Subsections of Homeworks
Week 1 HW: Principles and Practices
Biological Engineering Application
1. Description:
My idea is to develop a biosensor for heavy metal detection using engineered Escherichia coli bacteria as a chassis. Lead, cadmium, and mercury are the most problematic heavy metals, accumulating in organisms and slowly killing them. The tool would involve genetically modifying E. coli to incorporate metal-responsive genetic circuits that detect the presence of heavy metals and respond by producing an easily observable response. Right now I am considering green fluorescent protein (GFP) to be the best fit. When exposed to contaminated water, the bacteria would “light up” proportionally to the metal concentration, allowing for quick detection via a simple handheld fluorometer. Heavy metal pollution from industrial runoff, mining, and urban waste is a major global issue, leading to contaminated drinking water that causes neurological damage, kidney failure, and ecosystem disruption in affected communities. Traditional detection methods, like atomic absorption spectrometry, are expensive, lab-based, and time-consuming, often delaying response to pollution events. In contrast, biosensors are low-cost, portable, and provide real-time results, enabling faster interventions in polluted areas, such as rivers or municipal water supplies in developing regions. Iit promotes sustainable monitoring and could integrate with community science initiatives to empower local environmental stewardship.
2. Governance/Policy Goals
To ensure this biosensor contributes to an ethical future, the overarching goal is to promote responsible innovation that prioritizes non-malfeasance—preventing harm to human health, ecosystems, and society—while fostering beneficial uses. This adapts the synthetic genomics framework mentioned, emphasizing safety and security alongside equity to address potential disparities in access and impact. I break this into three specific sub-goals: Biosafety Sub-Goal: Minimize environmental and health risks from the release or unintended spread of engineered organisms. This includes preventing ecological disruption, such as engineered bacteria outcompeting native microbes or transferring genes, which could alter biodiversity in water systems. Policies should require containment strategies to ensure the biosensor is used in controlled settings without persistent environmental persistence. Biosecurity Sub-Goal: Prevent misuse or dual-use applications that could enable harm, like weaponizing metal-detection circuits for bioterrorism or creating biosensors that inadvertently aid in evading pollution regulations. This promotes secure handling of genetic designs to avoid exploitation by malicious actors, drawing on broader SynBio governance to safeguard public trust. Equity Sub-Goal: Ensure fair access and distribution of the technology, particularly for underserved communities facing water pollution. This involves policies that avoid exacerbating global inequalities, such as patent barriers limiting deployment in low-income countries, and instead encourage open-source designs or subsidies to promote inclusive benefits and autonomy in environmental monitoring.
3. Potential Governance Actions
Action 1: Mandatory Biosafety Certification for Biosensor Deployment (Regulatory Rule by Federal Regulators like the EPA or FDA)
Purpose: Currently, environmental biosensors fall under general biotech regulations like the Coordinated Framework for Biotechnology, but lack specific mandates for field-use certification. The proposed change requires pre-approval testing for ecological safety before commercial or research deployment, similar to EPA pesticide registrations, to prevent accidental harm. Design: Regulators would develop a certification process involving lab and field trials assessing containment, gene stability, and non-toxicity. Actors include federal agencies (implementers), companies/academics (applicants who must fund tests), and expert panels (approvers). Funding could come from application fees, with international harmonization via bodies like the OECD. Assumptions: Assumes regulators have sufficient expertise in SynBio and that testing protocols accurately predict real-world impacts; uncertainties include long-term ecological effects in diverse water environments. Risks of Failure & “Success”: Failure could occur if bureaucracy delays innovation, stifling biosensor adoption in pollution hotspots. “Success” might unintendedly create over-reliance on certified tools, marginalizing non-certified community innovations, or lead to complacency in broader pollution prevention.
Action 2: Grant Incentives for Equity-Focused Biosensor Development
Purpose: Today, funding for SynBio often prioritizes commercial viability over equity, leading to tech concentrated in wealthy nations. The proposal introduces grants rewarding projects with open-access designs in polluted developing regions, analogous to financial incentives for green tech under the Paris Agreement. Design: Organizations would allocate funds via competitive calls, requiring proposals to include equity metrics like tech transfer to low-income areas. Actors: Funders (providers), researchers/companies (opt-in applicants), and NGOs (implementers for distribution). Approval involves peer review emphasizing ethical impact. Assumptions: Assumes grantees will genuinely prioritize equity without gaming the system; uncertainties around measuring “equitable” outcomes in diverse cultural contexts. Risks of Failure & “Success”: Failure if funds are insufficient or misallocated, perpetuating inequities. “Success” could flood markets with subsidized tools, undercutting local innovations or creating dependency on international aid, potentially ignoring region-specific pollution needs.
Action 3: Integration of Genetic Kill Switches in Biosensor Designs
Purpose: Current SynBio designs sometimes lack built-in safeguards, risking uncontrolled spread. The change mandates researchers to embed “kill switches”—genetic circuits that deactivate the organism after use or under certain conditions (e.g., absence of a lab-provided nutrient)—inspired by 3D printing’s software locks for safety. Design: Researchers would incorporate switches during engineering, sharing protocols via open repositories like Addgene. Actors: Academics (implementers who opt-in for ethical best practices), funders (requiring it for grants), and journals (enforcing via publication guidelines). No major new funding needed, but training workshops could accelerate adoption. Assumptions: Assumes kill switches are foolproof and won’t evolve away; uncertainties include efficacy in variable field conditions like polluted water pH changes. Risks of Failure & “Success”: Failure if switches malfunction, allowing escape and ecological harm. “Success” might give a false sense of security, encouraging riskier deployments, or raise biosecurity concerns if switches are hacked for malicious persistence.
4. Score
| Criterion | Sub-Criteria | Action 1: Mandatory Biosafety Certification | Action 2: Grant Incentives for Equity-Focused | Action 3: Integration of Genetic Kill Switches |
|---|---|---|---|---|
| Enhance Biosecurity | By preventing incidents | 1 | 2 | 1 |
| By helping respond | 1 | 2 | 2 | |
| Foster Biosafety | By preventing incidents | 1 | 2 | 1 |
| By helping respond | 2 | 2 | 1 | |
| Promote Equity | By ensuring fair access | 2 | 3 | |
| By preventing disparities | 1 | 2 | 2 | |
| Protect the Environment | By preventing incidents | 1 | 2 | 1 |
| By helping respond | 1 | 2 | 1 | |
| Other Considerations | Minimizing costs and burdens | 2 | 2 | 3 |
| Feasibility | 2 | 2 | 2 | |
| Not impede research | 3 | 2 | 3 | |
| Promote constructive applications | 1 | 2 | 2 |
5. The most important governance option
Based on the scoring, I would prioritize a combination of Action 2 and Action 3 (Integration of Genetic Kill Switches), with Action 1 as a supportive but secondary measure. This hybrid approach balances prevention, response, and promotion of beneficial uses without excessive burdens. Action 2 excels in equity and constructive applications, addressing the critical need to make biosensors accessible in pollution-affected developing regions, while fostering innovation through incentives rather than mandates. Action 3 complements this by providing strong biosafety and biosecurity, offering technical safeguard that integrates seamlessly into development workflows. Together, they minimize risks like environmental harm. Action 1, while effective for prevention, scores poorly on burdens, feasibility for small actors, and not impeding research, making it better as an optional escalation for high-risk commercial applications rather than a blanket requirement. Trade-offs considered: This combo trades some regulatory rigor (Action 1’s strength) for flexibility and lower costs, potentially allowing minor incidents if voluntary adoption lags, but it avoids stifling research in a field like synthetic biology where rapid iteration is key. It also prioritizes equity over uniform enforcement, which might mean uneven global standards but better real-world impact in vulnerable areas.
Homework Questions from Professor Jacobson:
- Nature’s machinery for copying DNA is called polymerase. What is the error rate of polymerase? How does this compare to the length of the human genome. How does biology deal with that discrepancy?
- Polymerase error rate: 1:10^6
- The human genome has 4.2 billion base pairs, which means that polymerase makes 3200 errors at each celll division.
- In order to deal with this high error rate, cells have systems made of multiple proteins that correct those errors, although they are not perfect.
- How many different ways are there to code (DNA nucleotide code) for an average human protein? In practice what are some of the reasons that all of these different codes don’t work to code for the protein of interest?
- Human protein: ~1036bp –> ~345 amino acids W
- Different codons can enconde for the same AA, but not all these sequences are equally efficcinet. Depending on the organism, a codon might be better for correctly and fast/slow enough being translated into an amino acid.
Homework Questions from Dr. LeProust:
- What’s the most commonly used method for oligo synthesis currently?
- The solid-phase phosphoramidite chemical synthesis
- Why is it difficult to make oligos longer than 200nt via direct synthesis?
- error accumulation and chemical limitations
- Why can’t you make a 2000bp gene via direct oligo synthesis?
- mailny due to error accumulation
Homework Question from George Church:
What are the 10 essential amino acids in all animals and how does this affect your view of the “Lysine Contingency”?
The 10 essential amino acids that animals cannot synthesize on their own and must obtain through their diet are: Arginine, Histidine, Isoleucine, Leucine, Lysine, Methionine, Phenylalanine, Threonine, Tryptophan, Valine
The “Lysine Contingency” refers to the fictional failsafe mechanism in Michael Crichton’s Jurassic Park, where resurrected dinosaurs were genetically engineered to lack the ability to produce lysine, one of these essential amino acids. The idea was that without lab-supplied lysine supplements, the dinosaurs would die, preventing them from surviving if they escaped containment. Lysine is an essential amino acid for all animals which reinforces why this contingency is fundamentally flawed as a control measure. In reality, no animal can synthesize lysine endogenously—they all rely on dietary sources like plants, insects, or meat, which are abundant in natural ecosystems. If the engineered dinosaurs escaped, they could simply consume lysine-rich foods in the wild (as they do in the story), rendering the dependency moot. It highlights a clever narrative device but poor fictional science, as it overlooks basic nutritional biology and ecology. A more effective contingency might target a non-essential amino acid or something truly unique to the lab environment.
Week 2 Homework: DNA Read, Write & Edit
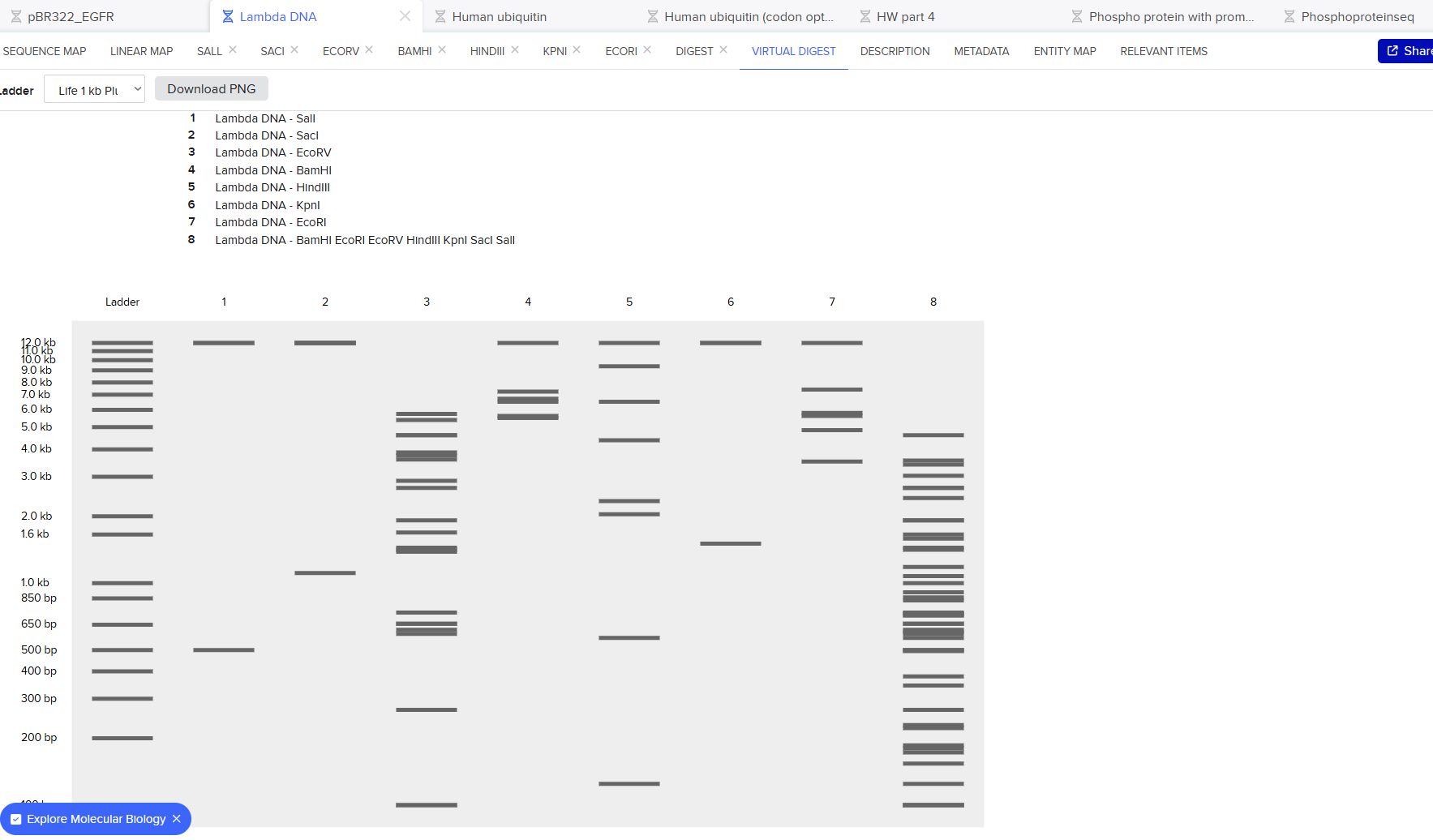
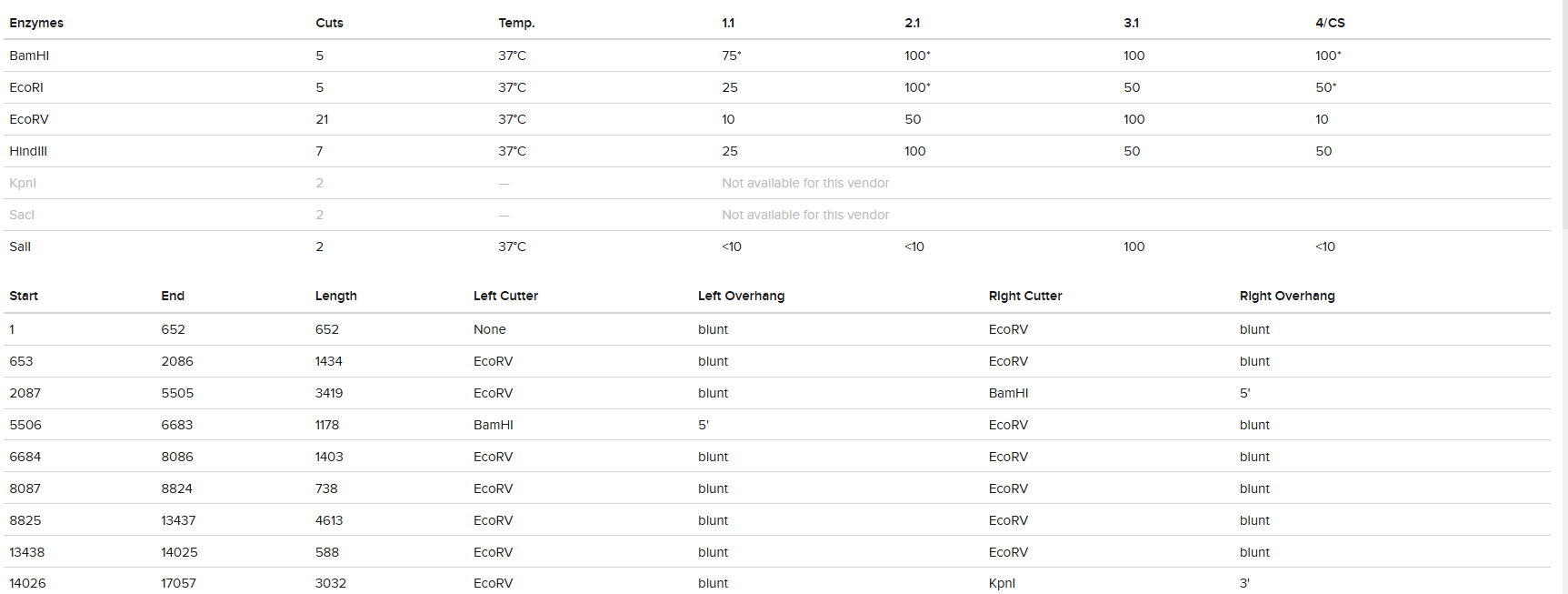
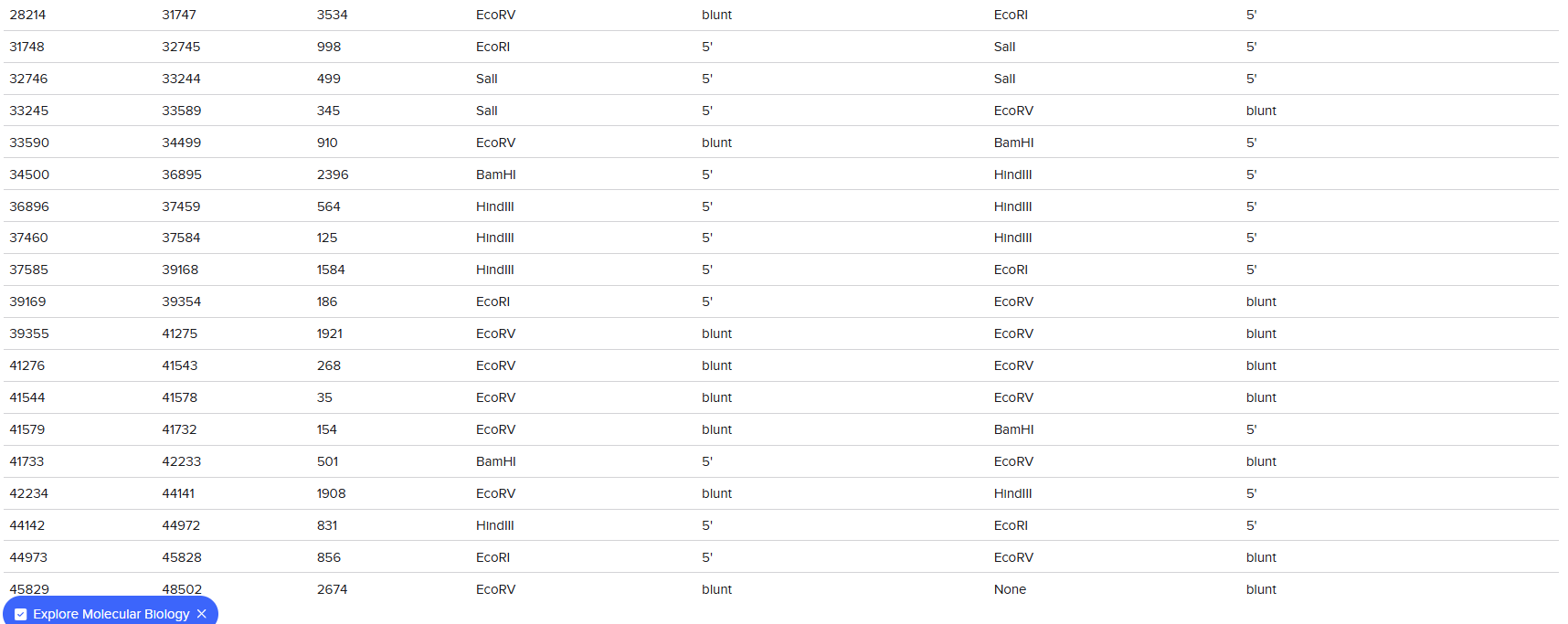
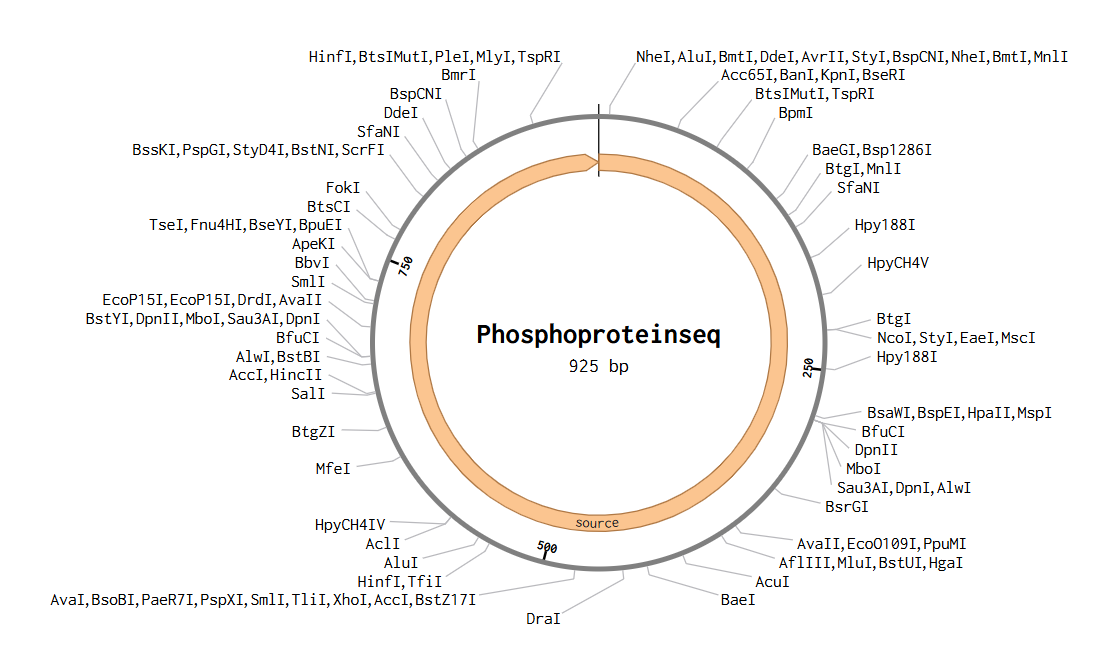
1. Gel Art:
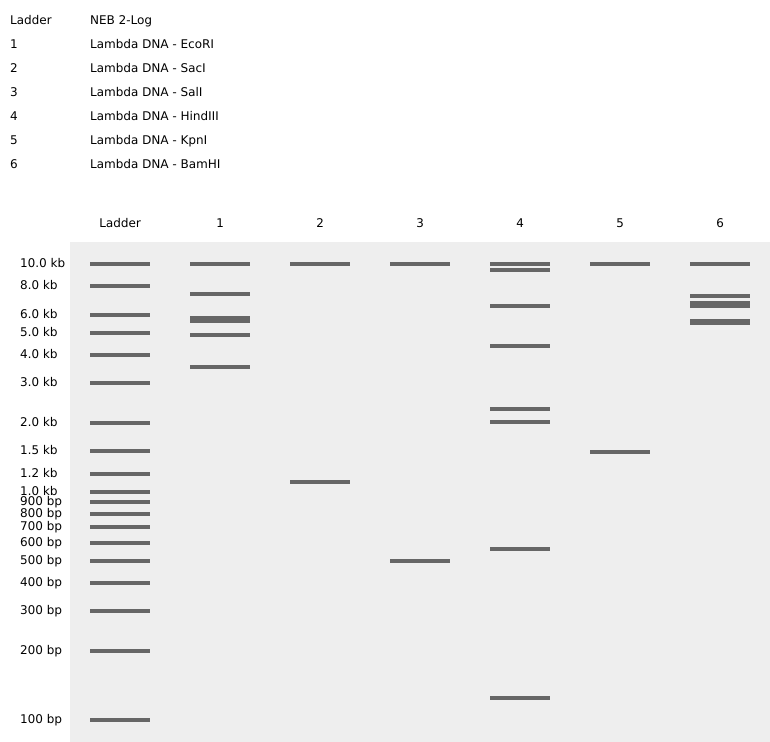


