Week 6 — Genetic Circuits Part I: Assembly Technologies
✨ DNA Assembly ✨
1. What are some components in the Phusion High-Fidelity PCR Master Mix and what is their purpose?
The Phusion High-Fidelity PCR Master Mix contains Phusion DNA polymerase, dNTPs, reaction buffer, and Mg²⁺ ions. These components allow accurate DNA amplification, with the polymerase synthesizing DNA, dNTPs acting as building blocks, and the buffer and Mg²⁺ providing optimal conditions for the reaction.
2. What are some factors that determine primer annealing temperature during PCR?
Primer annealing temperature during PCR is mainly determined by the melting temperature (Tm) of the primers, their length and GC content, and the specificity of primer binding to the template DNA. These factors influence how strongly the primers bind to the target sequence.
3. There are two methods from this class that create linear fragments of DNA: PCR, and restriction enzyme digests. Compare and contrast these two methods, both in terms of protocol as well as when one may be preferable to use over the other.
PCR and restriction enzyme digestion can both create linear fragments of DNA, but they differ in their protocol and purpose.
The PCR protocol is more complex because it requires different temperature steps during the process: denaturation, primer annealing, and extension. These steps are repeated many times in a thermocycler to amplify the DNA fragment between the primers. PCR also requires several components such as primers, DNA polymerase, and nucleotides.
In contrast, restriction enzyme digestion is simpler. In this method, DNA is incubated with restriction enzymes at an optimal temperature so that the enzymes can recognize specific sequences and cut the DNA at those sites. Usually, it only requires maintaining the right conditions for the enzyme to function properly.
The choice between these methods depends on the goal of the experiment. PCR is preferable when we want to amplify a specific DNA fragment from a small amount of DNA. Restriction enzyme digestion is preferable when we want to cut DNA into specific fragments.
4. How can you ensure that the DNA sequences that you have digested and PCR-ed will be appropriate for Gibson cloning?
You can ensure that DNA sequences are appropriate for Gibson cloning by designing primers during PCR that add overlapping sequences at the ends of your fragments. These overlaps are complementary to the adjacent fragment or vector, so after PCR, the fragments will already have ends compatible for Gibson assembly.
5. How does the plasmid DNA enter the E. coli cells during transformation?
The plasmid DNA cannot enter E. coli cells on its own because the membrane is a barrier. The cells must be made competent, which can be done in two ways:
- Chemical competence: Treating cells with salts such as CaCl₂, which makes the membrane more permeable, followed by a heat shock to allow the plasmid to enter.
- Electroporation: Applying a short electrical pulse to create temporary pores in the membrane, through which the plasmid can pass.
6. Describe another assembly method in detail (such as Golden Gate Assembly)
a. Explain the other method in 5 - 7 sentences plus diagrams (either handmade or online).
Golden Gate Assembly (GGA) is a molecular cloning method that allows multiple DNA fragments to be assembled in a defined order in a single reaction. Each fragment must have overhangs that are complementary to the vector or to the adjacent fragment, so they can fit together like puzzle pieces. These overhangs are created by Type IIS restriction enzymes, which cut outside of their recognition site, generating specific single-stranded ends.
Unlike Gibson Assembly, GGA does not require PCR to create overlaps if fragments already contain the enzyme sites. After the overhangs are formed, DNA ligase joins the fragments together, sealing the nicks and producing a continuous DNA molecule.
This method is very efficient for assembling multiple fragments at once, and it leaves no extra sequences (scarless) between fragments. It is particularly useful when precise order and orientation of fragments is required.
