Subsections of Anastasia Savic — HTGAA Spring 2026
Homework
Weekly homework submissions:
Week 1 HW: Principles and Practices
An application of biological engineering I see as a profound project is a in vivo vector for gene therapy delivery, particularly aimed to aid treatment development for Gastrointestinal disease. An application of policy aiming to sustaining non-malfeasance is developing a clear interpretation of legal parameters regarding in vivo vector R&D and applications outside of academia. This requires a multilayer approach within rigorous biosafety containment and ethical research guidelines. Policy should include legal measures taken in the case of violation, containment methodology or guidelines on how to obtain containment methodology in the case of misuse, as well as consistent regulatory observation by local, national, and international governance.
Week 2 HW: DNA Read, Write, and Edit
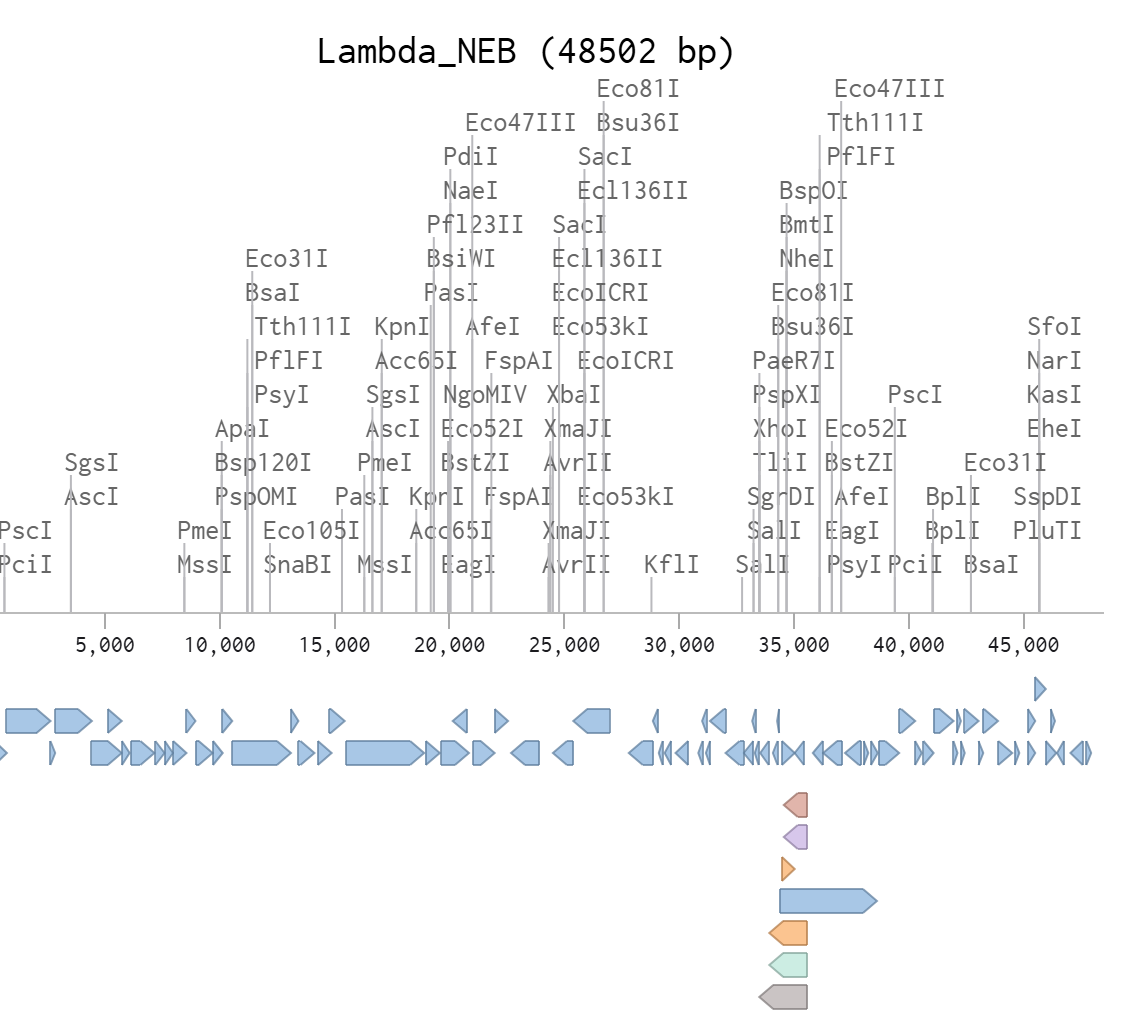
Part 1: Benchling & In-silico Gel Art Objective - For this assignment, I have imported lambda DNA as a .gb file into Benchling and performed in-silico restriction digests with the seven enzymes. I recorded the fragment sizes and generated simulated gels, then designed a multi-lane gel that forms an image.
