Week 3 HW: Lab Automation
Assignment 1: Python Script for Opentrons Artwork
Task 1.1: Create a Python file to run on an Opentrons liquid handling robot.

Also there is the link of the Usagi code for reference (Assisted with GeminiAI)
In addition,there is the result inoculated by opentrons (ty to SYNBIO USFQ node!) in black agar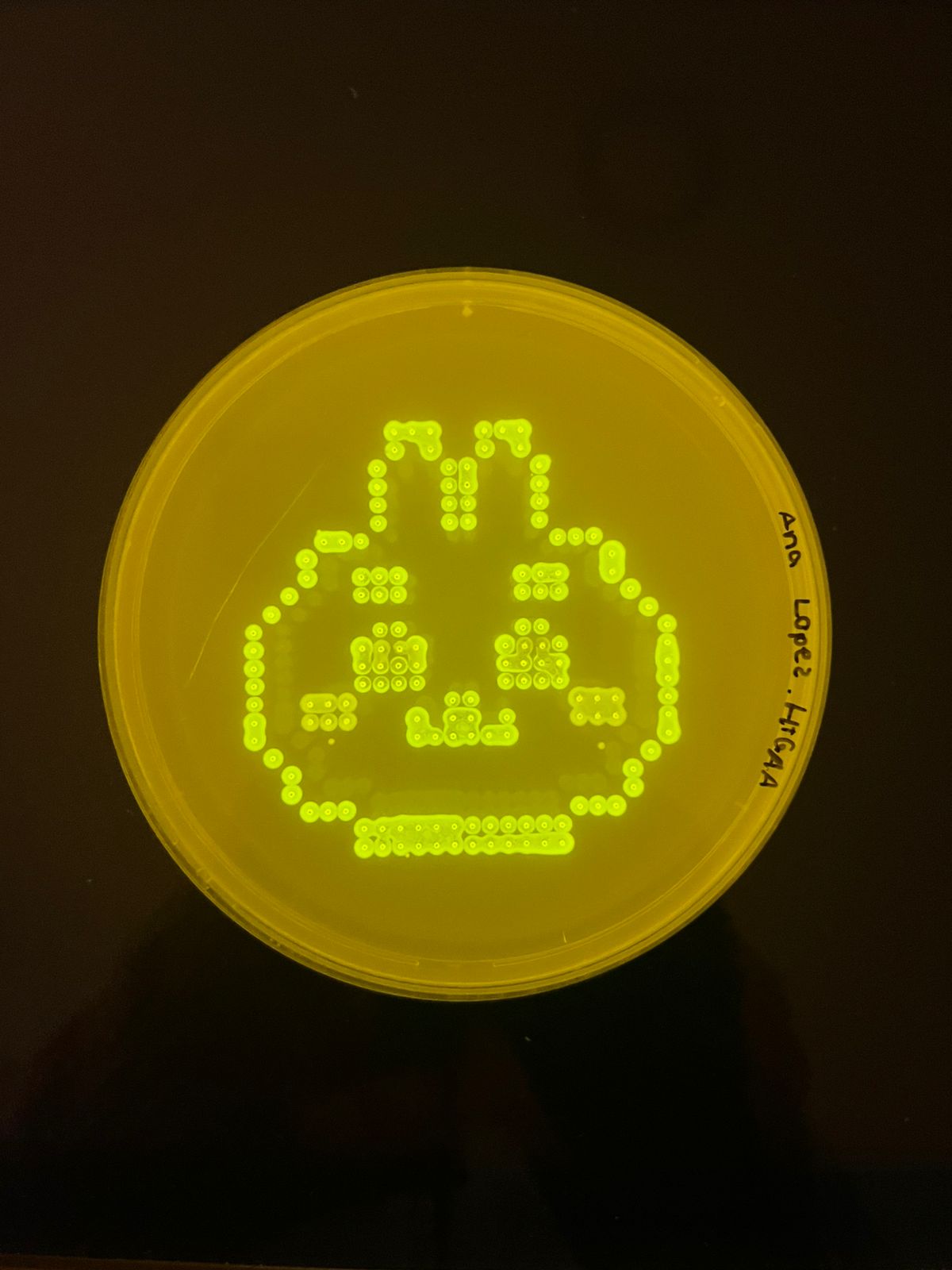 

Assignment 2: Post-Lab Questions
One of the great parts about having an automated robot is being able to precisely mix, deposit, and run reactions without much intervention, and design and deploy experiments remotely.
Task 2.1: Find and describe a published paper that utilizes the Opentrons or an automation tool to achieve novel biological applications.
For this part I choosed the following paper “AssemblyTron: flexible automation of DNA assembly with Opentrons OT-2 lab robots”


(Assisted with DeepseekAI) This paper addresses a common problem such as manual and DNA assembly-related human errors. The authors developed AssemblyTron, an open-source Python package that acts as a bridge between DNA design software and the physical automation of the Opentrons OT-2 liquid handling robot. The biological applications are core molecular cloning tasks:
- Automated PCR setup with an optimized annealing temperature gradient.
- Golden Gate assembly of multi-part DNA constructs (e.g., assembling four fragments to create chromoprotein expression plasmids).
- Homology-dependent assembly (like IVA and AQUA cloning) for site-directed mutagenesis and plasmid construction.
AssemblyTron automates the entire cloning workflow. Its utility was demonstrated by successfully assembling four-part chromoprotein plasmids and performing site-directed mutagenesis with efficiencies comparable to manual methods. The key achievement is creating an accessible, low-cost automation solution that reduces human error, training time, and the cost barrier for academic labs to engage in high-throughput synthetic biology.
Task 2.2: Write a description about what you intend to do with automation tools for your final project. You may include example pseudocode, Python scripts, 3D printed holders, a plan for how to use Ginkgo Nebula, and more. You may reference this week’s recitation slide deck for lab automation details.
The aim for this part is to automate the key steps required to extract metabolites from three distinct microbial consortia (complex samples). The workflow includes:
- Serial dilution and plating to isolate individual bacterial strains from each consortium.
- Preparation of agar plates for culturing.
- Colony picking and inoculation into liquid media.
- Preparation of samples for DNA extraction (for identification).
- Metabolite extraction from liquid cultures. (I have few informations for this step but the manage of samples requieres automatization).
Automation will increase throughput, reduce manual errors, and allow remote execution of experiments. The project will leverage an Opentrons OT-2 for liquid handling, custom 3D‑printed adapters to stabilise agar plates during colony manipulation, and the Ginkgo Bioworks Nebula platform for cloud‑based experiment design and remote protocol execution (such as interpretation of biochemical characterization and identification).
Final Project Ideas
I developt three ideas related to the use of microorganisms. I love microbiology and how microbes can be both versatile and resilient. Some species posses interesting features that can be used for human purposes. In this case, the first slide reflects the use of possible bioproducts from wild microbes as a natural alternative of pesticides/chemicals in the treatment/prevention of bacterial MOKO disease in economy-importance crops in Ecuador (musaceae and solanaceae).