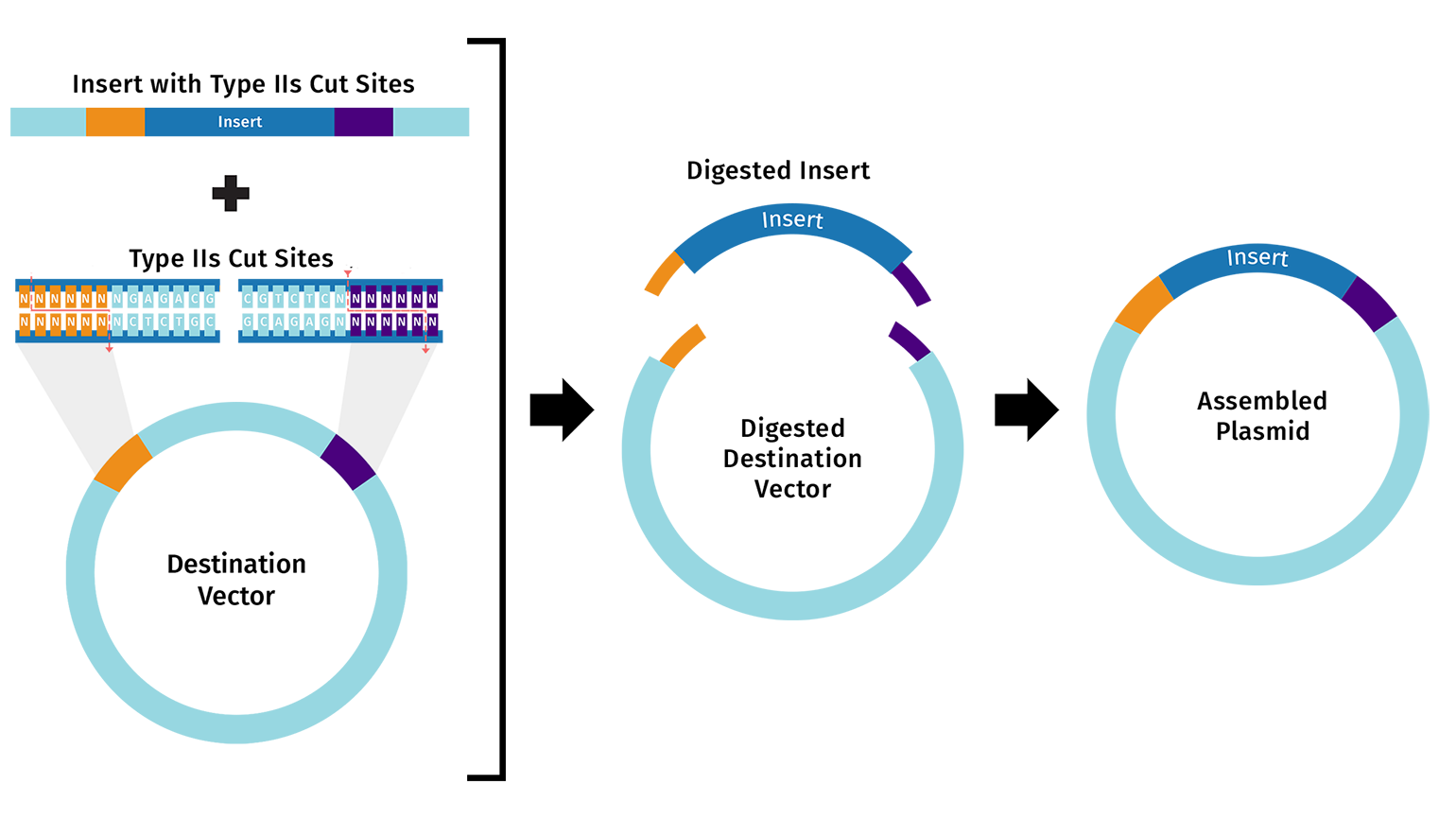
b. Model this assembly method with Benchling or Asimov Kernel!
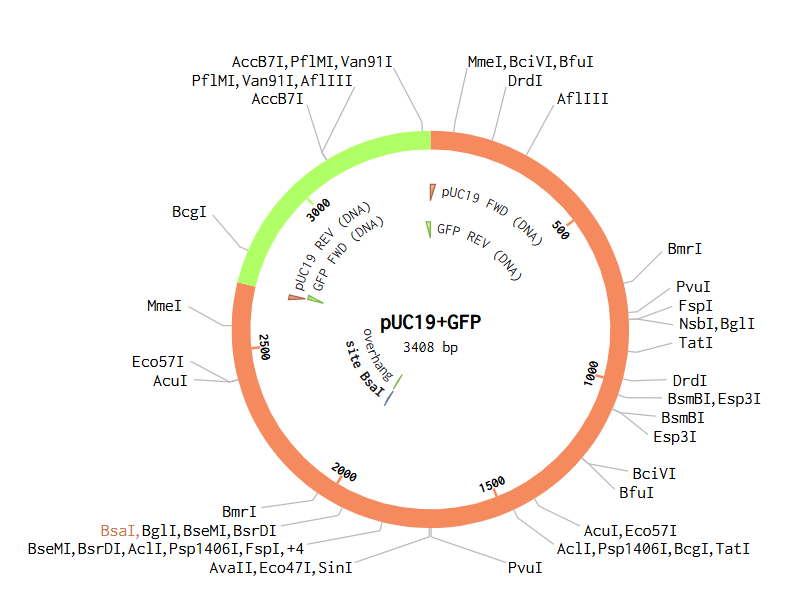
In Benchling, I created a new DNA sequence (type: DNA, topology: circular) to represent the pUC19 plasmid. I searched for its sequence and found it to be 2686 bp. I then added another DNA fragment that I wanted to insert into pUC19, which is GFP. From UniProt, I obtained the GFP sequence of 238 amino acids and used an online tool to perform reverse translation from amino acids to base pairs.
