week-09-hw-cell-free-systems
Week 9: Cell-Free systems!


Part A: General and Lecturer-Specific Questions
General questions:
- Explain the main advantages of cell-free protein synthesis over traditional in vivo methods, specifically in terms of flexibility and control over experimental variables. Name at least two cases where cell-free expression is more beneficial than cell production.
Cell-free protein synthesis (CFPS) offers important advantages over traditional in vivo expression because it provides a more open, flexible, and controllable reaction environment. Since there is no living cell to maintain, the researcher can directly adjust variables such as ionic strength, pH, redox conditions, DNA template concentration, cofactors, chaperones, detergents, lipids, or energy substrates without worrying about cell viability. CFPS is also typically faster, allowing protein production in hours rather than requiring cell growth, transformation, and induction steps over longer periods. In addition, it facilitates rapid prototyping of constructs and reaction conditions (Garenne et al., 2021; Jewett et al., 2008).
Another major advantage is that CFPS is particularly useful for proteins that are difficult to express in living cells, such as toxic proteins, membrane proteins, or proteins that require non-standard reaction environments. Because the system is open, reagents can be supplied directly and problematic cellular responses such as toxicity, growth inhibition, or proteolytic stress can be reduced (Garenne et al., 2021; Meyer et al., 2025).
Two cases where cell-free expression is more beneficial than cell-based production are:
| Cases | Description |
|---|---|
| 1) Toxic proteins | They may inhibit growth or kill the host cell during in vivo production (Chipman et al., 2025). |
| 2) Membrane proteins | CFPS allows co-translational insertion into detergents, nanodiscs, or liposomes under defined conditions, improving solubility and functional analysis (Meyer et al., 2025). |
- Describe the main components of a cell-free expression system and explain the role of each component
A cell-free expression system generally includes the following components:
| Component: | Description |
|---|---|
| 1) Cell extract or purified transcription–translation machinery | Provides ribosomes, translation factors, tRNAs, aminoacyl-tRNA synthetases, and often metabolic enzymes needed for protein synthesis. In extract-based systems, these components come from lysed cells; in reconstituted systems, they are added as purified factors. (1) |
| 2) DNA or mRNA template | Contains the coding sequence for the target protein and the regulatory elements needed for transcription and/or translation (1). |
| 3) Amino acids | Serve as the building blocks for protein synthesis (1). |
| 4) Nucleotides (ATP, GTP, CTP, UTP) | Required for transcription and for translation-associated energy consumption (1) |
| 5) Energy source and regeneration system | Maintains ATP and GTP availability during the reaction, which is essential because protein synthesis is highly energy demanding (2; 3) |
| 6) Salts and buffer components | Helps to keep suitable ionic strength and pH for enzyme activity and ribosome function, especially magnesium and potassium ions (3) |
| 7) Cofactors and additives | Include chaperones, disulfide-bond helpers, detergents, lipids, nanodiscs, or microsomes depending on the protein being expressed (4; 5) |
References 1. (Garenne et al., 2021); 2. (Jewett et al., 2008); 3. (Caschera, 2025); 4. (Harris et al., 2020); 5. (Meyer et al., 2025).
Additionally, a view of a CFPS by the article:
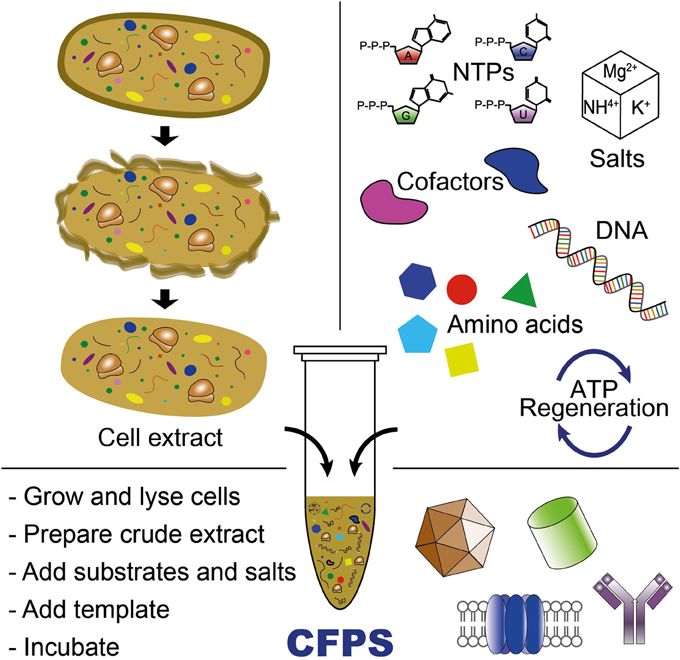

Figure 1. CFPS compounds from (Hong et al., 2014)
- Why is energy provision regeneration critical in cell-free systems? Describe a method you could use to ensure continuous ATP supply in your cell-free experiment.
