endoribonuclease, which integrates the first-layer signals.
- In the second layer, this regulator controls the expression of a fluorescent protein.
This architecture allows hierarchical processing, where intermediate regulators act as hidden nodes, enabling more complex and nonlinear decision-making compared to a single-layer system.
Conceptual Diagram:
🧠 Code info (click to expand)
Diagram:
flowchart LR
subgraph I["Input layer"]
X1["X1"]
X2["X2"]
end
subgraph L1["Layer 1"]
A["DNA → Tx/Tl → Csy4-A"]
B["DNA → Tx/Tl → Csy4-B"]
end
subgraph H["Hidden layer"]
C["Regulated transcript<br/>DNA → Tx/Tl → Csy4-C"]
end
subgraph O["Output layer"]
D["Regulated fluorescent protein mRNA"]
Y["Fluorescence output"]
end
X1 --> A
X2 --> B
A -- "RNA cleavage/regulation" --> C
B -- "RNA cleavage/regulation" --> C
C -- "RNA cleavage/regulation" --> D
D --> YDisclaimer: For the creation of the Multi-layer perceptron diagram it was used ChatGPT 5.2.


Part 2: Fungal Materials
1. Examples of fungal materials
Fungal materials are formed through the self-assembly of mycelial networks, which bind organic substrates into cohesive and structured biomaterials. These networks enable the formation of diverse materials with applications ranging from packaging to advanced functional systems. As shown in Figure 1, mycelium-based materials can be engineered into different formats depending on their processing and intended use.
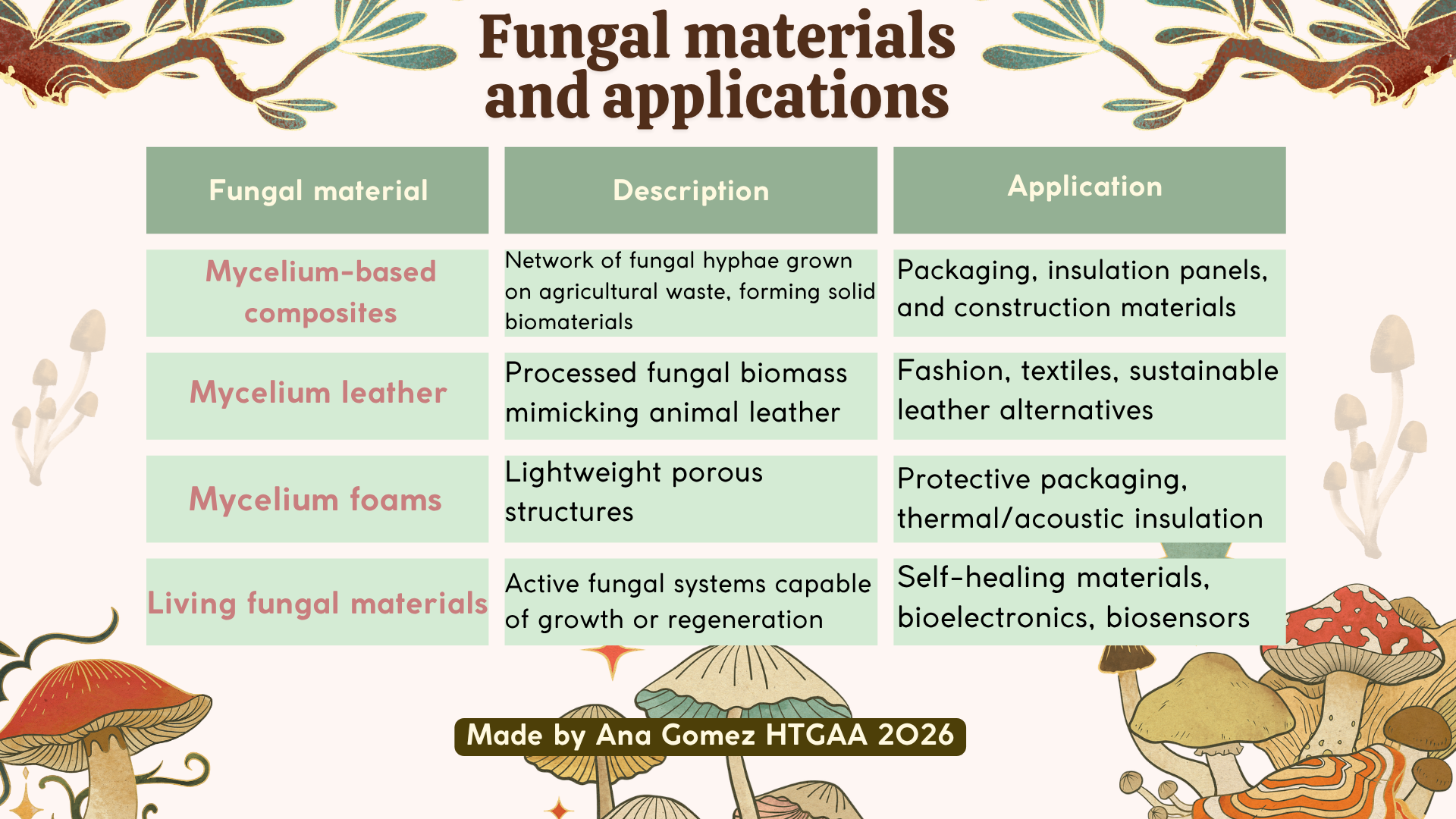

Figure 1. Fungal materials and applications table. (Based on Sharma et al., 2026)
Fungal materials offer several advantages, including low production cost, sustainability, and reduced environmental impact. Notably, during their growth phase, fungal systems can contribute to carbon sequestration. However, these benefits are accompanied by important limitations, such as susceptibility to degradation and moisture sensitivity, which can restrict their use in certain applications. These trade-offs are summarized in Table 1, which highlights both the advantages of fungal materials compared to traditional materials and their inherent limitations.