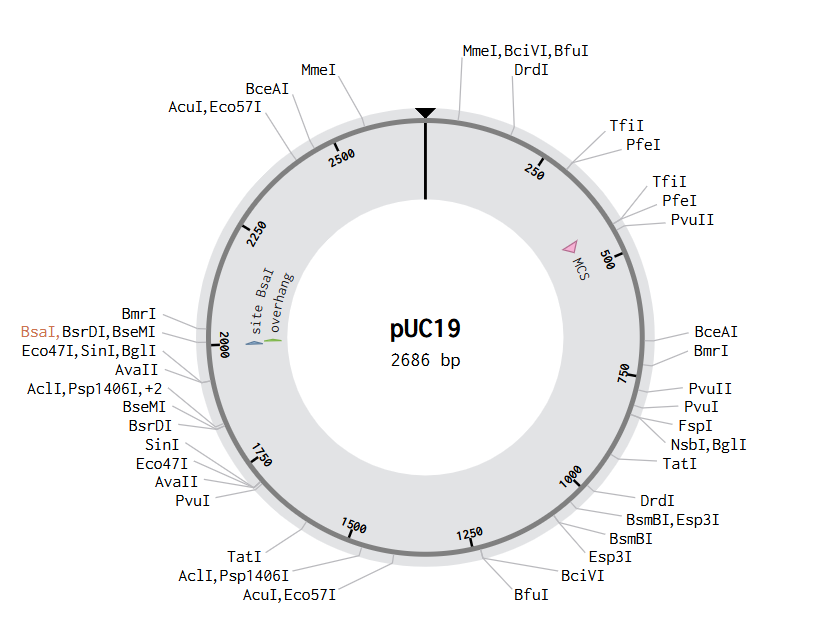
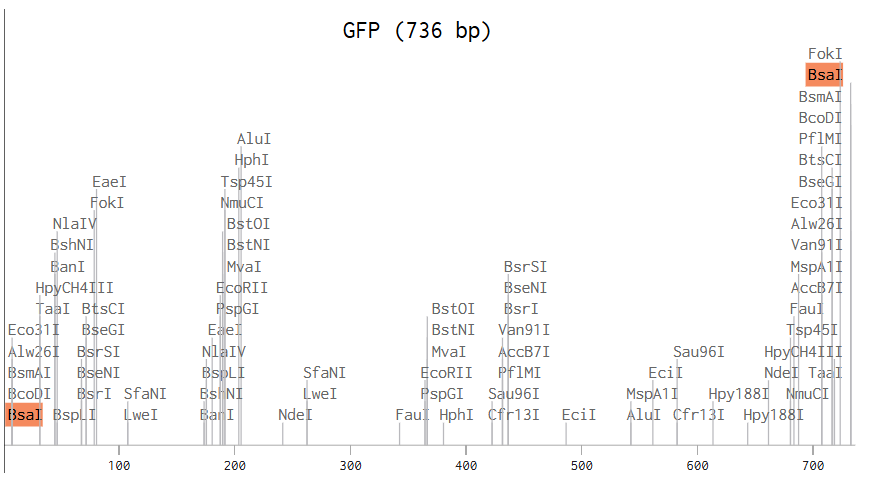
In the pUC19 sequence, I ran a digest using the Type IIS enzyme BsaI, and I found that it cuts at position 2006. I added annotations for the cut site: 1999–2004 for the BsaI recognition site, and 2006–2009 for the 4 bp overhang CGGT.
To link the GFP fragment, I added the complementary overhangs (GCCA) at both the 5’ and 3’ ends of the fragment. Then, using the Assembly Wizard, I selected the method I wanted (Golden Gate Assembly), assigned the backbone as pUC19 and the insert as GFP, and finally obtained the assembled plasmid with GFP correctly inserted.