Energy regeneration is critical in CFPS because transcription and translation consume large amounts of ATP and GTP. Without a continuous energy supply, the reaction quickly slows or stops, lowering protein yield. In addition, some simple high-energy substrates can accumulate inorganic phosphate, which chelates magnesium and impairs ribosomal activity, further reducing productivity (Yavad et al., 2025). One way to ensure continuous ATP supply is to use an ATP-regeneration system based on phosphoenolpyruvate (PEP), which donates phosphate groups for ATP resynthesis. Another effective strategy is to use maltodextrin/polyphosphate-based metabolism in crude extracts, which can support longer-lasting and more cost-effective ATP regeneration through endogenous metabolic enzymes (Caschera & Noireaux, 2015; Chen et al., 2019).
- Compare prokaryotic versus eukaryotic cell-free expression systems. Choose a protein to produce in each system and explain why.
In the following table, a comparison is made between different cell systems and the cell-free expression.
Table 1: Comparison of prokaryotic versus eukaryotic cell-free expression systems
| Feature | Prokaryotic Cell-Free System | Eukaryotic Cell-Free System | Example Proteins |
|---|---|---|---|
| Speed, cost & yield | Based on E. coli extracts; generally faster, cheaper, and higher yielding. Ideal for rapid screening and prototyping. | Typically slower, more expensive, and sometimes lower yielding compared to prokaryotic systems. | GFP or bacterial metabolic enzymes (efficient cytosolic expression). |
| Post-translational modifications (PTMs) | Limited capacity for PTMs; not suitable for complex modifications. | Capable of complex PTMs such as glycosylation and proper disulfide bond formation. | Glycosylated receptor fragments or secreted proteins. |
| Protein folding & complexity | Best suited for simple, soluble proteins; may struggle with complex folding. | Better suited for complex proteins requiring proper folding machinery. | Eukaryotic enzymes or multi-domain proteins. |
| Membrane protein expression | Limited ability; often requires artificial additives (detergents, liposomes). | More efficient due to the presence of microsomes and native-like membrane environments. | Eukaryotic membrane proteins (e.g., receptors, channels). |
Description: The table information was based on (Garenne et al., 2021; Jewett et al., 2008; Meyer et al., 2025), and (Fenz et al., 2014) for the eukaryotic field.
- How would you design a cell-free experiment to optimize the expression of a membrane protein? Discuss the challenges and how you would address them in your setup.
A rational way to design a cell-free experiment to express a membrane protein would be to optimize both the biochemical reaction conditions and the DNA construct. Based on the VDAC study by Zayni et al. (2021), I would use a prokaryotic cell-free expression system and focus particularly on the region surrounding the translation start site, since the paper showed that expression efficiency was mainly governed by translation initiation and mRNA conformation near the start codon, rather than by differences in transcription. (Zayni et al., 2021)
Steps:
- I would first select a plasmid, in this case a plasmid + T7 promoter and a properly positioned Shine-Dalgarno sequence.
- Then, I would evaluate the 5’ UTR and the first codons of the translated region. (Based on the paper of Zayni, it demonstrated that these sequence elements strongly influence whether the ribosome can properly dock and initiate translation) (Zayni et al., 2021)
- It is important to analyze the accessibility of the ribosome docking site and estimate the ΔEopen of the mRNA around the start codon.
- A lower ΔEopen would indicate that the ribosome-binding region is more accessible and therefore more likely to support efficient protein expression.