vs


The digestion results:
3.1. Choose your protein.
Human ubiquitin: one of the most highly conserved proteins across eukaryotic species, with an identical amino acid sequence in organisms ranging from yeast to humans. It marks misfolded proteins for degradation. Protein sequence - 76 amino acids sp|P0CG48|UBC_HUMAN Polyubiquitin-C OS=Homo sapiens OX=9606 GN=UBC PE=1 SV=2 MQIFVKTLTGKTITLEVEPSDTIENVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLSDYNIQKESTLHLVLRLRGG
3.2. Reverse Translation:
ATGCAGATCTTCGTGAAGACCCTGACCGGCAAGACCATCACTCTGGAGGTGGAGCCCAGTGACACCATCGAGAACGTGAAGGCCAAGATCCAGGACAAGGAGGGCATCCCTCCTGACCAGCAGAGGCTGATCTTTGCCGGCAAGCAGCTGGAAGATGGCCGCACCCTGTCTGACTACAACATCCAGAAGGAGTCCACCCTGCACCTGGTGCTGCGTCTGCGTGGCGGC
3.3. Codon optimization:
Different organisms have distinct preferences for which codons they use for each amino acid, due to differences in tRNA abundance and translation efficiency. The native human sequence may contain codons that are rare in other hosts, leading to slower translation and ribosomal stalling, which increase the risk of misfolding. I decided to optimize for Escherichia coli K-12, which is fast-growing, inexpensive, well-characterized, and widely used for high-yield protein expression in research and industry. Using Benchling codon optimization tool, I obtained:
ATGCAGATCTTTGTGAAAACCCTGACCGGTAAAACCATTACCCTGGAAGTGGAGCCGAGCGATACCATTGAAAACGTGAAAGCGAAAATTCAGGATAAAGAAGGCATTCCGCCGGATCAGCAGCGCCTGATTTTTGCCGGCAAACAGCTGGAAGATGGTCGTACCCTGAGCGACTATAACATTCAGAAAGAAAGCACCTTACATCTGGTGCTGCGTCTGCGTGGTGGT
3.4. You have a sequence! Now what?
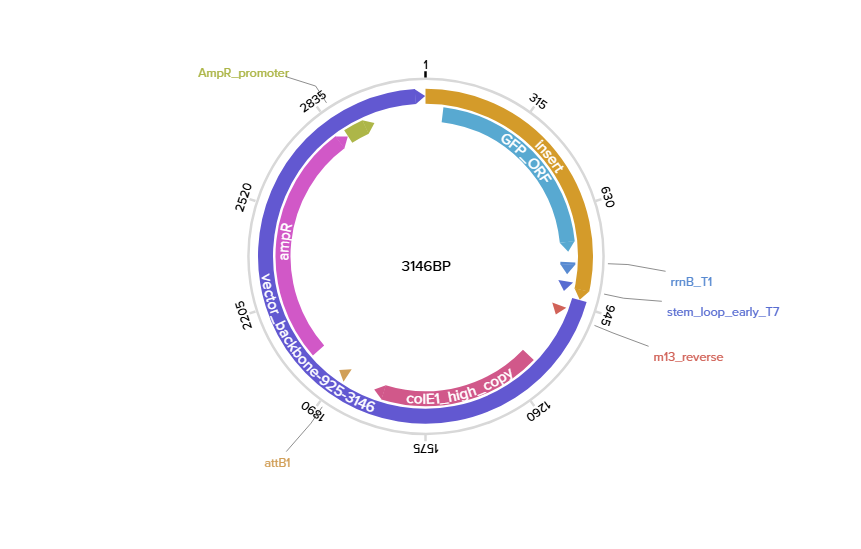
The optimized DNA sequence can be synthesized and cloned into a vector for protein production in E.coli K-12:
The DNA is inserted into a plasmid and transported into E. coli cells. Host RNA polymerase transcribes the DNA into mRNA. Ribosomes then translate the mRNA into the polypeptide chain using tRNAs, amino acids, and energy from GTP/ATP. The protein folds , and cells are lysed to purify it.
3.5. How does it work in nature/biological systems?
How a single gene codes for multiple proteins at the transcriptional level: One common mechanism is splicing, where a single pre-mRNA transcript is processed in different ways to include or exclude exons, producing multiple mature mRNA isoforms from the same gene. These isoforms are translated into distinct proteins with different functions. (Ubiquitin genes themselves primarily produce identical monomers via polyubiquitin precursors or fusions, with cleavage occurring post-translationally; alternative splicing is more prominent in other human genes, e.g., for diversity in receptors or enzymes.) Alignment of DNA, transcribed RNA, and translated Protein:
Important:
- in the genome there are multiple genes that code for ubiquitin (ribosomal fusion genes and polyubiquitin genes), due to the necessity of upregulating the production during periods of increased metabolic stress;
- in general, ubiquitin genes do not contain introns, most likely due to the need of quick production and low change for errors to occur.
- ubiquitin suffers post translation modifications
- ubiquitination is the process by which E1, E2 and E3 enzymes mark misfolded proteins or danaturated ones for protein degradation by attaching the damaged protein to ubiquitin as a substrate
- E1 activates ubiquitin
- E2 transports ubiquitin to the target protein
- E3 facilitates ubiquity-substrate binding
4.2
Complete sequence (promoter, RBS, Start codon, coding sequence, 7xHis Tag, end codon, terminator)
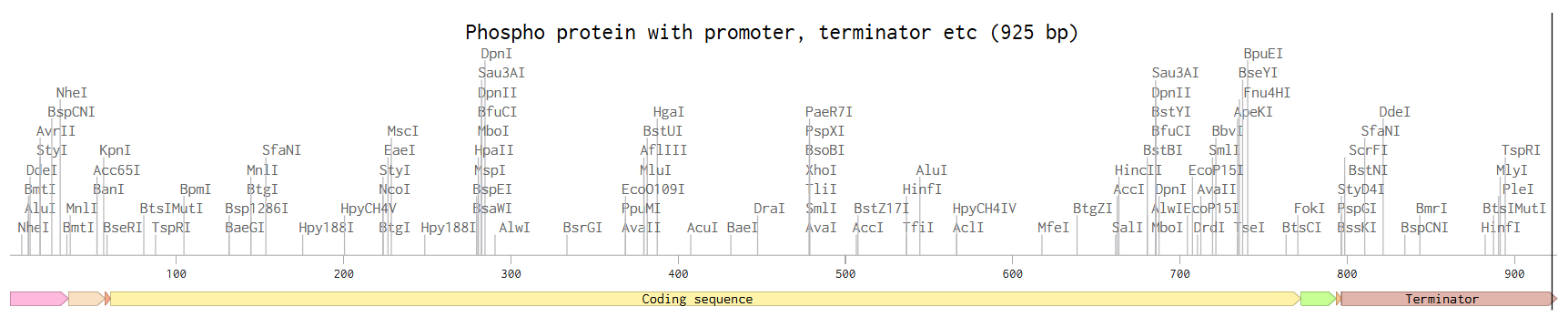