Subsections of Homework
Week 1 HW: Principles and Practices
An application of biological engineering I see as a profound project is a in vivo vector for gene therapy delivery, particularly aimed to aid treatment development for Gastrointestinal disease.
An application of policy aiming to sustaining non-malfeasance is developing a clear interpretation of legal parameters regarding in vivo vector R&D and applications outside of academia. This requires a multilayer approach within rigorous biosafety containment and ethical research guidelines. Policy should include legal measures taken in the case of violation, containment methodology or guidelines on how to obtain containment methodology in the case of misuse, as well as consistent regulatory observation by local, national, and international governance.
Currently vectors are under constant shift due to balancing the rapid innovation and safety risks associated to the research. Major regulators actively are updating guidelines in the efforts to accommodate advanced viral vectors. In these new proposals, governance should be further in depth with collaboration in early stage development. Incentive to collaborate with governing bodies should include gauranteed and contract research funding through the governing body. Labwork, required documentation, and final products will be continuously shared by the primary research group (companies or acadmemics) and it’s governing collaborators(federal regulators). To preserve intellectual property rights of the primary research group, governing bodies will divide teams within the federal regulators amongst the the competitors. Actors such as NIH staff who are working in collaboration with the primary research teams, the Board of the CDC and others will implement reflective policies of the International Council for Harmonisation (ICH) and the Cell and Gene Therapy Discussion Group (CGTDG). This reflective policies will umbrella all projects and will adjust current foundational guidelines to best optimize innovation, but the brunt of the regulation and documentation for safety and possible containment methodology will be based on the project details themselves. Each federal regulation team will include individuals representing the FDA, NIH, CDC, and others. Policy for each project will be determined on the vector’s original pathogenesis and virulence and organic changes or additions that make cause altering to those properties. With all proposals implemented, in the case of an accident, the regulatory teams will have first hand knowledge and documentation of the project, its content and the organic systems that started it. Failures of this proposal may include possible leaks of information or digital infiltration via cybersecurity concerns. Though the technology is fairly knew and I am limited in my knowledge base, this policy proposal could be oversimplified or indirectly unapplicable in certain areas of study due to the complex nature of the research and the vectors themselves.
| Does the option: | Option 1 | Option 2 | Option 3 |
|---|---|---|---|
| Enhance Biosecurity | 2 | 2 | 1 |
| • By preventing incidents | 3 | 2 | 1 |
| • By helping respond | 1 | 2 | 1 |
| Foster Lab Safety | 2 | 2 | 1 |
| • By preventing incident | 1 | 1 | 1 |
| • By helping respond | 3 | 3 | 1 |
| Protect the environment | 1.5 | 1 | 2 |
| • By preventing incidents | 2 | 1 | 1 |
| • By helping respond | 1 | 1 | 3 |
| Other considerations | 2 | 2 | 2 |
| • Minimizing costs and burdens to stakeholders | 3 | 3 | 3 |
| • Feasibility? | 3 | 3 | 3 |
| • Not impede research | 2 | 2 | 2 |
| • Promote constructive applications | 1 | 1 | 1 |
I think a combination between all the proposals is possible and feasible. I would prioritize minimizing the constant changing plane that we currently have in regulatory markets and use more of an umbrella approach with interaction between all the regulators and research groups individually or in collaboration. This flexibility will open doors for new approaches and prioritize the innovation itself. Federal regulations should be broken down upon the project’s guidelines, on that note, local governance, univiersity policies, and branches of federal agencies should be part of those teams. Prior to the research beginning, guidelines should be developed as a matrix surrounding this particular project or thesis.
Polymerase error rate is about 1 error per 10^6 bases. Human genome length is about 3.2 Billion base pairs. With that error rate, per genome copy, there would be 3,200 errors. Proofreading during replication reduces the mistakes, 3’ to 5’ by exonuclease. Mismatch repair after the replication catches more mistakes. Damage repair systems fix the lesions that would become mutations. If all else fails, checkpoints and apoptosis become key players in mitigating the risks.
Human protein coding length = 1036 base pairs. That is roughly 345 amino acids if we ignore stop codons. There is approximately 3 codons per amino acid. 61 sense codons for 20 amino acids. So 3.05345, about 10167 different DNA sequences. Common failures of protein synthesis can be codon bias, tRNA amounts present, high GC, and hidden or accidental splicing sites.
Week 2 HW: DNA Read, Write, and Edit
Part 1: Benchling & In-silico Gel Art Objective - For this assignment, I have imported lambda DNA as a .gb file into Benchling and performed in-silico restriction digests with the seven enzymes. I recorded the fragment sizes and generated simulated gels, then designed a multi-lane gel that forms an image.