Optimization
To optimize the construct, I would test the following different design strategies:
- Adjusting the spacing between the Shine-Dalgarno sequence and the start codon.
- Adding a translation enhancer if native expression is too weak.
- If I want to preserve the native amino acid sequence, introducing synonymous mutations (Substitutions; DNA changes that alter a codon’s nucleotide sequence but not the resulting amino acid) (Oelschlaeger, 2024) in the first several codons to reduce the inhibitory mRNA secondary structures without altering the protein itself.
This last strategy in the paper aims to substantially improve VDAC expression while preserving the WT protein sequence.
Main challenges
The main challenges in this setup could be:
- Low translation efficiency: Caused by poor ribosome access to the start region.
- Non-native N-terminal additions: Enhancers like His-tags or CAT- derived sequences are used.
- Persistent low expression of membrane proteins, since these proteins are inherently difficult to produce.
I would address these by first identifying whether the limitation is transcriptional or translational. Since the paper showed that mRNA levels remained similar across poorly and well-expressed constructs, I would prioritize troubleshooting the translation-initiation region rather than assuming transcription is the problem. I would then redesign the coding sequence near the start codon to improve RDS accessibility, ideally using synonymous codon optimization.
- Imagine you observe a low yield of your target protein in a cell-free system. Describe three possible reasons for this and suggest a troubleshooting strategy for each.
If I observed a low yield of my target membrane protein in a cell-free system, three possible reasons would be:
Case 1. Poor translation initiation due to inhibitory mRNA secondary structure:
| Description: | Troubleshooting strategy: |
|---|---|
| A major reason for low yield could be that the region around the start codon is too structured, preventing proper ribosome docking. In the VDAC paper, constructs with similar transcription levels still showed very different protein yields, indicating that the bottleneck was translation initiation rather than mRNA production | I would redesign the sequence around the start codon, especially the first several codons of the translated region, to reduce the ΔEopen and improve accessibility of the ribosome docking site. This could be done with synonymous mutations so that the amino acid sequence remains unchanged |
Case 2. Suboptimal construct architecture near the 5′UTR and RBS
| Description: | Troubleshooting strategy: |
|---|---|
| Another reason could be an ineffective arrangement of the 5′UTR, Shine–Dalgarno sequence, and initiation codon. The study showed that even small differences in construct design near the start region changed VDAC expression substantially. It also found that the most favorable arrangement involved an optimal RBS-to-start codon spacing, around 11 nucleotides upstream in their model | I would test alternative plasmid designs with improved RBS positioning and compare constructs with or without translation-enhancing elements. If native expression remains poor, I could temporarily use an enhancer-containing construct for screening, then later optimize an enhancer-free native version |
Case 3. Inadequate reaction conditions for cell-free synthesis
| Description: | Troubleshooting strategy: |
|---|---|
| A third reason could be that the biochemical environment is not ideal for the system. The paper notes that the E. coli-based cell-free platform depends on appropriate biochemical conditions, including high T7 RNA polymerase activity and sufficient amino acid supply, especially for rapidly degraded amino acids. Even though the authors conclude that the mRNA sequence was more decisive than the biochemical conditions in their study, these conditions still matter for successful expression | I would verify reaction composition, template concentration, incubation time, and amino acid supply, and confirm that the chosen cell-free kit is appropriate for the membrane protein. I would also compare performance across different constructs under the same reaction conditions to distinguish sequence-related effects from reaction-related effects |
Additionally:
Click here to download the paper: Enhancing the Cell-Free Expression of Native Membrane Proteins by In Silico Optimization of the Coding Sequence-An Experimental Study of the Human Voltage-Dependent Anion Channel


Homework question from Kate Adamala
Based on my final individual project, KitBi, I am looking to detect early Gram-negative bacteria biofilms from kitchen surfaces and utensils in an easy, portable, economic, and quick method similar to a pH paper.
- Pick a function and describe it.
a. What would your synthetic cell do? What is the input, and what is the output?
- My synthetic cell would detect quorum-sensing molecules from Gram-negative bacteria before mature biofilm formation and convert that signal into a visible reporter output.