4.3, 4.4, 4,5, 4,6 Results:
5.1 What DNA would you want to sequence (e.g., read) and why? DNA READ
i)I would be enthralled to sequence the genes responsible for ribonucleic vaults. A more profound understanding of how the MVPs (major vault proteins) create literal containers used for cellular transportation would help us better understand how to create intricate structures far more complex than what evolution could achieve in the coming millions of years.
ii) I would use Third-Generation sequencing. Why? Unlike second-generation, which requires plenty of repetitions on amplified clusters, third generation sequencing performs single-molecule, real-time sequencing, hence being a lot more efficient. & It does not require PCR amplification, meaning it bypasses the cycle of amplification that can lead to errors.
5.2 DNA Write:
i) A genetic circuit designed for microplastic detection that pollutes aquatic environments.
The sheer volume of plastic entering oceans has created a vicious cycle where macroplastic breaks down into microplastic that ends up in our food and has toxicological effects on our health. The design would be a genetic circuit that responds to Terephthalic Acid (TPA), a primary product of PET plastic degradation.
- The Sensor: A specific transcription factor (TpaR) that remains inactive until it binds to TPA.
- The Reporter: Once TpaR is activated, it triggers the expression of a protein that can be seen with ease.
ii)
- Synthesis Technology: Silicon-based Synthesis I would choose this technology because it uses a silicon platform to act as a solid support for the chemical reactions. Traditional plastic plates are inefficient; by moving to silicon, we can rebuke the wastefulness of older methods, reducing reagent use by over 99%.
- Scalability: It can synthesize 9,600 genes on a single chip Length Constraints: The longer the DNA strand is, the more the efficiency of adding new bases drops. It would also be preferred to integrate a skill switch for keeping the bacteria population under control.
5.3 DNA Edit:
(i) The Edit: Enabling Nitrogen Fixation in Cereals Currently, only legumes can naturally convert atmospheric nitrogen into a usable form via a secretive symbiotic relationship with specialized bacteria. Most cereal crops lack this ability, forcing farmers into a vicious cycle of applying synthetic nitrogen fertilizers. The planet’s soil and waterways bear the brunt of this practice, leading to massive nutrient runoff and greenhouse gas emissions.
I would want to edit the Symbiosis Receptor-Like Kinase (SYMRK) and Nod Factor Receptor (NFR) genes in wheat. These edits would “re-tune” the plant’s root receptors to recognize the signaling molecules of nitrogen-fixing bacteria.
(ii) The Technology: Prime Editing To achieve these precise changes without causing accidental damage to the rest of the genome, I would use Prime Editing (PE).
While standard CRISPR-Cas9 is effective for cutting genes, Prime Editing is a search-and-replace tool that can rewrite specific DNA letters with manifold precision.
How it Edits DNA (simplified): Targeting: The Prime Editor is a protein consisting of a Cas9 nickase and a Reverse Transcriptase is guided to the specific root receptor gene. Nicking: Instead of cutting both strands of DNA, it creates a small nick in only one strand. Reverse Transcription: The pegRNA (prime editing guide RNA) provides a template. The RT enzyme reads this RNA and synthesizes a new strand of DNA that contains the desired edit. Flap Competition: The newly synthesized DNA flap competes with the original unedited DNA flap.
Incorporation: Through natural cellular repair, the old flap is removed, and the new, edited sequence is permanently integrated into the genome.
Inputs: The Prime Editor Construct: Usually delivered as a plasmid or mRNA encoding the Cas9-RT fusion protein. The pegRNA: The custom RNA sequence that searches and replaces. Delivery System: For plants, we often use Agrobacterium-mediated transformation to shoot these components into the plant tissue. Target Cells: Embryogenic callus cells from the wheat plant, which can be grown back into a whole, fertile plant. Limitations: Efficiency and Precision Although Prime Editing is highly precise, it is not yet perfect: Efficiency: In many plant species, the editing efficiency is still quite low (often <10%). Getting the “search-and-replace” to actually stick across millions of cells remains a bottleneck. Size Constraints: Because the Prime Editor protein is so large, it is difficult to deliver it. Target Range: The editor must be near a specific sequence called a PAM site. If the receptor gene we want to edit isn’t near a PAM, the tool is effectively blind to that location.
Week 3 Homework: Lab automation
Lab Automation Homework Assignment:
“Your task this week is to Create a Python file to run on an Opentrons liquid handling robot. Review this week’s recitation and this week’s lab for details on the Opentrons and programming it. Generate an artistic design using the GUI at opentrons-art.rcdonovan.com. Using the coordinates from the GUI, follow the instructions in the HTGAA26 Opentrons Colab to write your own Python script which draws your design using the Opentrons.”
Colab code:
Generated Design:

Final Colab Barni design:

It is to be noted that I have used Gemini AI thinking mode for the generation of the entire script.
1. Find and describe a published paper that utilizes the Opentrons or an automation tool to achieve novel biological applications.
Paper name: “BOTany Methods: Accessible Automation for Plant Synthetic Biology” by Mingqi Xie and Qian de la Tour (bioRxiv, August 22, 2025) Short description: democratizing access to laboratory automation in plant synthetic biology by developing a suite of modular protocols called BOTany Methods, which leverage the affordable Opentrons OT-2 liquid-handling robot to streamline common molecular biology workflows. The authors highlight how traditional biofoundries are costly and inaccessible, limiting high-throughput experimentation, reproducibility, and the Design-Build-Test-Learn (DBTL) cycle in plant science. By using the OT-2, they enable automation for tasks that are typically manual and error-prone, making them suitable for educational settings and nre researchers.
2. Write a description about what you intend to do with automation tools for your final project. You may include example pseudocode, Python scripts, 3D printed holders, a plan for how to use Ginkgo Nebula, and more. You may reference this week’s recitation slide deck for lab automation details.
For my final project, I plan to automate the development and testing of an aptamer-based electrochemical biosensor integrated into a portable electrical device for automatic detection of heavy metals in water samples. The biosensor will use DNA aptamers as the biorecognition element, immobilized on gold screen-printed electrodes, to bind target metals and produce measurable changes in electrical impedance or current. This will be integrated into a compact, Arduino-based device with a microfluidic channel for sample flow, enabling automatic sensing: water is drawn in via a pump, interacts with the biosensor, and triggers real-time readout via a potentiostat module, with data logged to a cloud app for alerts if thresholds are exceeded. The goal is to create a low-cost, field-deployable system for environmental monitoring, optimizing aptamer sequences and immobilization for high sensitivity (sub-ppb levels). I’ll use the Opentrons OT-2 for local automation to handle high-throughput screening of aptamer variants and electrode functionalization, while leveraging Ginkgo Nebula’s cloud lab for large-scale DNA synthesis and initial validation. Specifically:
Aptamer design and synthesis: Design aptamer libraries targeting heavy metals using tools like SELEX simulation software, then submit to Ginkgo Nebula for automated synthesis and amplification. Nebula’s high-throughput platform can produce 96-384 variants in plates, shipped for local use. Local automation with Opentrons: The OT-2 will prepare functionalization solutions (e.g., thiol-modified aptamers mixed with linkers), deposit them precisely onto electrode arrays in a 96-well format for immobilization, and perform washing/incubation steps. It will also simulate testing by dispensing metal-spiked water samples and cofactors, allowing batch optimization of binding conditions.
Final project ideas:
1. Cristalin clearance:
- the inner cells that made up the crystalline keep accumulating glucose molecules that, over time, leading to decreased lens clarity.
- these cells autodestroyed their organelles to become almost completely transparent, therefore they no longer possess the internal machinery for doing basically anything;
- they still have some types of chaperone cells, but unlike normal chaperones that are degraded by ubiquitin-proteosome complexes, the chaperones inside these cells enter “kamikaze mode” and once they capture a denatured protein, they sacrifice themselves and remain in the cell captive.
- Glycolization occurs when a glucose molecule binds to the N-terminal of a protein (this happens to chaperones in crystallin cells).
- the cells membrane still has a few “ports” where molecules smaller than 1200 daltons can enter (like glucose)
If I were to choose these subject for my final project I would research a way for how we could use engineered viruses (VLPs) to inject proteins (not nucleic acid) into the cristallin cells. The proteins injected can range from proteases to any protein that can help the crystallin become clear again. Or if the virus method fails I would most likely try using perforin or similar substances to introduces the proteins in the cell and out of the cell.
2. Aging neurons:
- Neurons in the brain rely heavily on mitochondria for energy production, but over time these organelles accumulate mutations in their DNA, leading to impaired function and cellular senescence.
- Unlike nuclear DNA, mitochondrial DNA lacks robust repair mechanisms, making it prone to oxidative damage from reactive oxygen species generated during metabolism.
- Aging neurons exhibit reduced mitophagy, the process where damaged mitochondria are selectively degraded and recycled, resulting in a buildup of dysfunctional units.
- Certain proteins like Parkin and PINK1 play key roles in tagging damaged mitochondria for removal, but their activity declines with age due to factors like protein misfolding.
- Neurons have limited ability to divide, so they must maintain long-term homeostasis through mechanisms like mitochondrial fusion and fission to distribute healthy organelles.
- If I were to choose this subject for my final project I would research a way to enhance mitophagy using CRISPR-based gene editing to upregulate Parkin and PINK1 expression in neuronal cell models. This could involve delivering the editing tools via adeno-associated viruses (AAVs) targeted to brain tissue. Or if gene editing proves inefficient, I would explore small molecule activators that mimic the effects of these proteins to clear damaged mitochondria and restore neuronal energy levels.
3. Bacterial biofilms
- Bacterial biofilms form protective communities on surfaces where cells are embedded in a matrix of extracellular polymeric substances, making them highly resistant to antibiotics. Within biofilms, bacteria enter a dormant state with slowed metabolism, reducing the effectiveness of drugs that target active processes like cell wall synthesis.
- Quorum sensing allows bacteria in biofilms to communicate and coordinate behaviors, including the upregulation of efflux pumps that expel antibiotics from cells.
- Persister cells, a subpopulation in biofilms, survive antibiotic treatment by entering a non-replicating phase and can repopulate the community once the threat is gone.
- The biofilm matrix acts as a physical barrier, limiting antibiotic penetration and binding to charged components that neutralize drug activity.
- If I were to choose this subject for my final project I would research a way to disrupt biofilm integrity using engineered bacteriophages that deliver enzymes like dispersin B to degrade the extracellular matrix. This could expose bacteria to standard antibiotics for better efficacy. Or if phage delivery is challenging, I would investigate nanoparticle carriers loaded with quorum-sensing inhibitors to prevent communication and weaken the biofilm structure from within.
Protein design part 1
Homework 4: Protein design part 1:
Part A:
- How many molecules of amino acids do you take with a piece of 500 grams of meat? (on average an amino acid is ~100 Daltons)
- 500g of meat => ~25% grams of protein => 125g of protein
- 125g of protein / 100 g per mole amino acids = 1.25 moles of AAs
- Nr of amino acids: 6.022 * 1023 * 1.25 = 7.5275 * 1023 amino acids in 500g of meat;
- Why do humans eat beef but do not become a cow, eat fish but do not become fish?
- What does it even mean to become a cow??? To act like a cow? To look like a cow? Both? I guess it means that your genome is indistinguishable from the one of a cow. Everything that you ingest is broken down into the monomers of that thing (or very close), because otherwise your cells will struggle to obtain the nutrients that they need. The nutrients present in a cow are the same as the ones found in humans..so it’s not the molecules that make us what we are, but the way they are arranged and function in our organism. Cells do not invent new genes or proteins (only accidentally). They use the nutrients to follow the instructions already present in them. In conclusion, to become a cow, you would need to rewrite your genome, and we can’t do that JUST by eating cows.
Why are there only 20 natural amino acids? Why would there be more than 20 alpha-AAs? Life found a way to work with those 20 amino acids, adding a new amino acid means increasing complexity and causing a lot of new structural problems. “The simpler, the better” - it applies to any system, because it minimizes problems. There aren’t just 20 “standard amino acids” because it is physically impossible for life to exist in any other way, there are only 20 because it worked.
Can you make other non-natural amino acids? Design some new amino acids.
A side chain containing an azobenzene group, which changes shape (from straight to bent) when hit by specific wavelengths of light. (answer generated with gemini ai thinking mode)
Further research:
Where did amino acids come from before enzymes that make them, and before life started? - Probably they appeared through spontaneous processes that occurred under the conditions that existed at that time on Earth. Conditions that were pretty extreme.
If you make an α-helix using D-amino acids, what handedness (right or left) would you expect? –>right-handed (L-amino acids are left-handed)
Can you discover additional helices in proteins? While the standard alpha helix is the most common, the sheer diversity of protein folding allows for variations that differ in their vertical rise and number of residues per rotation
Why are most molecular helices right-handed? Because to prevent overlapping, they form right-handed twists.
Why do β-sheets tend to aggregate? What is the driving force for β-sheet aggregation? beta-sheets tend to aggregate because they possess sticky exposed edges with unsatisfied hydrogen-bonding sites that seek to link with neighboring strands, creating a propitious environment for the formation of large, stable structures.
Why do many amyloid diseases form β-sheets? Can you use amyloid β-sheets as materials?
Design a β-sheet motif that forms a well-ordered structure.
Part B:
1.
Chosen protein: MAO B (Monoamine oxidase B)
- made of two identical subunits of 520 amino acids each; –> Further research:
- function: amine oxidation
- Because it creates ROS species, the overexpression of this protein damages surrounding tissue and is associated with Alzheimer’s and Parkinson’s;
2.
- Protein sequence: *
MSGDHLHNDS QIEADFRLND SHKHKDKHKD REHRHKEHKK EKDREKSKHS NSERDKERDK ERDKERDKER DREKDKDRER EREKEREKDS GKDRAKDKDR ERDRERDRER DREKDKEKDR DKDREKEKEK DREKDKEKDR DREKEKDRDK DKEKDRDKDK EKDRDKDKEK DRDKDKEKDR DKDKEKDRDK DKEKDRDKDK EKDRDKDKEK DRDKDKEKDR DKDKEKDRDK DKEKDRDKDK EKDRDKDKEK DRDKDKEKDR DKDKEKDRDK DKEKDRDKDK EKDRDKDKEK DRDKDKEKDR DKDKEKDRDK DKEKDRDKDK EKDRDKDKEK DRDKDKEKDR DKDKEKDRDK DKEKDRDKDK EKDRDKDKEK DRDKDKEKDR DKDKEKDRDK DKEKDRDKDK EKDRDKDKEK DRDKDKEKDR DKDKEKDRDK DKEKDRDKDK EKDRDKDKEK DRDKDKEKDR DKDKEKDRDK DKEKDRDKDK EKDRDKDKEK DRDKDKEKDR DKDKEKDRDK DKEKDRDKDK EKDRDKDKEK DRDKDKEKDR DKDKEKDRDK DKEKDRDKDK EKDRDKDKEK DRDKDKEKDR DKDKEKDRDK DKEKDRDKDK EKDRDKDKEK DRDKDKEKDR DKDKEKDRDK DKEKDRDKDK EKDRDKDKEK DRDKDKEKDR DKDKEKDRDK DKEKDRDKDK EKDRDKDKEK DRDKDKEKDR DKDKEKDRDK DKEKDRDKDK EKDRDKDKEK DRDKDKEKDR DKDKEKDRDK DKEKDRDKDK EKDRDKDKEK DRDKDKEKDR
- Leucine is the most frequent AA;
- 250 homologous proteins in other species;
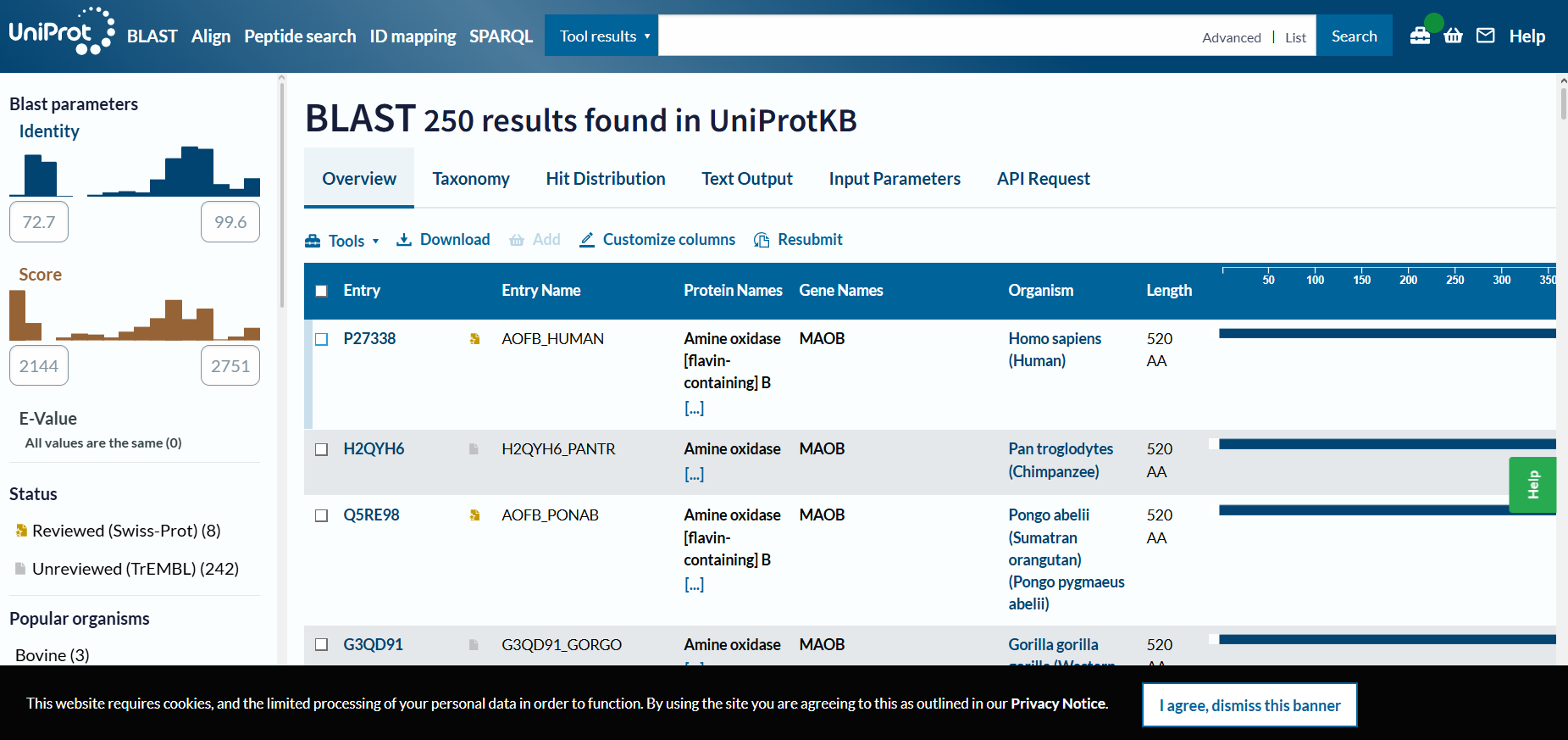