Table 1. Properties, advantages, and limitations of fungal materials compared to traditional materials
| Property | Fungal materials (advantage) | Compared to traditional materials | Limitation | Explanation |
|---|---|---|---|---|
| Sustainability | Biodegradable and low environmental impact | Plastics and synthetic materials are non-biodegradable and polluting | Limited durability | Faster degradation reduces lifespan in long-term applications |
| Production process | Grown from agricultural waste with low energy input | Conventional materials require energy-intensive industrial processes | Scalability challenges | Difficult to standardize growth conditions at industrial scale |
| Density and weight | Lightweight and porous structure | Concrete and polymers are often denser and heavier | Lower mechanical strength | Not suitable for load-bearing structures |
| Carbon footprint | Can sequester CO₂ during growth | Traditional materials often emit CO₂ during production | Limited structural performance | Trade-off between sustainability and strength |
| Customization | Can be molded during growth into complex shapes | Requires machining or molding after production | Growth variability | Results depend on environmental conditions (humidity, nutrients) |
| Biological activity | Can be functionalized (self-healing, sensing, antimicrobial) | Traditional materials are inert | Stability issues | Living or semi-living systems may change over time |
| Resource efficiency | Uses renewable substrates (e.g., lignocellulosic waste) | Relies on fossil-based or mined resources | Moisture sensitivity | High water absorption can compromise integrity |
| End-of-life impact | Fully compostable and circular | Waste accumulation and landfill persistence | Risk of contamination | Susceptible to microbial degradation if not treated |
Description: These trade-offs highlight the need for further optimization through material engineering and synthetic biology approaches, particularly to improve mechanical strength, stability, and scalability. Information based on (Alemu et al., 2022; Xia, 2024; Bitting et al., 2022; Parhizi et al., 2025)
2. Genetic engineering in fungi
Fungi represent a promising platform for synthetic biology due to their natural ability to grow as interconnected networks and secrete a wide range of enzymes. These characteristics make them particularly suitable for the development of functional and adaptive biomaterials. Through genetic engineering, fungi can be designed to perform specific tasks that enhance their utility in material science and environmental applications.
Potential applications of engineered fungi include:
- Self-healing materials → Fungi that regrow after damage
- Bioremediation → Degradation of plastics, hydrocarbons, or toxins
- Responsive materials → Materials that change color or fluorescence in response to stimuli
- Antimicrobial surfaces → Production of antifungal or antibacterial compounds
In addition, fungi offer several advantages over bacteria for material-based applications. As summarized in Figure 2, fungi are capable of forming multicellular, macroscopic structures through mycelial networks, which enables the development of biomaterials at larger scales. In contrast, bacteria are primarily suited for molecular-level engineering due to their unicellular nature.