- Input: AHL molecules released by Gram-negative bacteria.
- Output: fluorescence or colorimetric signal produced by the synthetic cell. AHLs are a practical early target because they are extracellular signals associated with quorum sensing in Gram-negative bacteria, and quorum sensing is closely tied to virulence and biofilm-related behaviors.
(Lentini et al., 2014; Miller & Gilmore, 2020; Galloway et al., 2010)
b. Could this function be realized by cell-free Tx/Tl alone, without encapsulation?
Yes, theoretically. There are published cell-free biosensors for quorum-sensing molecules, so detection itself does not strictly require encapsulation. A paper by Wen et al. describes a cell-free biosensor for quorum-sensing biomarkers in infectious disease contexts.
Additionally, talking about the membrane, it gives more of an artificial cell logic, similar to Kate’s example: the vesicle becomes a defined sensing unit, can protect the Tx/Tl mix, and about selective exchange with the environment. Reviews on synthetic cell–living cell communication also support liposome-based systems as useful platforms for chemical communication.
(Mukwaya et al., 2021; Rampioni et al., 2019)
c. Could this function be realized by a genetically modified natural cell?
Yes. Natural bacterial biosensors for AHLs exist, and whole-cell AHL sensor systems are well established.
d. Describe the desired outcome of your synthetic cell operation.
In the presence of Gram-negative quorum-sensing signals, the synthetic cell turns on a reporter and gives an early warning that a biofilm-prone bacterial population may be emerging on the surface being tested. This follows the objectives of detecting early and intervening before eradication becomes harder. The review on quorum-sensing molecule detection explicitly frames QS signals as potentially useful early diagnostic indicators.
- Design all components that would need to be part of your synthetic cell.
a. What would the membrane be made of?
The membrane will be a phospholipid membrane with cholesterol, for example, POPC + cholesterol. Since this is a standard and defensible artificial-cell style membrane in liposome-based systems. Also, based on Lentini’s example, it used phospholipids plus cholesterol as a simple artificial-cell membrane concept.
b. What would you encapsulate inside? Enzymes, small molecules.
Inside the vesicle, I would encapsulate:
- a bacterial cell-free Tx/Tl system
- a DNA circuit containing an AHL-responsive transcription factor
- a reporter gene such as sfGFP or lacZ
This is realistic because cell-free AHL biosensing has already been demonstrated, and bacterial lysate-based cell-free systems are commonly used for such biosensors
(Wen et al., 2017; Didovyk et al., 2017)
c. Which organism your Tx/Tl system will come from? Is bacterial OK, or do you need a mammalian system for some reason? (hint: for example, if you want to use small molecule modulated promotors, like Tet-ON, you need mammalian)
A bacterial system, ideally E. coli-based, is the most reasonable choice here, because AHL quorum-sensing modules like LuxR/plux are bacterial and do not require a mammalian expression background.
d. How will your synthetic cell communicate with the environment? (hint: are substrates permeable? or do you need to express the membrane channel?)
It will follow the communication logic:
(Ding et al., 2014; Mukwaya et al., 2021)
- Experimental details
a. List all lipids and genes. (bonus: find the specific genes; for example, instead of just saying “small molecule membrane channel” pick the actual gene.)
For my project, I decided to focus on a liposome-based synthetic cell encapsulating an E. coli cell-free expression system and a LuxR-responsive reporter circuit to detect AHL molecules released by Gram-negative bacteria as an early warning of biofilm development.
Lipids:
- POPC
- Cholesterol
Genes:
- luxR
- sfGFP under a LuxR-activated promoter such as PluxI
(Lentini et al., 2014; Miller & Gilmore, 2020)
b. How will you measure the function of your system?
Fluorescence output from GFP, measured by plate reader, microscopy, or bulk fluorescence.
Homework question from Peter Nguyen
Freeze-dried cell-free systems can be incorporated into all kinds of materials as biological sensors or as inducible enzymes to modify the material itself or the surrounding environment. Choose one application field — Architecture, Textiles/Fashion, or Robotics — and propose an application using cell-free systems that are functionally integrated into the material. Answer each of these key questions for your proposal pitch:
For this section, I am exploring the fashion/textile option with UV clothes based on: (Lawrynowicz et al., 2024; Sąsiadek-Andrzejczak & Kozicki, 2023)
- Write a one-sentence summary pitch sentence describing your concept.