- MAO-B is part of amine-oxidase family and is a mitochondrial membrane-bound protein;
3.
- Resolution: 3.00 Å => the resolution is not the best (<2 A)
- Released Date: 2001-11-29
- MAO-B is part of amine-oxidase family
4.
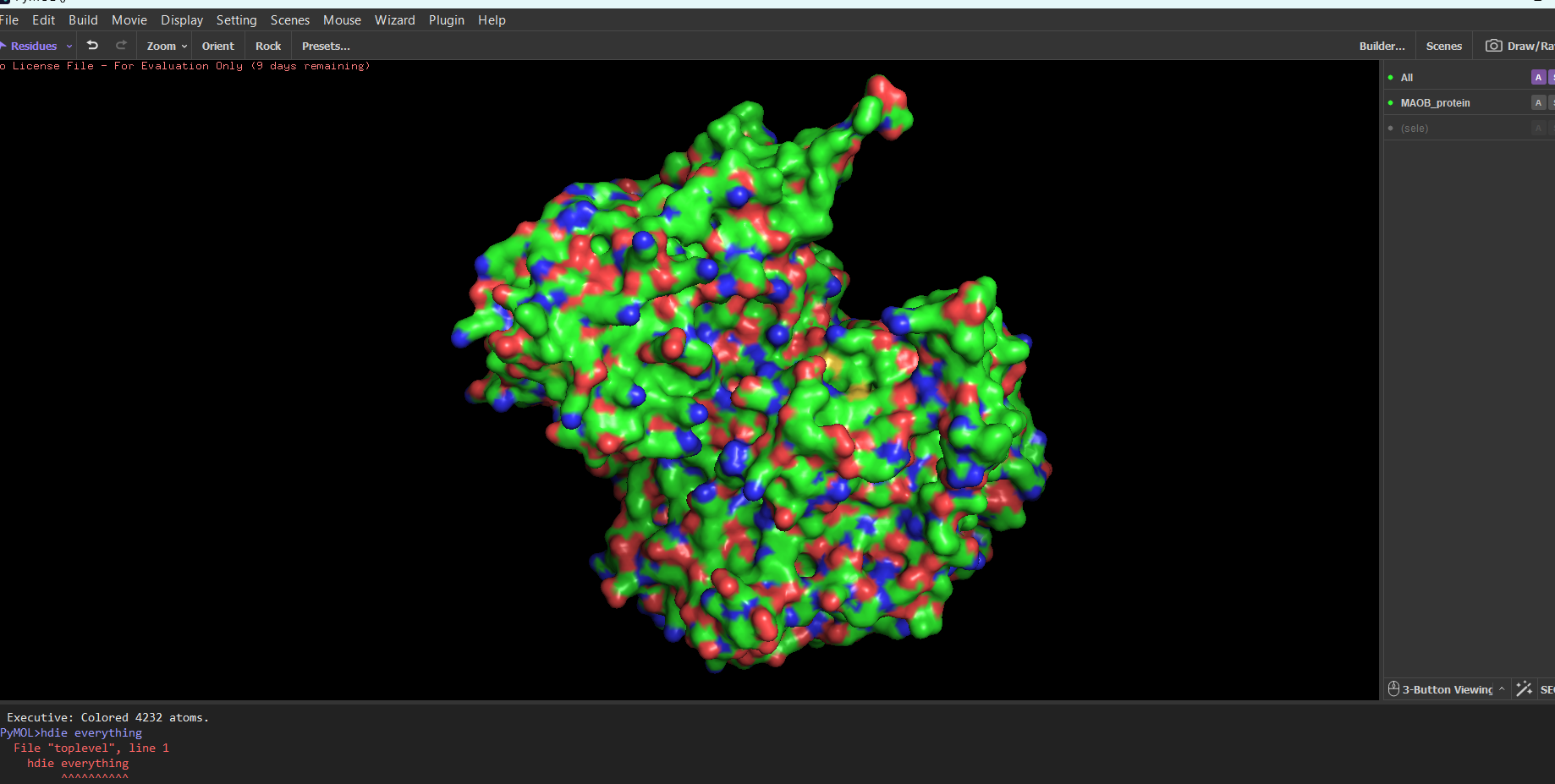
MAOB in pyMOL:
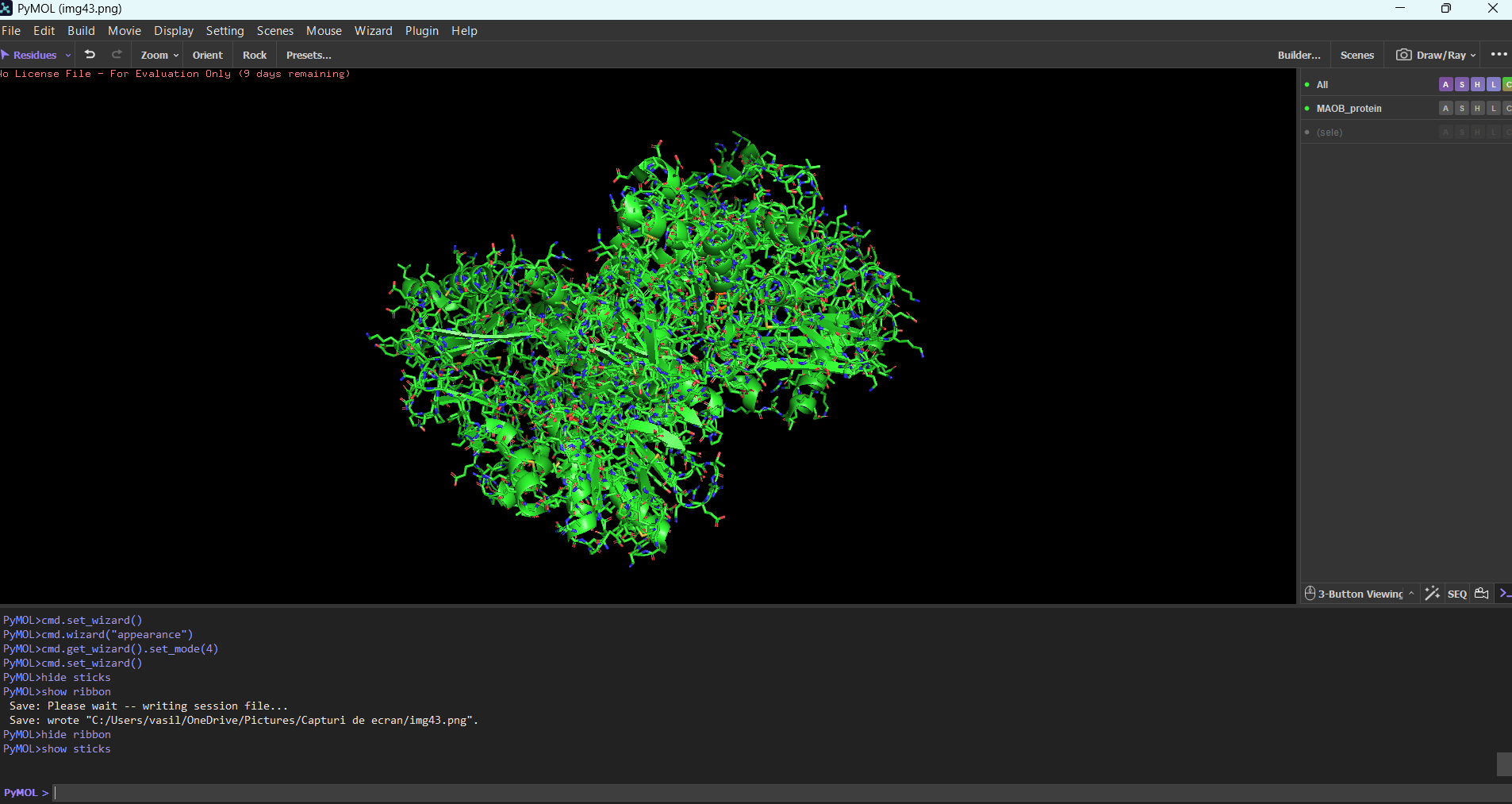

Ribbon mode:

Sticks mdoe:

Cartoon:

Note: The first time I tried to visualize the protein, pyMOL showed an overlapped version of multiple modes combined. For example, to see only the ribbon, I needed to type in the integrated command line: “hide everything” then you type “show ribbon”. Normally you should be able to do this using the GUI, but in my case the GUI didn’t work for some reason.
Protein colored by secondary structure:
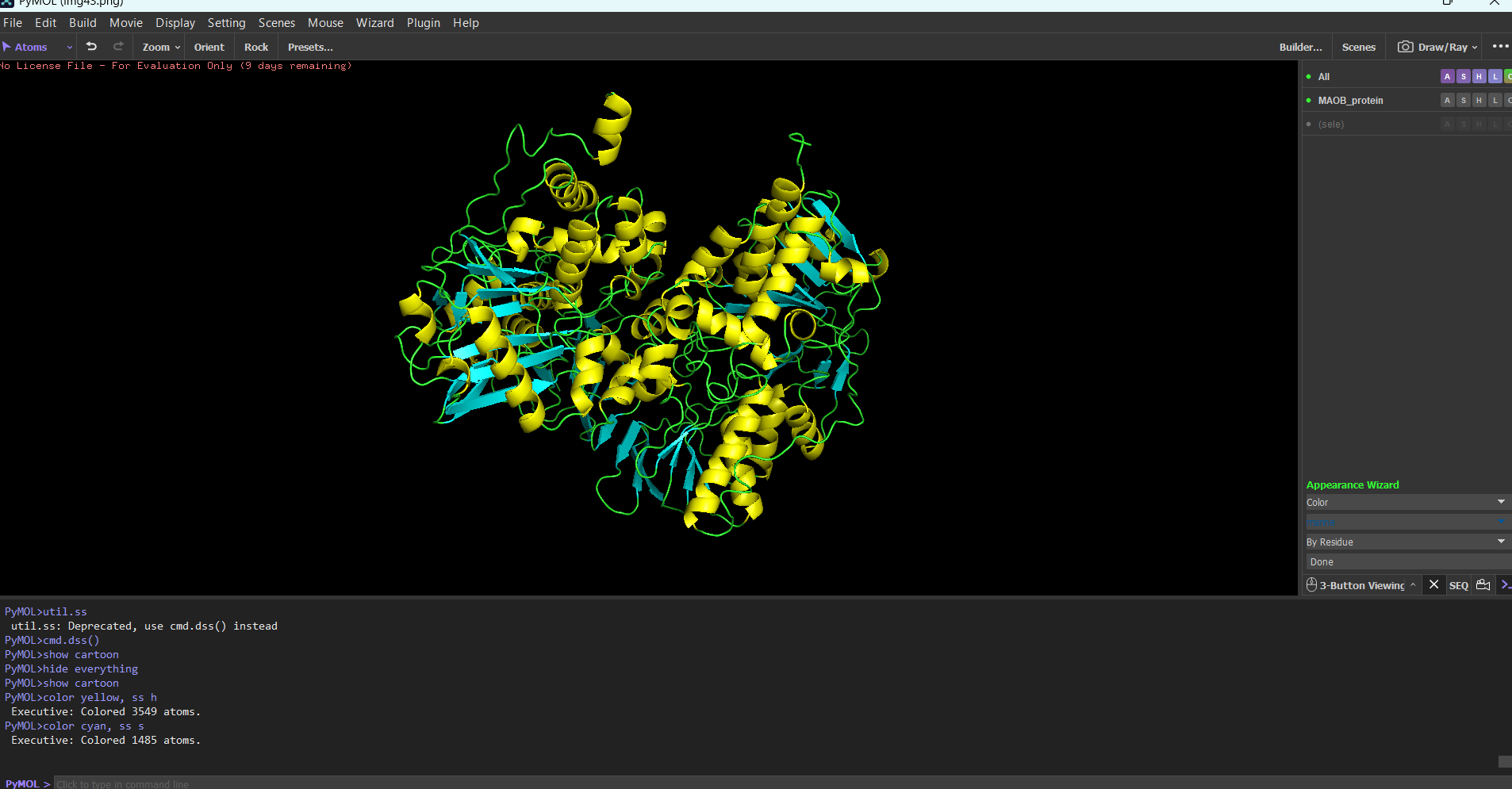
 It has roughly the same number of helices and beta sheets (maybe a few more beta sheets, but it differs from organism to organism most likely)
It has roughly the same number of helices and beta sheets (maybe a few more beta sheets, but it differs from organism to organism most likely)
Note: I used “color yellow, ss h” and “cyan, ss s” to color the secondary structures.
Further research: From what I read, proteins rich in helices often have a function that involves dynamic processes (enzymes, DNA binding mechanisms), and proteins rich in beta-sheets often provide mechanical support (structure). This statement is not always true (collagen).
Protein residue distribution:
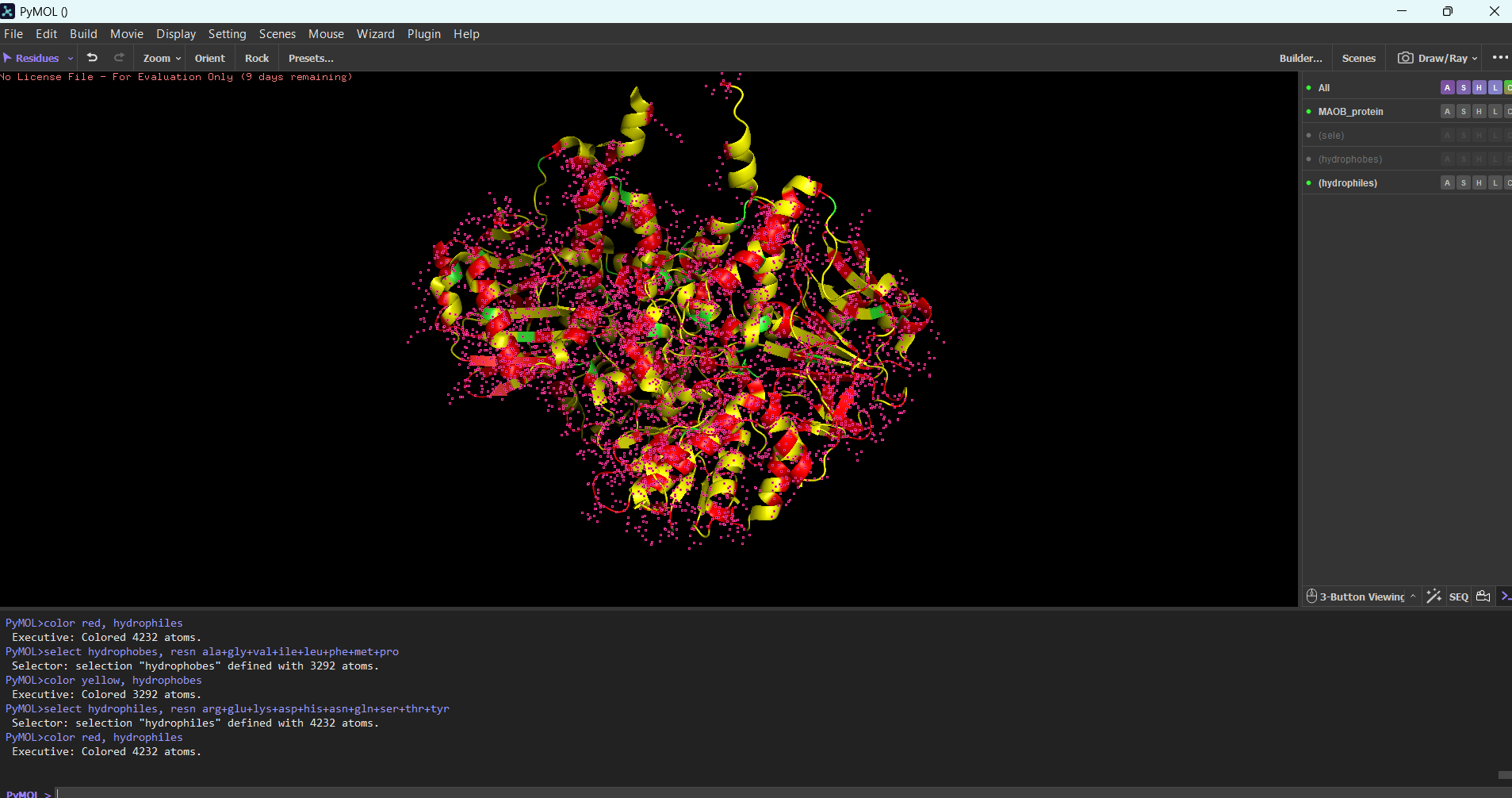

