Week 3 – Lab Automation
✨ Week 3 - homework ✨
Here is the link to my Automation Art (2026 HTGAA Bacteriophage): https://opentrons-art.rcdonovan.com/?id=jy86j81azdyuadc
I manually entered the coordinates from the link above, step by step, to reconstruct the design inside the protocol. After completing the process, I obtained the following image.
✨ Post-Lab Questions ✨
1. Find and describe a published paper that utilizes the Opentrons or an automation tool to achieve novel biological applications.
Article Title: AssemblyTron: Flexible automation of DNA assembly with Opentrons OT‑2 lab robots Authors: John A. Bryant Jr., Mason Kellinger, Cameron Longmire, Ryan Miller, R. Clay Wright Year: 2022 DOI: https://doi.org/10.1101/2022.09.29.510219
Article 1In the article “AssemblyTron: Flexible automation of DNA assembly with Opentrons OT‑2 lab robots,” the authors aim to demonstrate the value of using automation tools in scientific research. Their main objective is to show how the Opentrons OT‑2 platform can simplify laboratory workflows, reduce human error, and significantly shorten reaction setup time. The study uses plasmid DNA, including E. coli plasmids carrying chromoprotein genes and plasmids encoding plant transcription factors. By working with these constructs, the authors illustrate several capabilities of the OT‑2 robot. For example, the system automatically prepares PCR reactions with optimized annealing temperatures, performs Golden Gate Assembly to build multi‑fragment plasmids with high accuracy, and executes homology‑based assembly methods such as AQUA and IVA for cloning and site‑directed mutagenesis. Overall, the article highlights how integrating Opentrons automation into the Design‑Build‑Test‑Learn cycle can make molecular biology more efficient, reliable, and accessible to researchers. | Article 2However, the second article, “Real‑time AI‑driven quality control for laboratory automation,” demonstrates that even though automation brings many advantages, systems like the Opentrons OT‑2 are not completely error‑free. The authors highlight that issues such as missing pipette tips, incorrect liquid volumes, or failed aspiration steps can still occur during automated workflows. To address these limitations, the study introduces an AI‑based computer‑vision system capable of detecting such errors in real time. By integrating a YOLOv8 deep‑learning model with a camera mounted on the OT‑2, the system continuously monitors pipetting actions and alerts the user when something goes wrong. This shows that while automation improves efficiency and reproducibility, additional quality‑control tools are essential to ensure reliability, especially in sensitive biological experiments. |
2. Write a description about what you intend to do with automation tools for your final project.
For my final project, I want to explore how automation tools can support a workflow focused on improving plant-based strategies for reducing heavy metal contamination in soil. I’m also interested in how genetic modifications could help plants grow with fewer pesticides. To make the experimental steps more reliable and easier to repeat, I plan to automate several parts of the Design–Build–Test cycle.
I would use the Opentrons OT‑2 to automate tasks such as preparing PCR reactions, assembling genetic constructs, and setting up transformation mixes. Automating these steps would reduce pipetting errors and make it easier to test multiple gene variants in parallel. I may also design a 3D‑printed holder to keep plant DNA extraction tubes stable on the OT‑2 deck, since plant samples often come in irregular formats.
Here is an example of how part of the workflow could look in pseudocode:
load_labware(“PCR_plate”) load_reagents([“master_mix”, “template_DNA”, “primer_sets”])
for variant in gene_variants: pipette.transfer(master_mix, PCR_well[variant]) pipette.transfer(template_DNA, PCR_well[variant]) pipette.transfer(primer_sets[variant], PCR_well[variant])
run_thermocycler(“PCR_program”)
Later, I could use Ginkgo Nebula to explore or simulate different gene designs related to metal uptake or pest resistance, helping me decide which constructs are worth testing. Overall, automation would make the workflow more efficient and reproducible, allowing me to focus more on analyzing plant performance rather than repeating manual prep steps.
✨ Final Project Ideas ✨
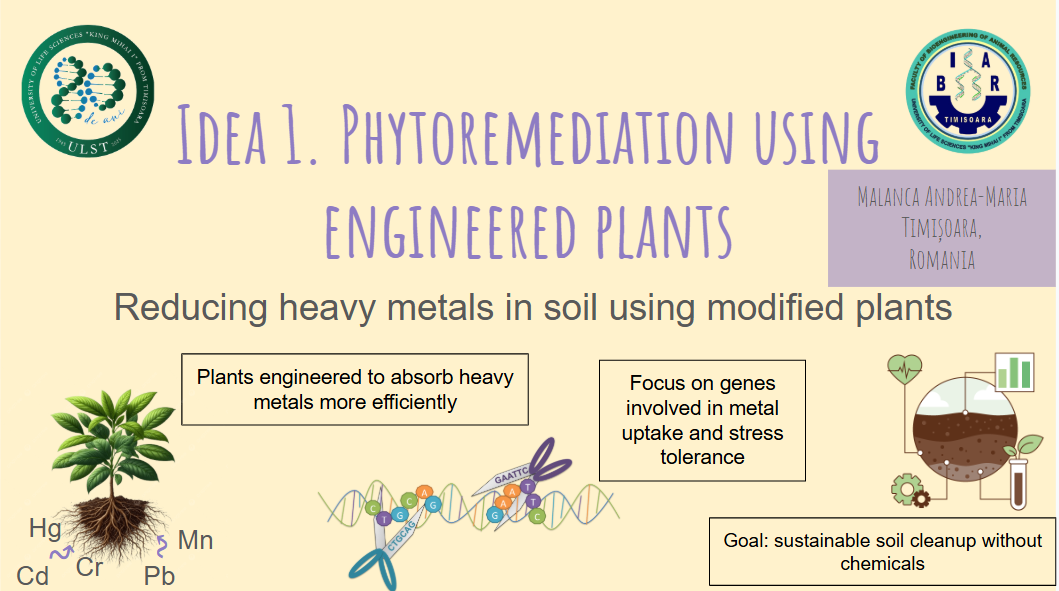


These ideas reflect different ways I could combine synthetic biology, environmental applications, and automation tools for my final project.