A sun-responsive shirt embedded with freeze-dried cell-free systems that activate under UV exposure, and produce a visible color change to reduce heat absorption and signal high solar intensity.
- How will the idea work, in more detail? Write 3-4 sentences or more.
The textile would contain microencapsulated, freeze-dried cell-free systems embedded within its fibers that respond to UV radiation. Upon exposure to sunlight, the system becomes activated (for example: heat or humidity) and produces a colorimetric output, where higher UV intensity generates a deep purple pigment, while lower exposure results in a softer pastel tone. This gradient response allows the textile to visually indicate different levels of solar exposure rather than a simple on/off signal. As a result, the material functions both as a real-time UV indicator and as an adaptive aesthetic element in fashion.
- What societal challenge or market need will this address?
This system addresses increasing UV exposure and heat stress, especially in regions like Latin America, especially here in Ecuador, where solar radiation is intense due to altitude.
High UV exposure is linked to skin damage and long-term health risks, but people often lack real-time awareness of exposure levels. A responsive textile could act as a personal UV sensor, helping individuals make better decisions about sun protection while also improving comfort and awareness.
- How do you envision addressing the limitations of cell-free reactions (e.g., activation with water, stability, one-time use)?
Table 2. Addressing limitations of cell-free systems
| Limitation: | Description: |
|---|---|
| Activation | Designed to activate with sweat or humidity instead of external water addition |
| Stability | Improved through lyophilization and stabilizers (e.g., trehalose) for long-term storage |
| One-time use | Implemented as microencapsulated, replaceable patches to allow renewal after activation |
| Reusability | Use of modular or layered textile design, where only sensing components are replaced |
| Washing limitations | Cell-free systems are water-sensitive, so patches should be removable before washing |
| Durability improvement | Protective encapsulation strategies could enhance resistance to moisture and extend usability |
Homework question from Ally Huang
Freeze-dried cell-free reactions have great potential in space, where resources are constrained. As described in my talk, the Genes in Space competition challenges students to consider how biotechnology, including cell-free reactions, can be used to solve biological problems encountered in space. While the competition is limited to only high school students, your assignment will be to develop your own mock Genes in Space proposal to practice thinking about biotech applications in space!
For this particular assignment, your proposal is required to incorporate the BioBits® cell-free protein expression system, but you may also use the other tools in the Genes in Space toolkit (the miniPCR® thermal cycler and the P51 Molecular Fluorescence Viewer). For more inspiration, check out https://www.genesinspace.org/ .
1. Provide background information that describes the space biology question or challenge you propose to address. Explain why this topic is significant for humanity, relevant for space exploration, and scientifically interesting. (Maximum 100 words)
Biofilms are a real challenge in spacecraft because microbes can colonize surfaces and water systems, threatening hardware reliability and potentially crew health. NASA documents note that biofilm formation has been observed in ISS systems, including water lines, where it contributed to clogging and pump issues. This is significant for humanity because long-duration missions will require reliable, low-resource methods for monitoring contamination. It is also scientifically interesting because microgravity and spaceflight conditions can alter microbial behavior, including biofilm-related traits.
Recommended lectures from NASA

 Figure 5. Microbial Research Guide from Colorado et al., 2021 Figure 5. Microbial Research Guide from Colorado et al., 2021 | 
 Figure 6. Conference of 2022 about Redefining Microbiological Risk Mitigation During Spaceflight from Ott, 2022 Figure 6. Conference of 2022 about Redefining Microbiological Risk Mitigation During Spaceflight from Ott, 2022 |
Link to access the following documents in the Sources section
2. Name the molecular or genetic target that you propose to study. Examples of molecular targets include individual genes and proteins, DNA and RNA sequences, or broader -omics approaches. (Maximum 30 words)
Biofilm- and quorum-sensing-related RNA targets from Gram-negative bacteria, such as luxS, lasI/lasR, or pslA transcripts
3. Describe how your molecular or genetic target relates to the space biology question or challenge your proposal addresses. (Maximum 100 words)
Biofilm formation does not begin as a visible layer; it starts with changes in gene expression and cell-cell signaling. Quorum-sensing and biofilm-associated transcripts can therefore serve as early molecular indicators of biofilm development before major fouling occurs. Detecting these RNA targets would help identify when bacteria are shifting from planktonic growth toward surface-associated communities, which is exactly the stage where intervention is most useful in spacecraft systems with limited maintenance capacity.