Hydrophobic and hydrophilic residues are distributed uniformly in this protein.
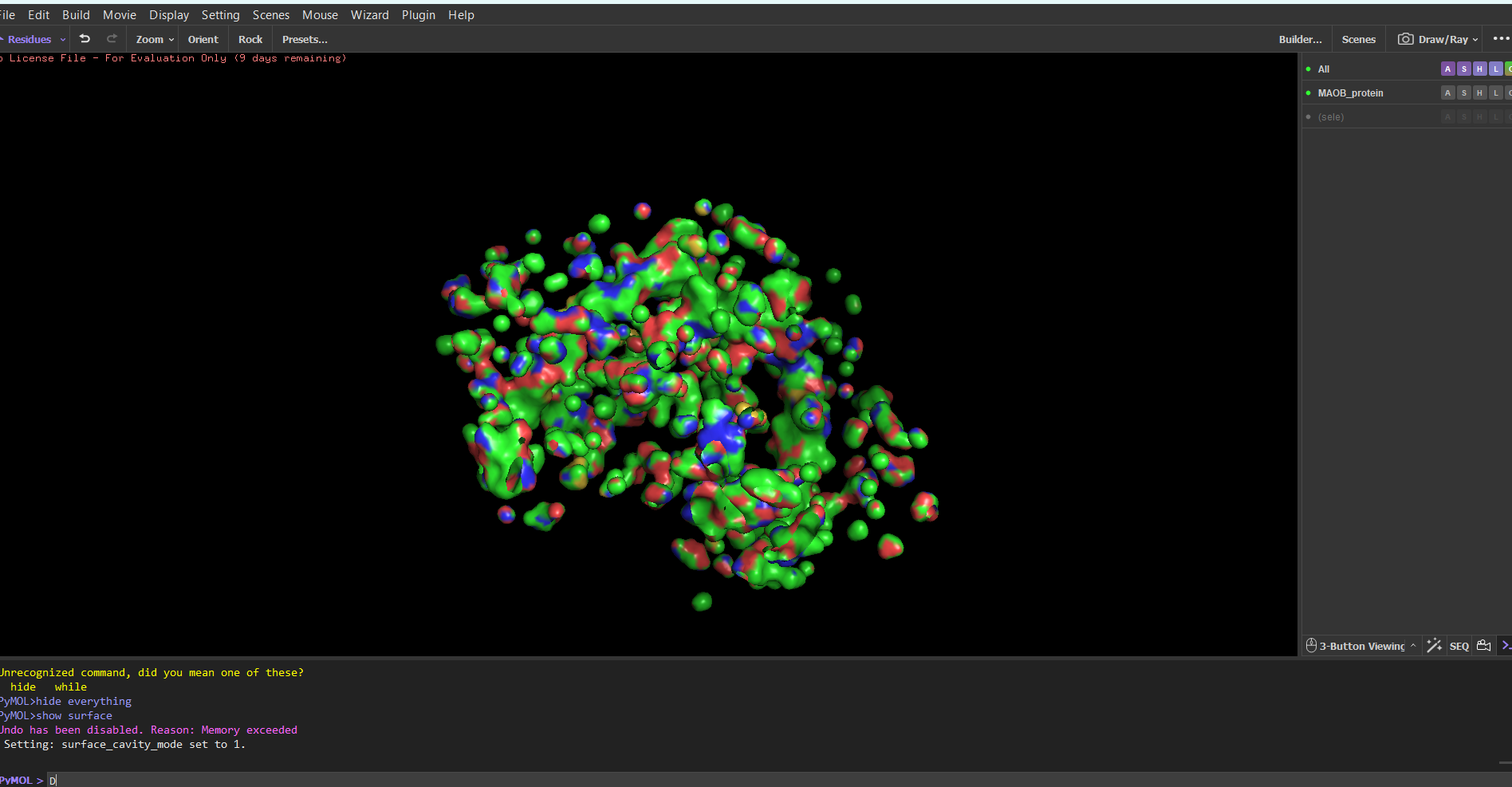
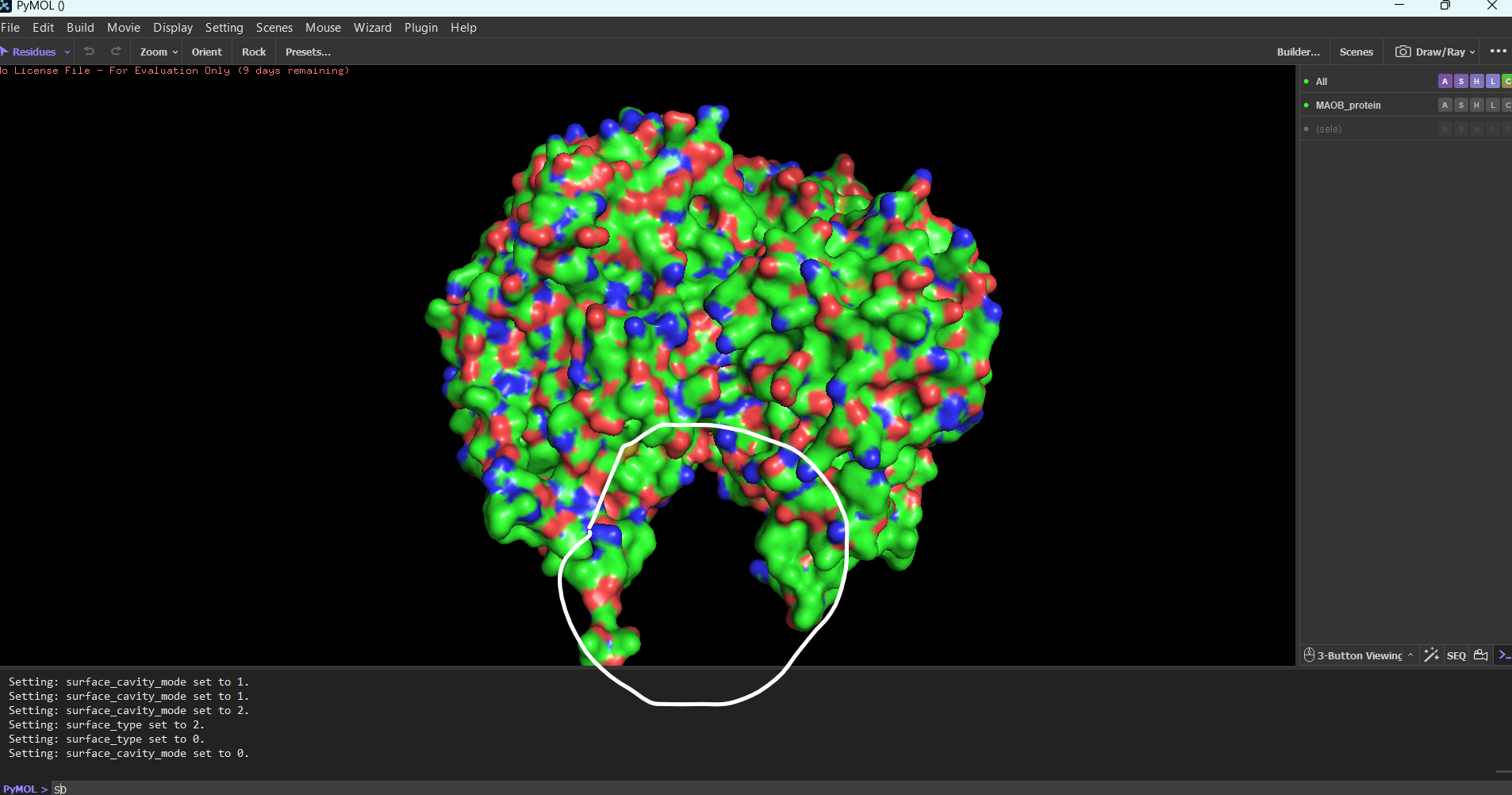
Surface mode:
Note: I tried showing only the cavities, because I thought it would be easier to find the binding sites. I still don’t know which ones are the biding sites.
Part C:
C1:
Glu Glu Leu Ile Thr Gly Val Leu Gly Ile Ser Ile Asp Leu Gly Met Val Thr Gly Ser Asp Leu Ala Lys Ala Val Lys Leu Ala Thr Gly Leu Gly Glu Ala Val Val Glu Gly Ala Lys Ala Val Gly Ser Val Leu Ala Leu Ser Thr Ala Leu Val Leu Ala Leu Leu Gly Leu Ala Leu Gly Leu Gly Leu Ala Leu Gly Leu Leu Gly Leu Ala Leu Gly Leu Leu Gly Leu Gly Leu Ala Leu Gly Leu Leu Gly Leu Ala Leu Gly Leu Leu Gly Leu Gly Leu Ala Leu Gly Leu Leu Gly Leu Ala Leu Gly Leu Leu Gly Leu Gly Leu Ala Leu Gly Leu Leu Gly Leu Ala Leu Gly Leu Leu Gly Leu Gly Leu Ala Leu Gly Leu Leu Gly Leu Ala Leu Gly Leu Leu Gly Leu Gly Leu Ala Leu Gly Leu Leu Gly Leu Ala Leu Gly Leu Leu Gly Leu Gly Leu Ala Leu Gly Leu Leu Gly Leu Ala Leu Gly Leu Leu Gly Leu Gly Leu Ala Leu Gly Leu Leu Gly Leu Ala Leu Gly Leu Leu Gly Leu Gly Leu Ala Leu Gly Leu Leu Gly Leu Ala Leu Gly Leu Leu Gly Leu Gly Leu Ala Leu Gly Leu Leu Gly Leu Ala Leu Gly Leu Leu Gly Leu Gly Leu Ala Leu Gly Leu Leu Gly Leu Ala Leu Gly Leu Leu Gly Leu Gly Leu Ala Leu Gly Leu Leu Gly Leu Ala Leu Gly Leu Leu Gly Leu Gly Leu Ala Leu Gly Leu Leu Gly Leu Ala Leu Gly Leu Leu Gly Leu
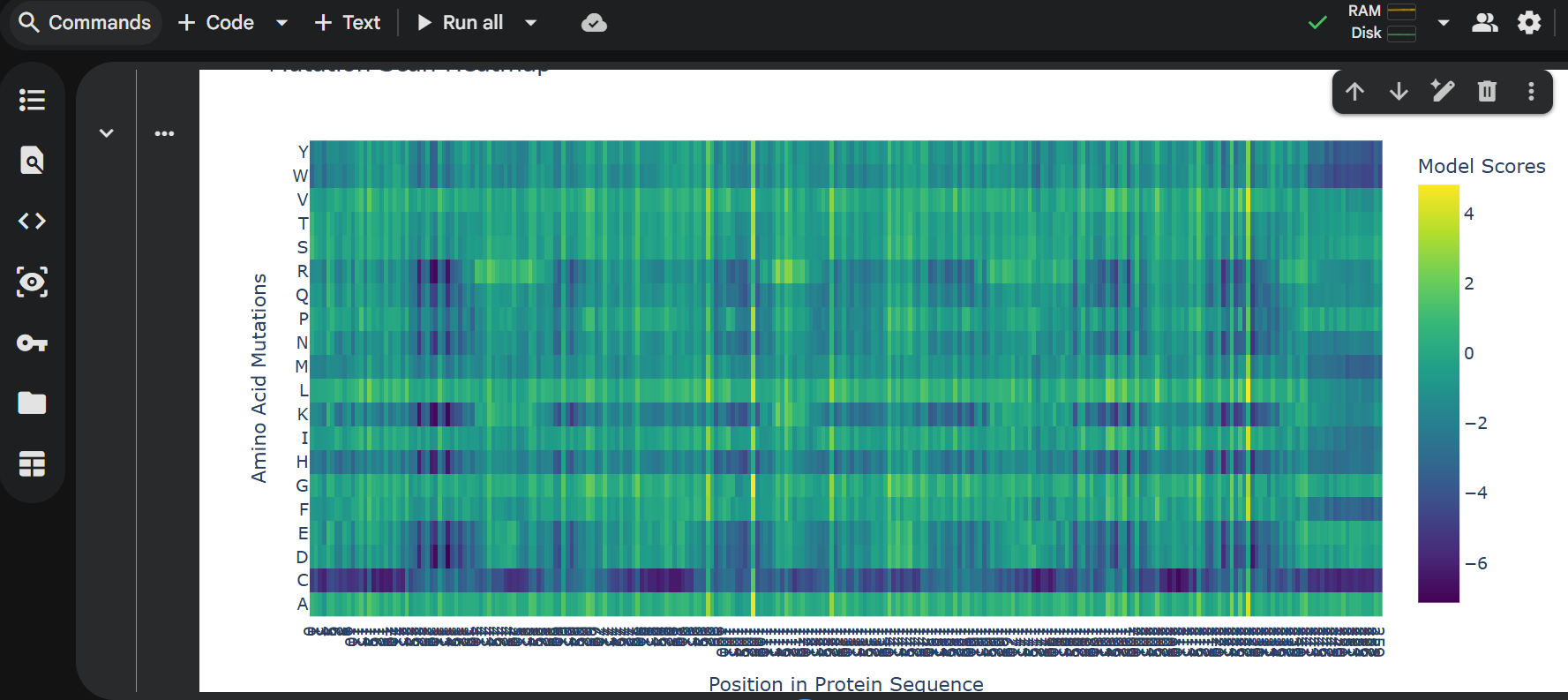