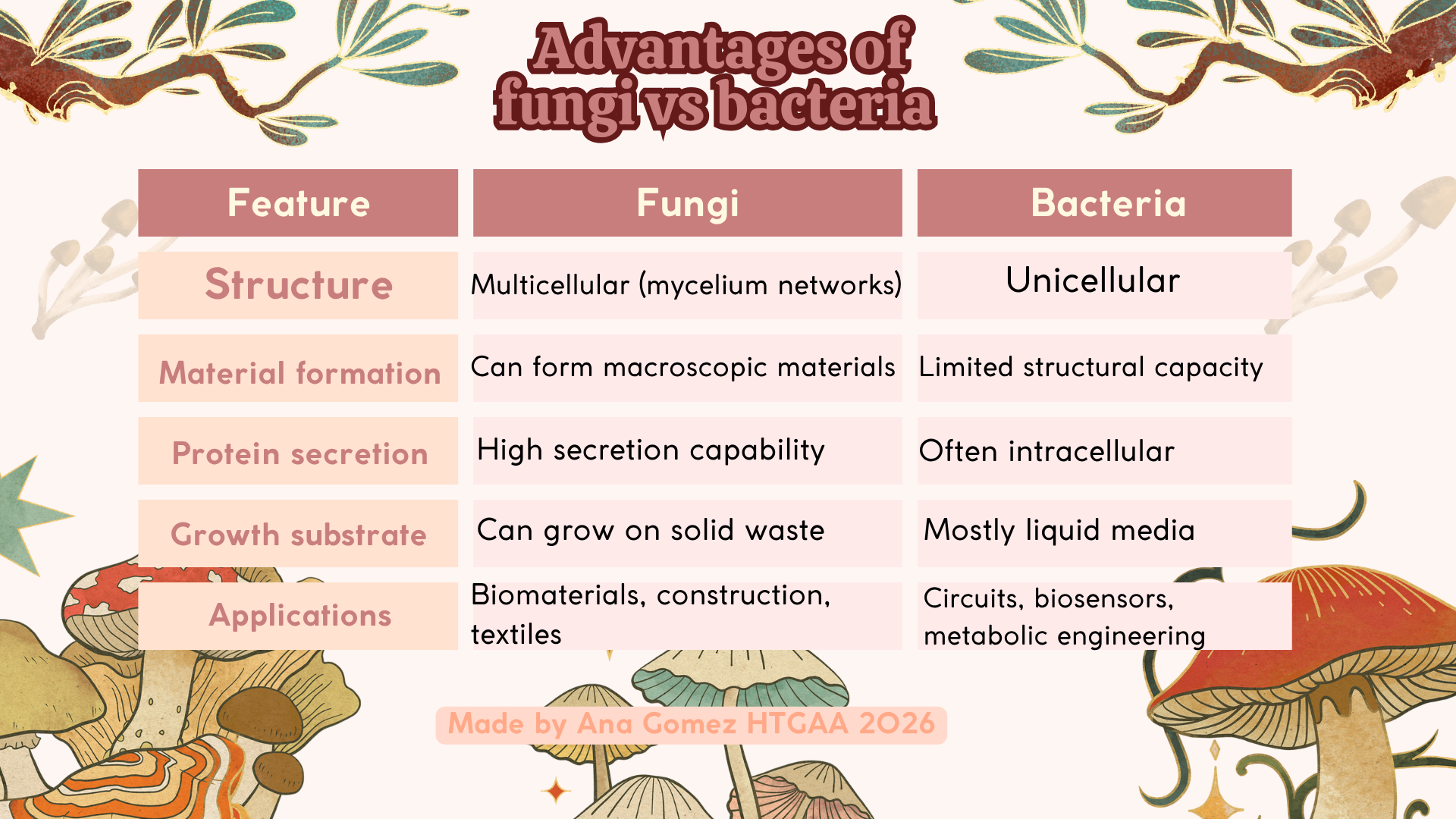

Figure 2. Advantage of Fungi vs. Bacteria table. Based on (Li et al., 2024; Pérez-Pazos et al., 2024)
Overall, fungal materials represent a promising platform for sustainable and programmable biomaterials. Their unique ability to grow, self-assemble, and interact dynamically with their environment positions them as a powerful alternative to traditional materials, particularly when combined with synthetic biology strategies.


Part 3: Individual projects!
Final Idea and first draft!
Title: KitBi. An Early-Warning Fluorescent Biosensor for Early Biofilm Commitment on Food-Contact Surfaces
Summary:
KitBi is a synthetic biology early-warning biosensor designed to report early biofilm commitment on food-contact surfaces before mature biofilm establishment. The project uses a promoter associated with biofilm regulation, such as PcsgD, driving sfGFP expression in non-pathogenic E. coli K-12. The goal is to shift from post-formation eradication to earlier risk detection, especially for Gram-negative foodborne contamination contexts relevant to stainless-steel and kitchen surfaces. Initial validation will be performed in silico through DNA design and simulation, with future translation toward portable or cell-free formats.
Validation- Benchling
Insert designs:
- First design:
- Second design:
Aim 1 draft
The first aim of my final project is to design and computationally validate a biofilm-responsive DNA construct in non-pathogenic E. coli that produces a fluorescent signal under early biofilm-inducing conditions relevant to food-contact surfaces, using Benchling for DNA construct design and Asimov Kernel for expression simulation.


Weekly Reflection:
This week felt a bit different because the concepts (IANNs and fungal materials) were interesting, but at first, they didn’t feel very connected to my project.