(Flores et al., 2024; Vélez et al., 2023)
4. Clearly state your hypothesis or research goal and explain the reasoning behind it. (Maximum 150 words)
I hypothesize that Gram-negative bacteria exposed to space-relevant stress conditions will show an increased abundance of quorum-sensing or biofilm-associated RNA targets compared with non-biofilm controls, and that these changes can be detected using a compact toolkit that combines miniPCR, BioBits®, and fluorescence readout.
- My goal is to develop an early-warning molecular screening strategy for spacecraft biofilm risk. The reasoning is that freeze-dried cell-free systems are portable and low-resource, while the Genes in Space toolkit already includes fluorescence-based tools designed for constrained environments.
If successful, this approach could support routine monitoring of microbial contamination during long-duration missions.
5. Outline your experimental plan - identify the sample(s) you will test in your experiment, including any necessary controls, the type of data or measurements that will be collected, etc. (Maximum 100 words)
I would test RNA extracted from Gram-negative bacterial cultures grown under a biofilm-promoting condition versus planktonic control cultures. miniPCR would amplify cDNA corresponding to selected biofilm-related targets, and BioBits® plus the P51 fluorescence viewer would be used to visualize signal output. Controls would include a no-template negative control and a non-biofilm bacterial condition. Data would consist of fluorescence presence or relative intensity across samples, indicating whether biofilm-associated targets are enriched under spacecraft-relevant stress conditions.


Part B: Homework Part B: Individual Final Project
Title:
KitBi: An Early-Warning Fluorescent Biosensor for Early Biofilm Commitment on Food-Contact Surfaces
Aims!
Aim 1 — Experimental aim
- Design and computationally validate a PcsgD-driven fluorescent reporter construct in non-pathogenic E. coli K-12 to detect early biofilm-inducing physiological states relevant to food-contact surfaces.
Aim 2 — Development aim
- Test the reporter under controlled early-attachment conditions using simulated or future experimental comparisons across planktonic, surface-exposed, and stainless-steel-associated growth states, and optimize the system with an internal normalization module or alternative promoters such as PcsgBAC.
Aim 3 — Visionary aim
- Translate KitBi into a portable early-warning platform for food-contact surface monitoring, potentially through freeze-dried, paper-based, or cell-free-compatible readouts that support preventive hygiene decisions before mature biofilm establishment.
Final Slide
Coming soon!


Weekly reflection:
This week made me realize how universal biofilms are. I initially thought of them mainly in food and clinical contexts, but learning about their presence in space environments really changed my perspective. It was surprising to see that biofilms can form even under microgravity conditions and still represent a risk for systems like water lines and surfaces in spacecraft.
This reinforced the relevance of my project, since early detection is not only important on Earth but also in highly controlled environments like space missions. It made me think that KitBi could have broader applications beyond food safety, especially in settings where prevention is critical and intervention is limited.