C2:
sp|P02144|MYG_HUMAN Myoglobin OS=Homo sapiens OX=9606 GN=MB PE=1 SV=2 MGLSDGEWQLVLNVWGKVEADIPGHGQEVLIRLFKGHPETLEKFDKFKHLKSEDEMKASE DLKKHGATVLTALGGILKKKGHHEAEIKPLAQSHATKHKIPVKYLEFISECIIQVLQSKH PGDFGADAQGAMNKALELFRKDMASNYKELGFQG
Folded with ESMFold:
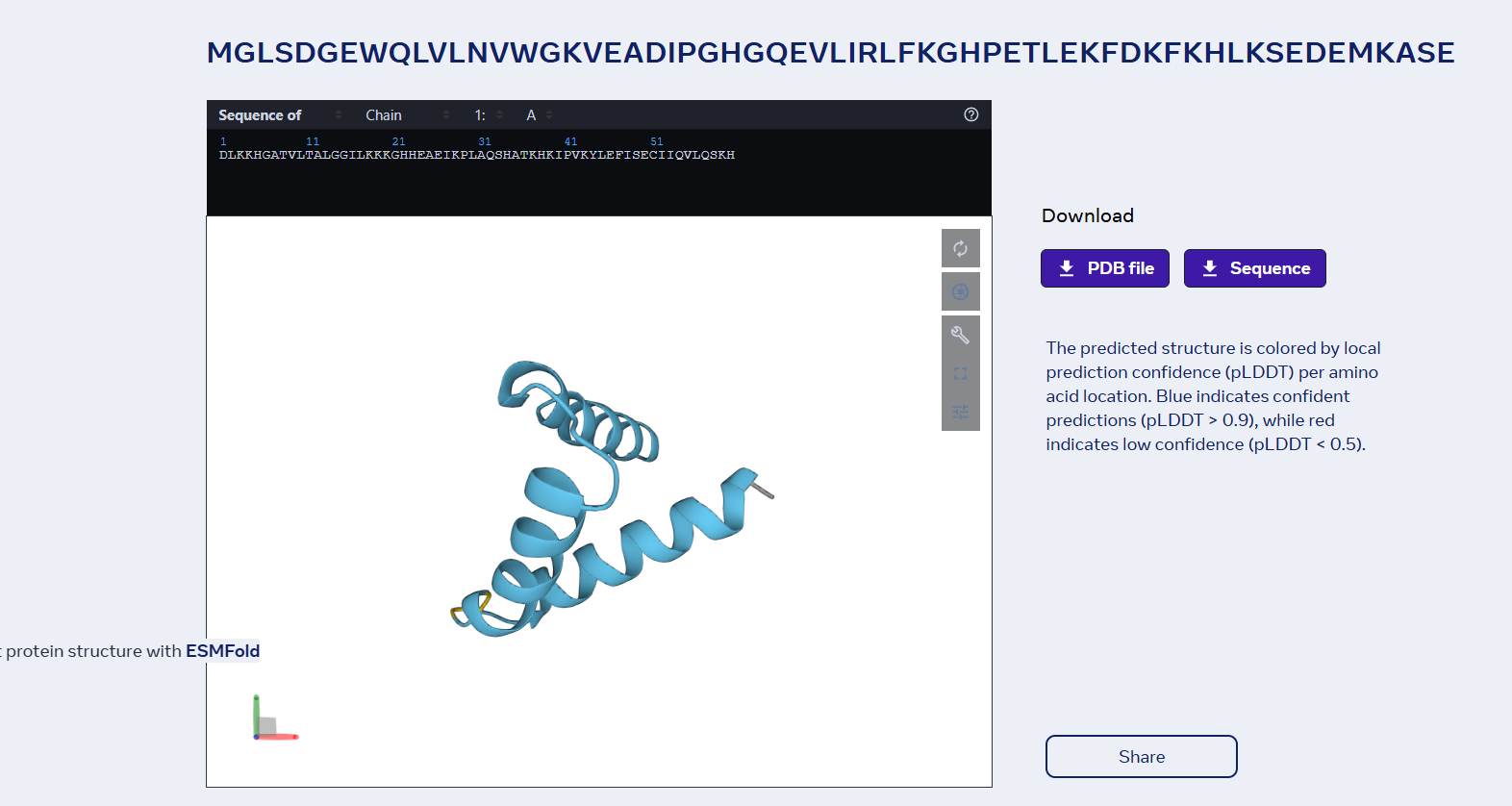