At the beginning, I honestly struggled with IANNs:
- They felt very abstract and kind of far from real applications
- The idea of implementing neural networks inside cells sounded cool, but also complicated
- I wasn’t sure how to connect that to what I’m doing
But after thinking about it more, I did take away something important:
- Biological systems don’t always work in simple ON/OFF logic
- They can integrate signals in a more gradual and layered way
- That actually relates to how biofilm-related promoters behave
Even if I’m not directly using IANNs in my design, it changed how I think about:
- promoter strength
- signal integration
- and how cells “decide” to activate certain pathways
For fungal materials, I found it way more intuitive and honestly really interesting:
| - Fungi can form actual macroscopic structures (not just molecular systems) |
| - Their mycelium works like a natural network that can bind materials together |
| - This made me think of biology not just as sensing, but also as material design |
- Biofilms forming on plastic in marine environments - These systems include bacteria, fungi, and other organisms
- What if those biofilm-forming organisms could be engineered? - Instead of just colonizing plastic, they could actually degrade it
It’s still a very early idea, but I liked that perspective:
→ biofilms are not just a problem, they could also be part of the solution
In terms of my project, this week was also important because I finalized my main idea: KitBi, an early-warning biosensor for biofilm formation. I initially struggled with deciding whether my idea was sufficiently innovative or too simple compared to other approaches. However, I realized that focusing on early detection rather than eradication aligns strongly with my background in biofilm research and gives the project a clear and meaningful direction. Choosing a problem that I understand well has made the design process more grounded and feasible.
Overall, this week helped me move from uncertainty to clarity. While some concepts remain challenging, I now feel more confident about my project direction and how it connects to broader themes in synthetic biology, such as sensing, regulation, and the design of living systems for real-world applications.
Thanks for reading it! This information is also in my personal Notion webpage, you can check it in: Notion- Week 7
References & sources:
PART 1
Cai, Y., Wang, Y., & Hu, S. (2025). Synthetic Gene Circuits Enable Sensing in Engineered Living Materials. Biosensors, 15(9), 556. https://doi.org/10.3390/bios15090556
Müller MM, Arndt KM and Hoffmann SA (2025) Genetic circuits in synthetic biology: broadening the toolbox of regulatory devices. Front. Synth. Biol. 3:1548572. https://doi.org/10.3389/fsybi.2025.1548572
Nilsson, A., Peters, J. M., Meimetis, N., Bryson, B., & Lauffenburger, D. A. (2022). Artificial neural networks enable genome-scale simulations of intracellular signaling. Nature communications, 13(1), 3069. https://doi.org/10.1038/s41467-022-30684-y
PART 2
Alemu, D., Tafesse, M., & Mondal, A. K. (2022). Mycelium-Based Composite: The Future Sustainable Biomaterial. International journal of biomaterials, 2022, 8401528. https://doi.org/10.1155/2022/8401528
Bitting, S., Derme, T., Lee, J., Van Mele, T., Dillenburger, B., & Block, P. (2022). Challenges and Opportunities in Scaling up Architectural Applications of Mycelium-Based Materials with Digital Fabrication. Biomimetics (Basel, Switzerland), 7(2), 44. https://doi.org/10.3390/biomimetics7020044
Gantenbein, S., Colucci, E., Käch, J., Trachsel, E., Coulter, F. B., Rühs, P. A., Masania, K., & Studart, A. R. (2022). Three-dimensional Printing of Mycelium Hydrogels into Living Complex Materials. arXiv (Cornell University). https://doi.org/10.48550/arxiv.2203.00976
Li, J., Yang, H., Duan, Y. Y., Sun, X. D., Pang, X. P., & Guo, Z. G. (2024). Fungi contribute more than bacteria to the ecological uniqueness of soil microbial communities in alpine meadows. Global Ecology and Conservation, 55, e03246. https://doi.org/10.1016/j.gecco.2024.e03246
Parhizi, Z., Dearnaley, J., Kauter, K., Mikkelsen, D., Pal, P., Shelley, T., & Burey, P. (2025). The Fungus Among Us: Innovations and Applications of Mycelium-Based Composites. Journal of Fungi, 11(8), 549. https://doi.org/10.3390/jof11080549
Pérez-Pazos, E., Beidler, K. V., Narayanan, A., Beatty, B. H., Maillard, F., Bancos, A., Heckman, K. A., & Kennedy, P. G. (2024). Fungi rather than bacteria drive early mass loss from fungal necromass regardless of particle size. Environmental microbiology reports, 16(3), e13280. https://doi.org/10.1111/1758-2229.13280
Sharma, M., Lim, L., & Kaur, G. (2025). Tailoring structure-property relationships of fungal mycelium for material applications: A process engineering approach for pure mycelium-based biomaterials. New Biotechnology, 91, 156–169. https://doi.org/10.1016/j.nbt.2025.12.006
Xia,Q. (2024). Utilizing mycelium-based materials for sustainable construction. Applied and Computational Engineering,63,10-15. **https://doi.org/10.54254/2755-2721/63/20240967**