Thank you for reading! I updated this entry also on my personal Notion webpage. To check it, please enter here! Notion W9
References and sources
PART A:
General Questions:
Caschera, F. (2025). Cell-free protein synthesis platforms for accelerating drug discovery. Biotechnology Notes, 6, 126-132. https://doi.org/10.1016/j.biotno.2025.02.001
Caschera, F., & Noireaux, V. (2015). A cost-effective polyphosphate-based metabolism fuels an all E. coli cell-free expression system. Metabolic Engineering, 27, 29-37. https://doi.org/10.1016/j.ymben.2014.10.007
Chen, J., Mitra, R., Zhang, S., Zuo, Z., Lin, L., Zhao, D., Xiang, H., & Han, J. (2019). Unusual Phosphoenolpyruvate (PEP) Synthetase-Like Protein Crucial to Enhancement of Polyhydroxyalkanoate Accumulation in Haloferax mediterranei Revealed by Dissection of PEP-Pyruvate Interconversion Mechanism. Applied and environmental microbiology, 85(19), e00984-19. https://doi.org/10.1128/AEM.00984-19
Chipman, D. M., Woolley, A. C., Chau, D. N., Lance, W. A., Talley, J. P., Green, T. P., Robbins, B. C., & Bundy, B. C. (2025). Cell-Free Protein Synthesis Reactor Formats: A Brief History and Analysis. SynBio, 3(3), 10. https://doi.org/10.3390/synbio303001
Fenz, S. F., Sachse, R., Schmidt, T., & Kubick, S. (2013). Cell-free synthesis of membrane proteins: Tailored cell models out of microsomes. Biochimica Et Biophysica Acta (BBA) - Biomembranes, 1838(5), 1382-1388. https://doi.org/10.1016/j.bbamem.2013.12.009
Garenne, D., Haines, M.C., Romantseva, E.F. et al. Cell-free gene expression. Nat Rev Methods Primers 1, 49 (2021). https://doi.org/10.1038/s43586-021-00046-x
Harris, N.J., Pellowe, G.A. & Booth, P.J. Cell-free expression tools to study co-translational folding of alpha helical membrane transporters. Sci Rep 10, 9125 (2020). https://doi.org/10.1038/s41598-020-66097-4
Jewett, M.C., Calhoun, K.A., Voloshin, A. et al. An integrated cell‐free metabolic platform for protein production and synthetic biology. Mol Syst Biol 4, MSB200857 (2008). https://doi.org/10.1038/msb.2008.57
Meyer, C., Arizzi, A., Henson, T. et al. Designer artificial environments for membrane protein synthesis. Nat Commun 16, 4363 (2025). https://doi.org/10.1038/s41467-025-59471-1
Oelschlaeger P. (2024). Molecular Mechanisms and the Significance of Synonymous Mutations. Biomolecules, 14(1), 132. https://doi.org/10.3390/biom14010132
Yadav, S., Perkins, A. J. P., Liyanagedera, S. B. W., Bougas, A., & Laohakunakorn, N. (2025). ATP Regeneration from Pyruvate in the PURE System. ACS synthetic biology, 14(1), 247–256. https://doi.org/10.1021/acssynbio.4c00697
Zayni, S., Damiati, S., Moreno-Flores, S., Amman, F., Hofacker, I., Jin, D., & Ehmoser, E. K. (2021). Enhancing the Cell-Free Expression of Native Membrane Proteins by In Silico Optimization of the Coding Sequence-An Experimental Study of the Human Voltage-Dependent Anion Channel. Membranes, 11(10), 741. https://doi.org/10.3390/membranes11100741
Kate Adamala:
Didovyk, A., Tonooka, T., Tsimring, L., & Hasty, J. (2017). Rapid and Scalable Preparation of Bacterial Lysates for Cell-Free Gene Expression. ACS Synthetic Biology, 6(12), 2198-2208. https://doi.org/10.1021/acssynbio.7b00253
Ding, Y., Wu, F., & Tan, C. (2014). Synthetic Biology: A Bridge between Artificial and Natural Cells. Life, 4(4), 1092-1116. https://doi.org/10.3390/life4041092
Galloway, W. R. J. D., Hodgkinson, J. T., Bowden, S. D., Welch, M., & Spring, D. R. (2010). Quorum Sensing in Gram-Negative Bacteria: Small-Molecule Modulation of AHL and AI-2 Quorum Sensing Pathways. Chemical Reviews, 111(1), 28-67. https://doi.org/10.1021/cr100109t