Folded with Boltz-1:
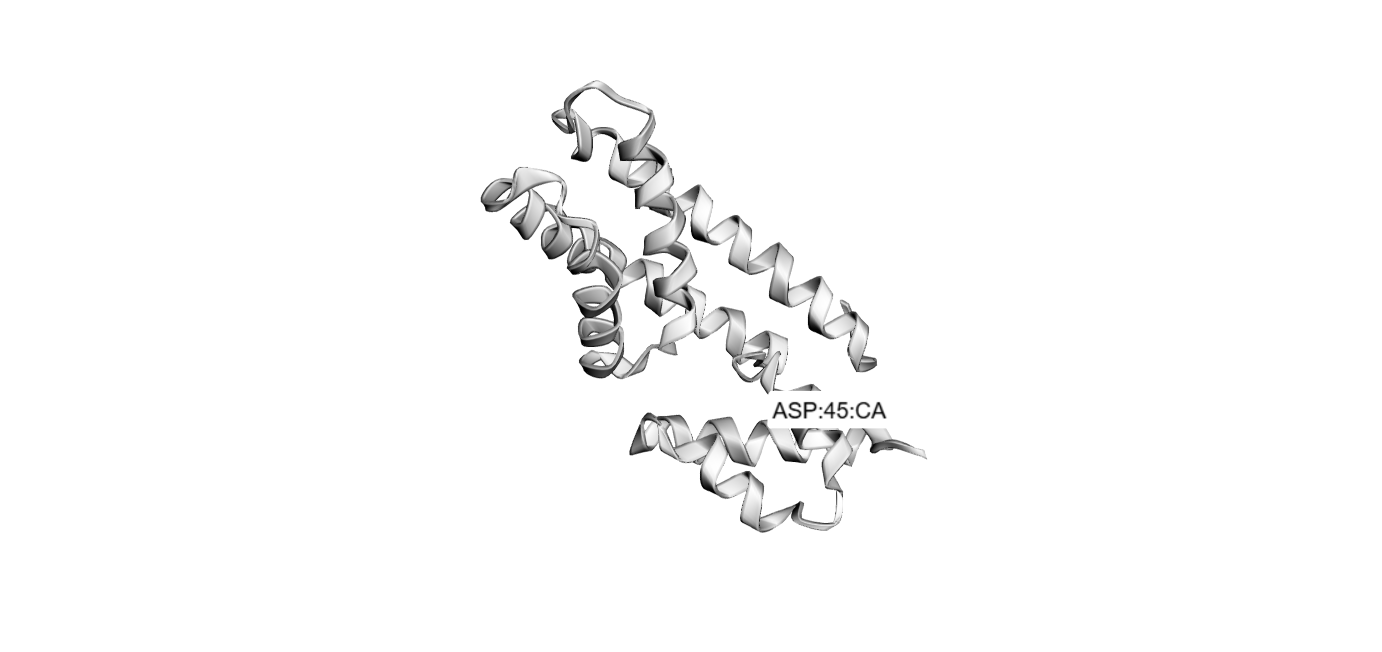

Folded with SwissDock:
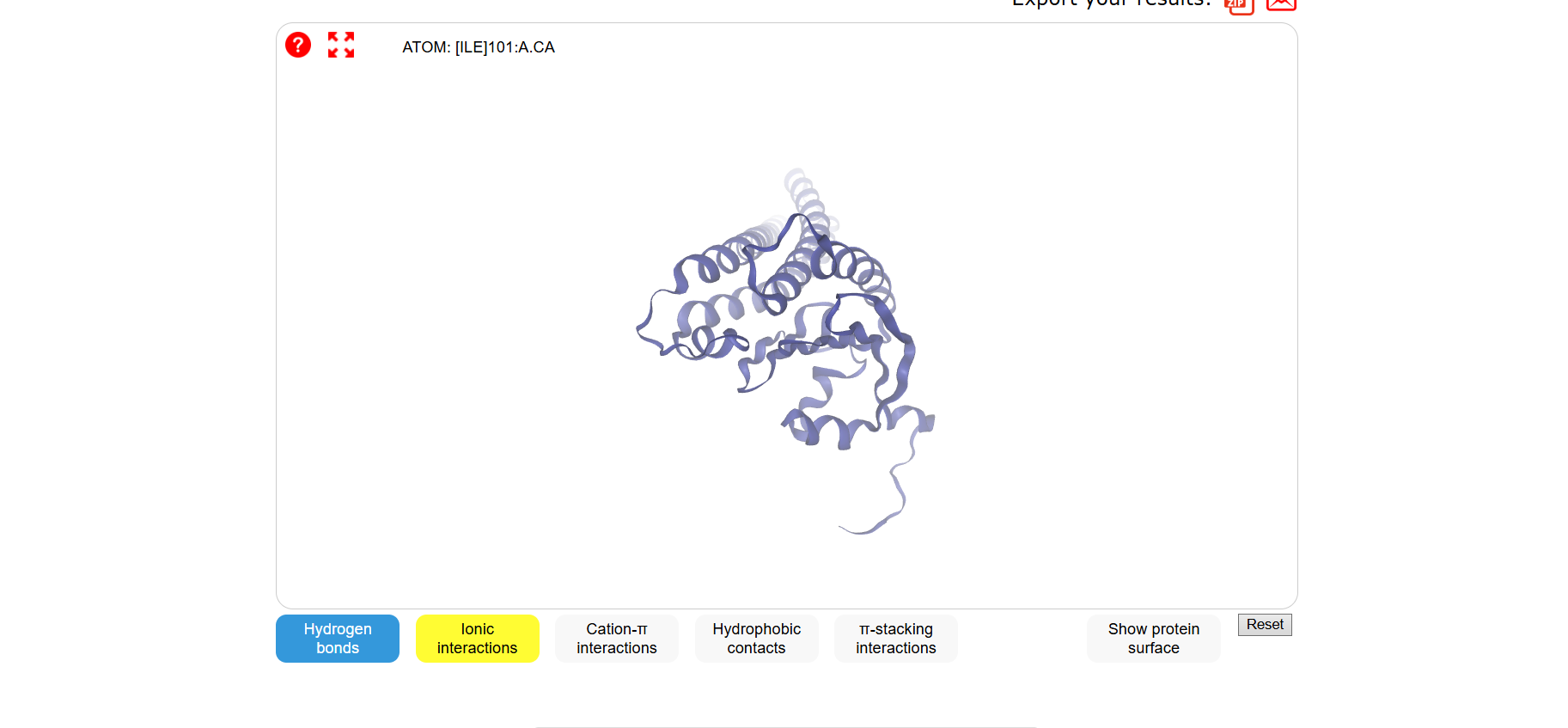

3 one point mutations:
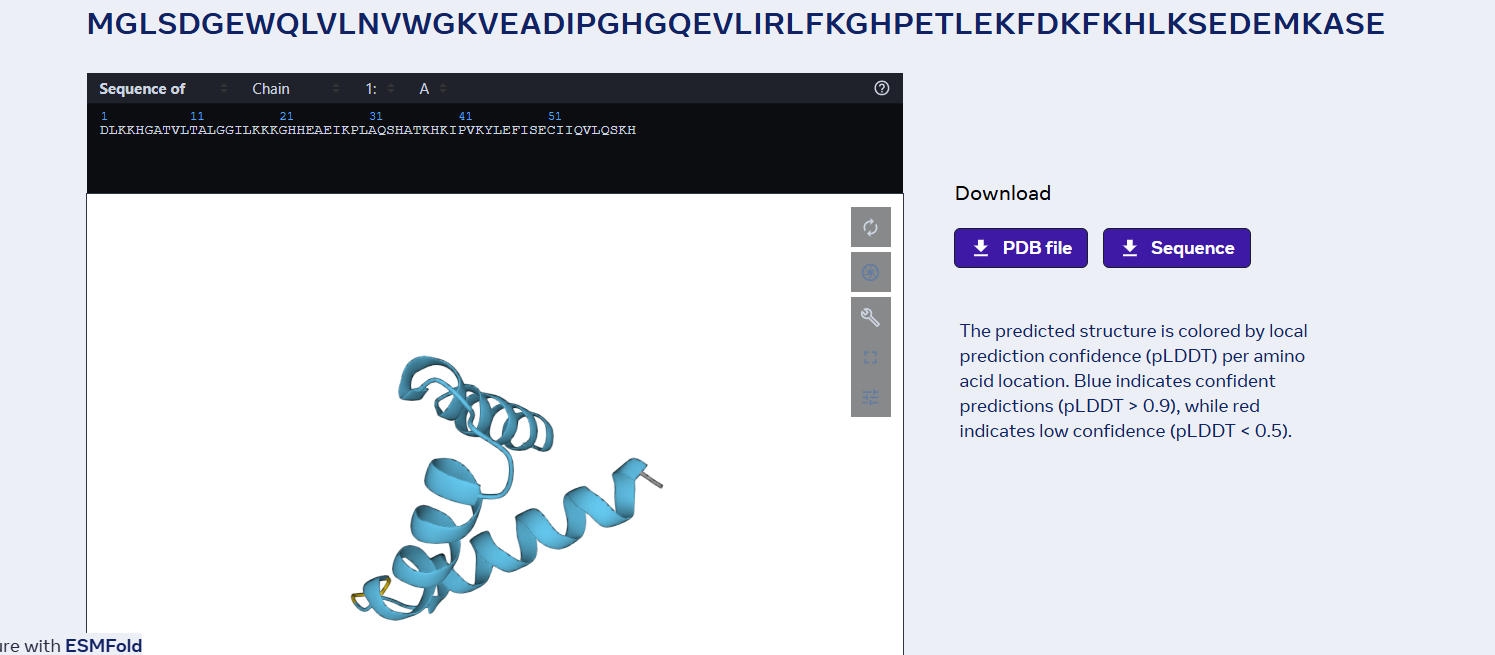

last 6 proteins mutation:
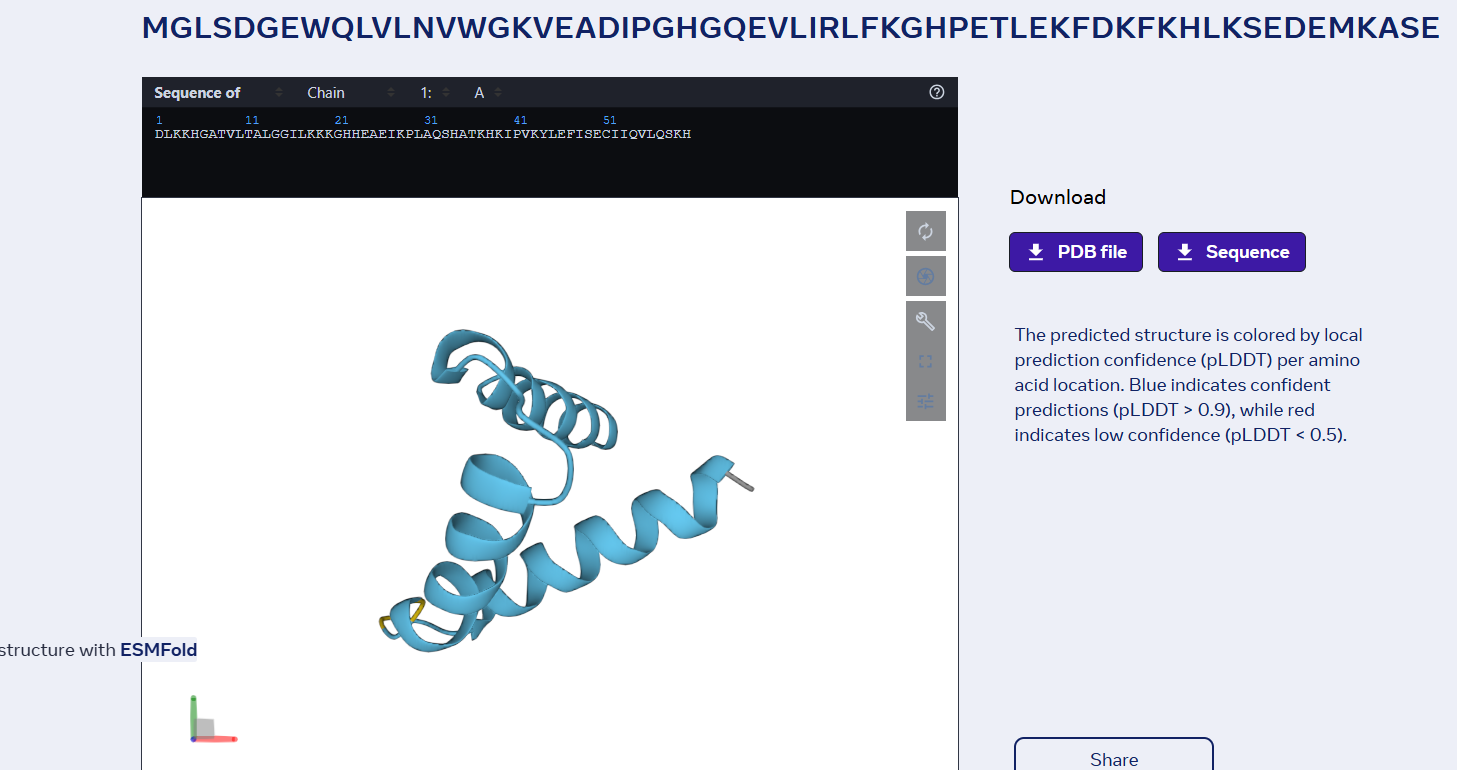

Part D:
Project Proposal: Engineering a Minimal MS2-L Lysis Engine
Primary Goal: Our group aims to increase the functional stability of the MS2 lysis (L) protein. We will achieve this by eliminating the N-terminal domain (residues 1–36). This truncation removes the “regulatory brake” that normally makes lysis dependent on the host chaperone DnaJ, resulting in a more potent, autonomous lysis protein, thus lysis will be achieved a lot faster, beacuse MS2-L will be functional from the moment of translation which gives less time for the proteases to degrade it before it attached to the cellular membrane.
Tools & Approaches We propose a computational pipeline to validate and optimize this truncated variant:
AlphaFold3 / AlphaFold-Multimer: We will model the truncated L protein in a cramped lipid bilayer environment to predict how the remaining transmembrane helix (TMH) inserts itself. We will also use it to confirm the loss of binding affinity to E. coli DnaJ.
Protein Language Models (ESM-2 / ESM-1v): We will use these models to perform in silico mutagenesis on the remaining C-terminal sequence. Our goal is to identify “stabilizing” mutations that strengthen the alpha-helical propensity of the membrane-spanning region.
Molecular Dynamics (MD) Simulations (OpenMM or Gromacs): Since lysis involves membrane distortion, we will simulate the truncated protein within a model bacterial membrane to observe its ability to disrupt lipid packing.
- Potential Pitfalls Membrane Toxicity in in silico models: Most protein design tools are trained on soluble proteins. Modeling a protein whose entire job is to destroy the membrane (lysis) may lead to unstable simulations or “unphysical” results.
Expression Levels: In a real-world lab setting, a highly toxic protein might kill the host cells so quickly that we cannot produce high enough titers of the phage for study.
- Pipeline Schematic Design: Truncate N-terminus (1-36).
Optimize: Run ESM-1v to find high-probability stabilizing mutations.
Predict: Fold top candidates in AlphaFold3 to ensure TMH orientation.
Simulate: Insert into a virtual membrane to verify disruptive “toxicity.”
Homework 5 Protein Design Part II
Protein Design Part II Homework:
Part 1
- protein sequence (with A4V mutation): ATKVVCVLKGDGPVQGIINFEQKESNGPVKVWGSIKGLTEGLHGFHVHEFGDNTAGATSAGPHFNPLSRKHGGPKDEERHVGDLGNVTADKDGVADVSIEDSVISLSGDHCIIGRTLVVHEKADDLGKGGNEESTKTGNAGSRLAAGVIGIAQ

Part 2:
- Firstly, I want to highlight a few hallmarks of the human protein SD01:
- The N-terminus is located near the C-terminus
- The residues 3 to 9, thus including the A4V mutation, form a beta-strand.
- Human SD01 is rich in beta-strands
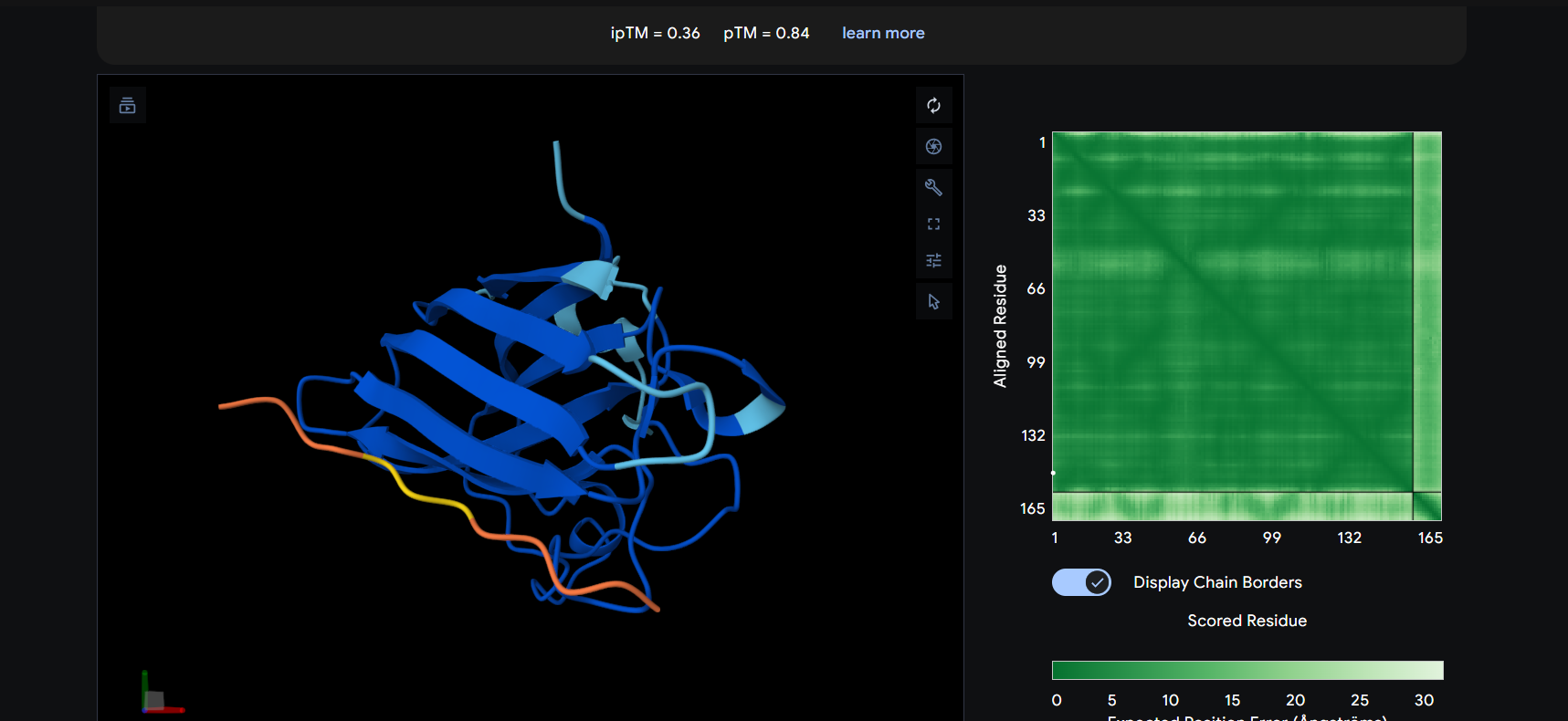

The peptide appears to bind on the surface of the protein in the opposite part of the N-terminus.
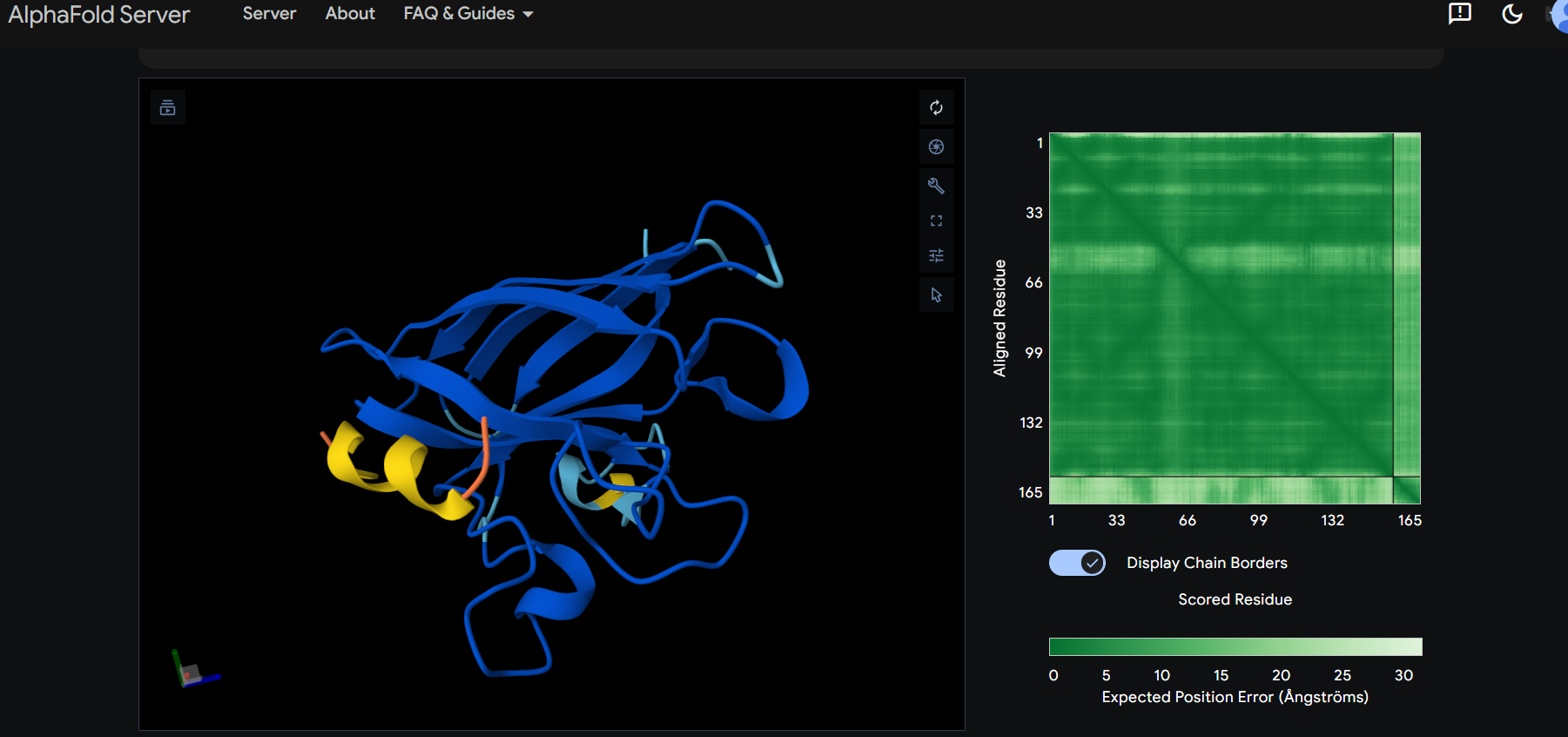

The peptide appears to bind on the surface of the protein in the opposite part of the N-terminus.
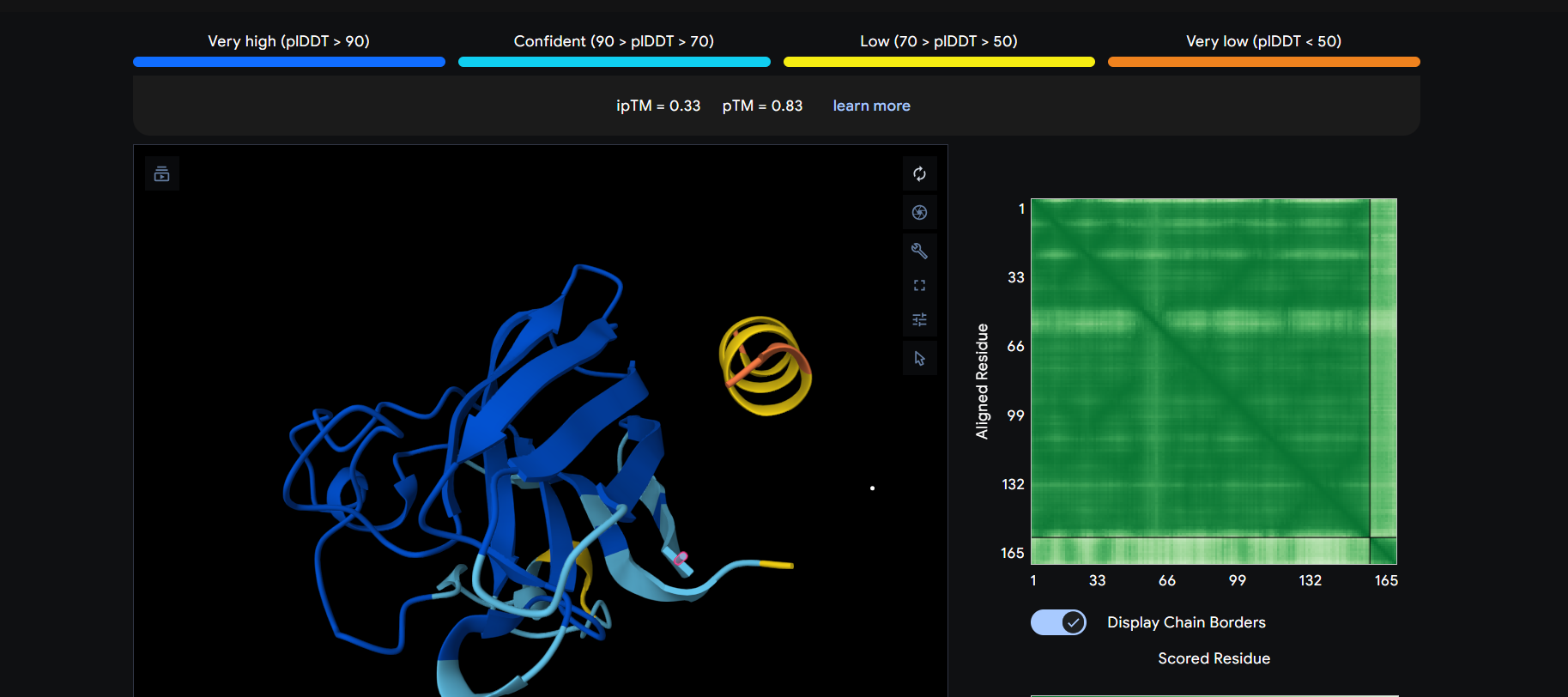