Kumari, A., Pasini, P., Deo, S. K., Flomenhoft, D., Shashidhar, H., & Daunert, S. (2006). Biosensing Systems for the Detection of Bacterial Quorum Signaling Molecules. Analytical Chemistry, 78(22), 7603-7609. https://doi.org/10.1021/ac061421n
Lentini, R., Santero, S., Chizzolini, F. et al. Integrating artificial with natural cells to translate chemical messages that direct E. coli behaviour. Nat Commun 5, 4012 (2014). https://doi.org/10.1038/ncomms5012
Miller, C., & Gilmore, J. (2020). Detection of Quorum-Sensing Molecules for Pathogenic Molecules Using Cell-Based and Cell-Free Biosensors. Antibiotics, 9(5), 259. https://doi.org/10.3390/antibiotics9050259
Mukwaya, V., Mann, S. & Dou, H. Chemical communication at the synthetic cell/living cell interface. Commun Chem 4, 161 (2021). https://doi.org/10.1038/s42004-021-00597-w
Rampioni, G., D’Angelo, F., Leoni, L., & Stano, P. (2019). Gene-Expressing Liposomes as Synthetic Cells for Molecular Communication Studies. Frontiers in bioengineering and biotechnology, 7, 1. https://doi.org/10.3389/fbioe.2019.00001
Wen, K. Y., Cameron, L., Chappell, J., Jensen, K., Bell, D. J., Kelwick, R., Kopniczky, M., Davies, J. C., Filloux, A., & Freemont, P. S. (2017). A Cell-Free Biosensor for Detecting Quorum Sensing Molecules in P. aeruginosa-Infected Respiratory Samples. ACS Synthetic Biology, 6(12), 2293-2301. https://doi.org/10.1021/acssynbio.7b00219
Peter Nguyen
Lawrynowicz, A., Vuori, S., Palo, E., Winther, M., Lastusaari, M., & Miettunen, K. (2024). Transforming fabrics into UV-sensing wearables: A photochromic hackmanite coating for repeatable detection. Chemical Engineering Journal, 494, 153069. https://doi.org/10.1016/j.cej.2024.153069
Sąsiadek-Andrzejczak, E., & Kozicki, M. (2023). Multi-Color Printed Textiles for Ultraviolet Radiation Measurements, Creative Designing, and Stimuli-Sensitive Garments. Materials, 16(16), 5622. https://doi.org/10.3390/ma16165622
Ally Huang
Flores, P., Luo, J., Mueller, D. W., Muecklich, F., & Zea, L. (2024). Space biofilms - An overview of the morphology of Pseudomonas aeruginosa biofilms grown on silicone and cellulose membranes on board the international space station. Biofilm, 7, 100182. https://doi.org/10.1016/j.bioflm.2024.100182
Jung, J. K., Rasor, B. J., Rybnicky, G. A., Silverman, A. D., Standeven, J., Kuhn, R., Granito, T., Ekas, H. M., Wang, B. M., Karim, A. S., Lucks, J. B., & Jewett, M. C. (2023). At-Home, Cell-Free Synthetic Biology Education Modules for Transcriptional Regulation and Environmental Water Quality Monitoring. ACS synthetic biology, 12(10), 2909–2921. https://doi.org/10.1021/acssynbio.3c00223
Ravichandran, V., Krishnan, B., Tinwala, M., Kumar, A. S., & Jobby, R. (2025). Microbial resilience in space: Biofilms, risks and strategies for space exploration. Life Sciences In Space Research, 47, 1-13. https://doi.org/10.1016/j.lssr.2025.05.004
Vélez Justiniano, Y. A., Goeres, D. M., Sandvik, E. L., Kjellerup, B. V., Sysoeva, T. A., Harris, J. S., Warnat, S., McGlennen, M., Foreman, C. M., Yang, J., Li, W., Cassilly, C. D., Lott, K., & HerrNeckar, L. E. (2023). Mitigation and use of biofilms in space for the benefit of human space exploration. Biofilm, 5, 100102. https://doi.org/10.1016/j.bioflm.2022.100102
SOURCES:
Image PART A:
Hong, S. H., Kwon, Y. C., & Jewett, M. C. (2014). Non-standard amino acid incorporation into proteins using Escherichia coli cell-free protein synthesis. Frontiers in chemistry, 2, 34. https://doi.org/10.3389/fchem.2014.00034
Figure 5 link: https://www.nasa.gov/wp-content/uploads/2021/10/microbial_research_2021_tagged.pdf
Figure 6 link: https://ntrs.nasa.gov/api/citations/20220008788/downloads/Redefining Microbiological Risk Mitigation during Spaceflight.pdf