The peptide appears to bind on the surface of the protein near the N-terminus.
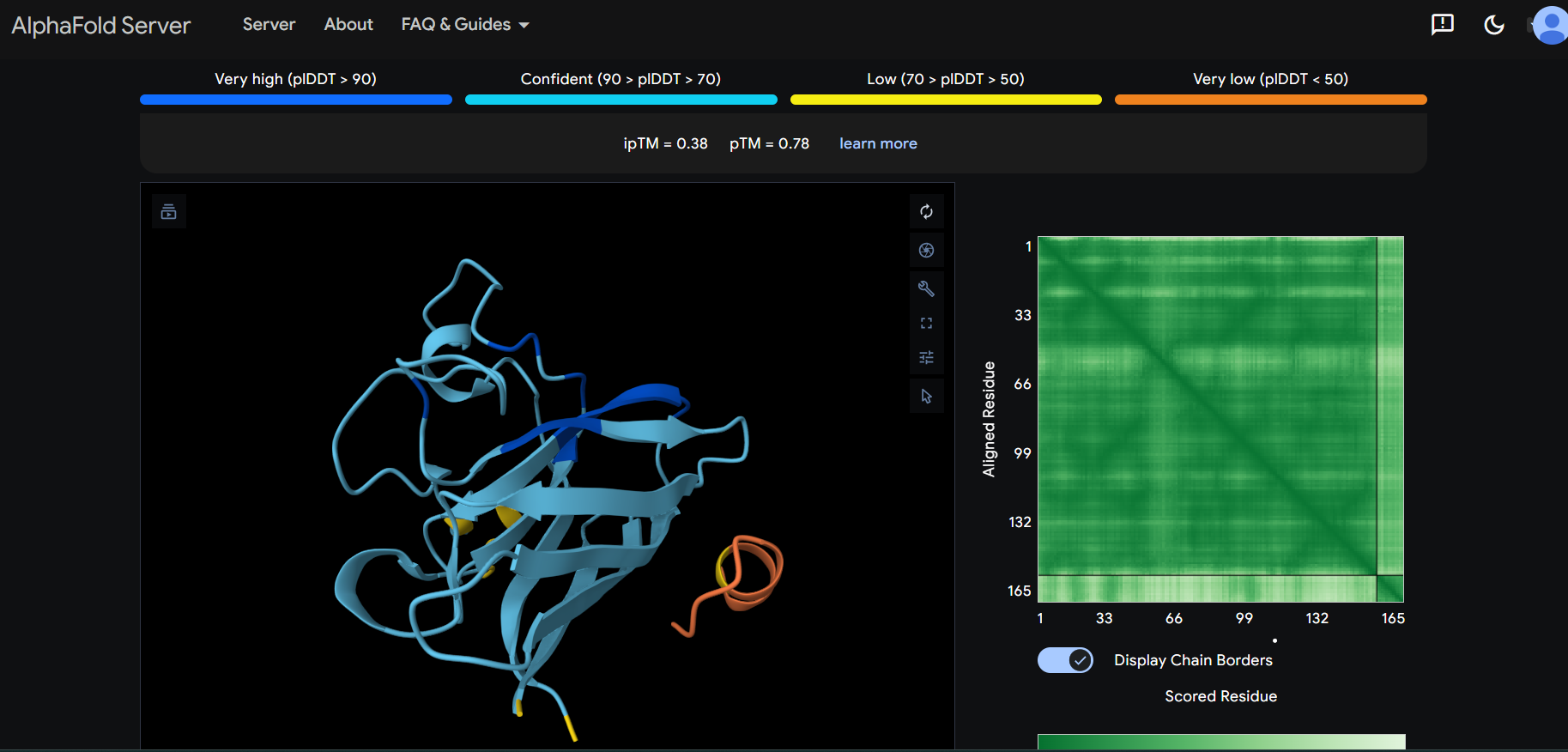

The peptide appears to bind on the surface of the protein aproximatevely near the N-terminus.
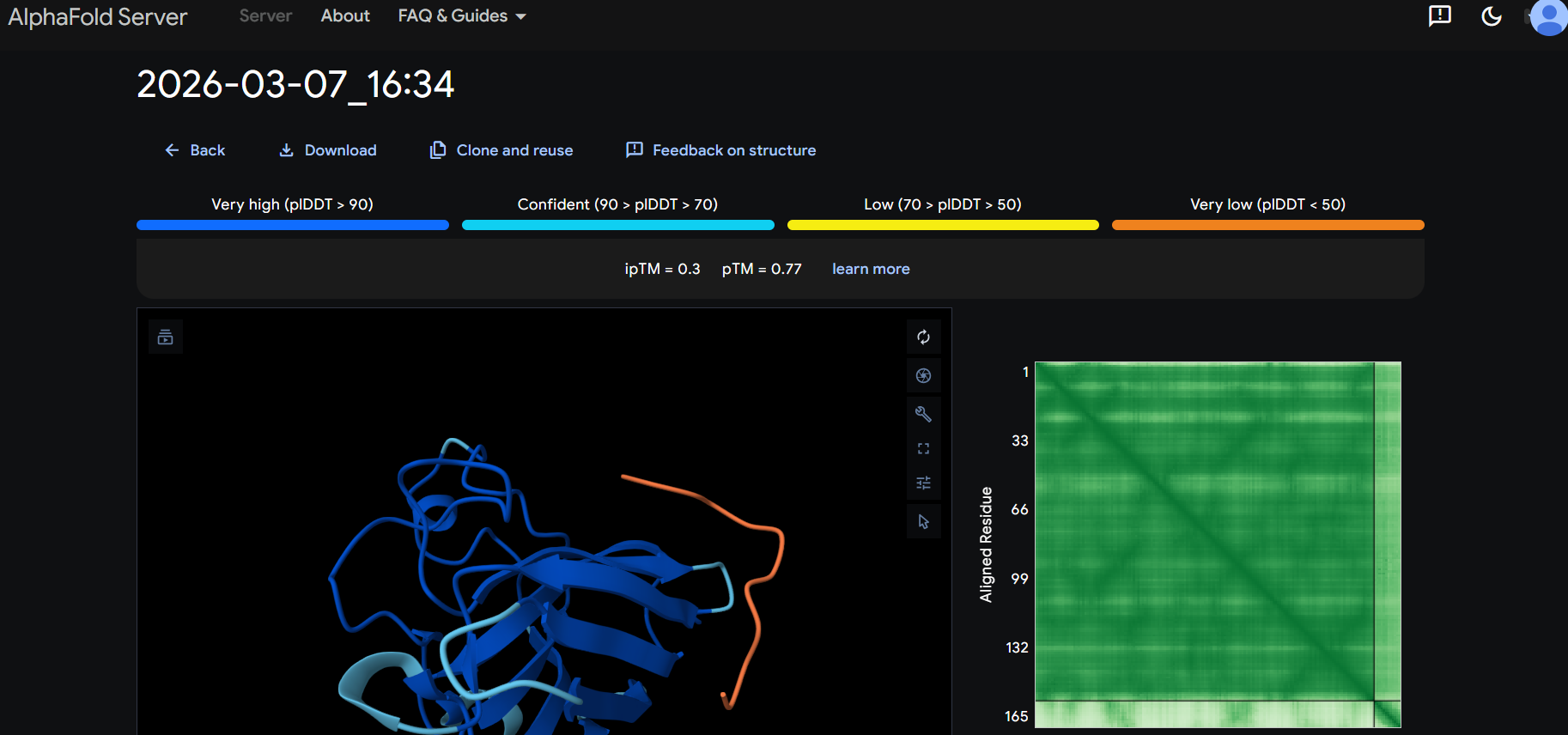

The peptide appears to bind on the surface of the protein aproximatevely near the N-terminus, near the beta-sheet formed by SD01.
The ipTM score for all the predicted SDo1-peptide interactions is below what it would be considered reliable.
The prediction are incongruent with the knwon binding sites.
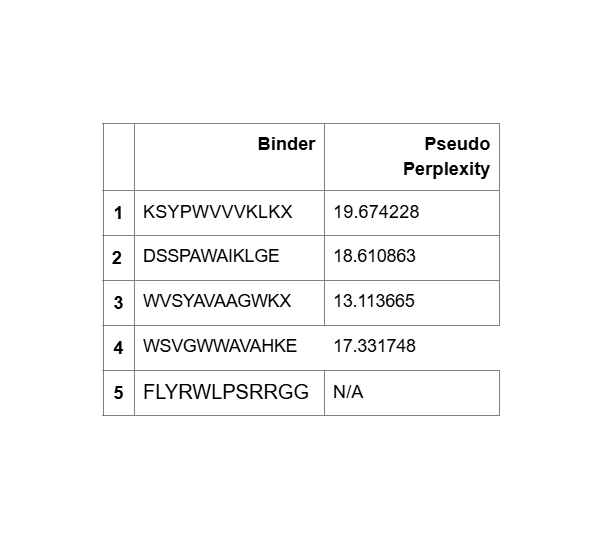
Part 3:
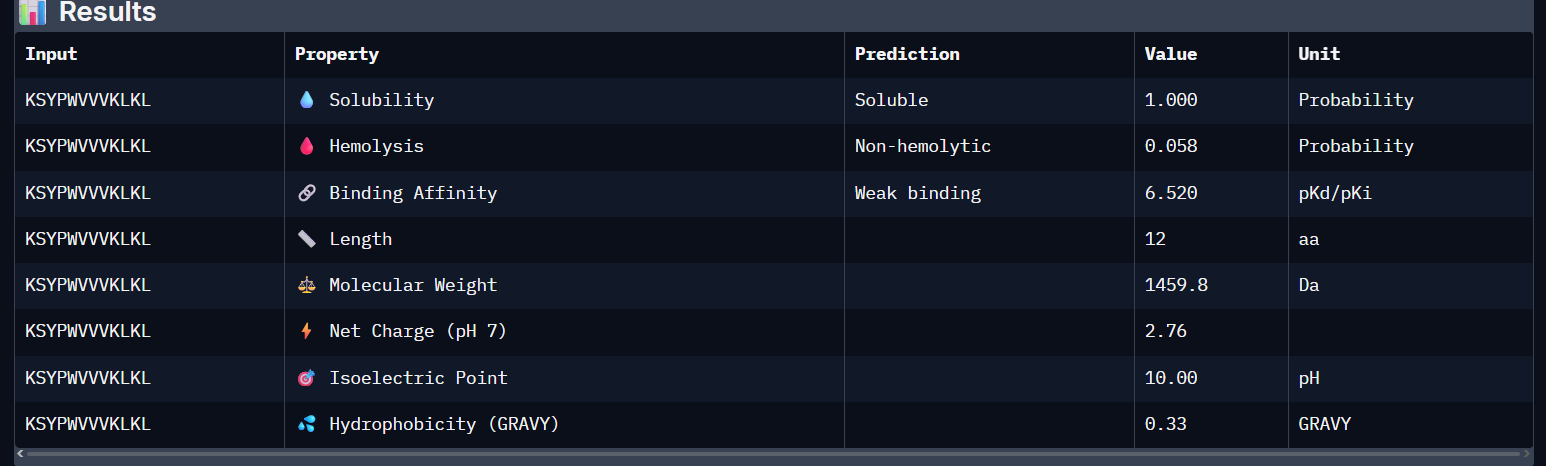
KSYPWVVVKLKL:
 DSSPAWAIKLGE:
DSSPAWAIKLGE:
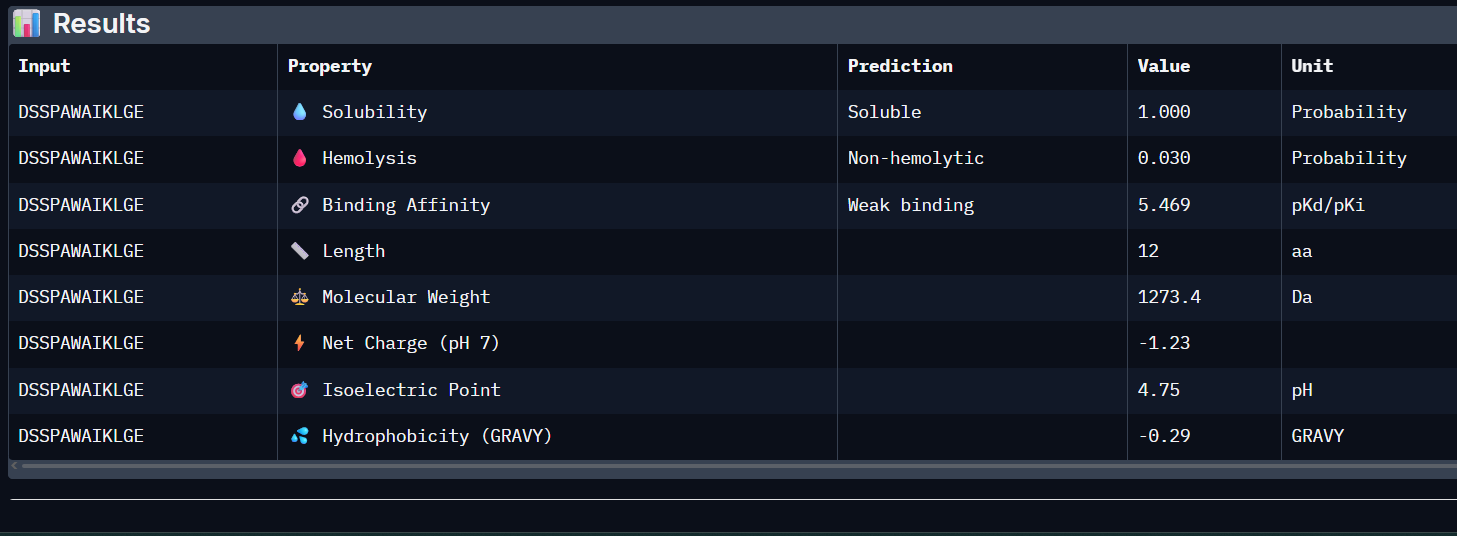
 WVSYAVAAGWKL:
WVSYAVAAGWKL:
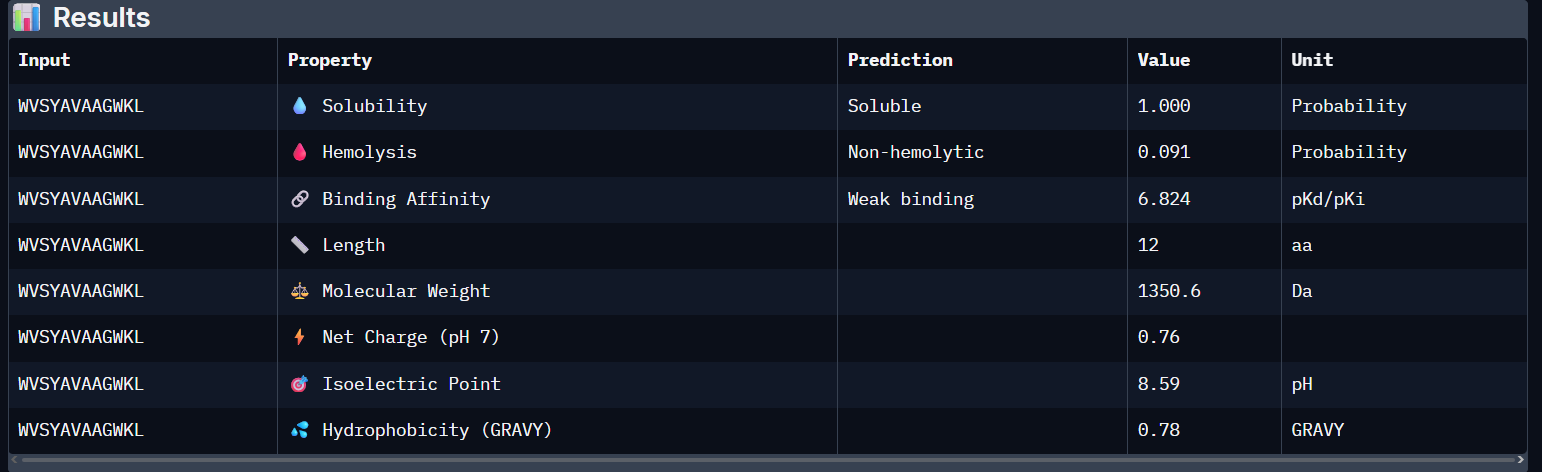
 WSVGWWAVAHKE:
WSVGWWAVAHKE:
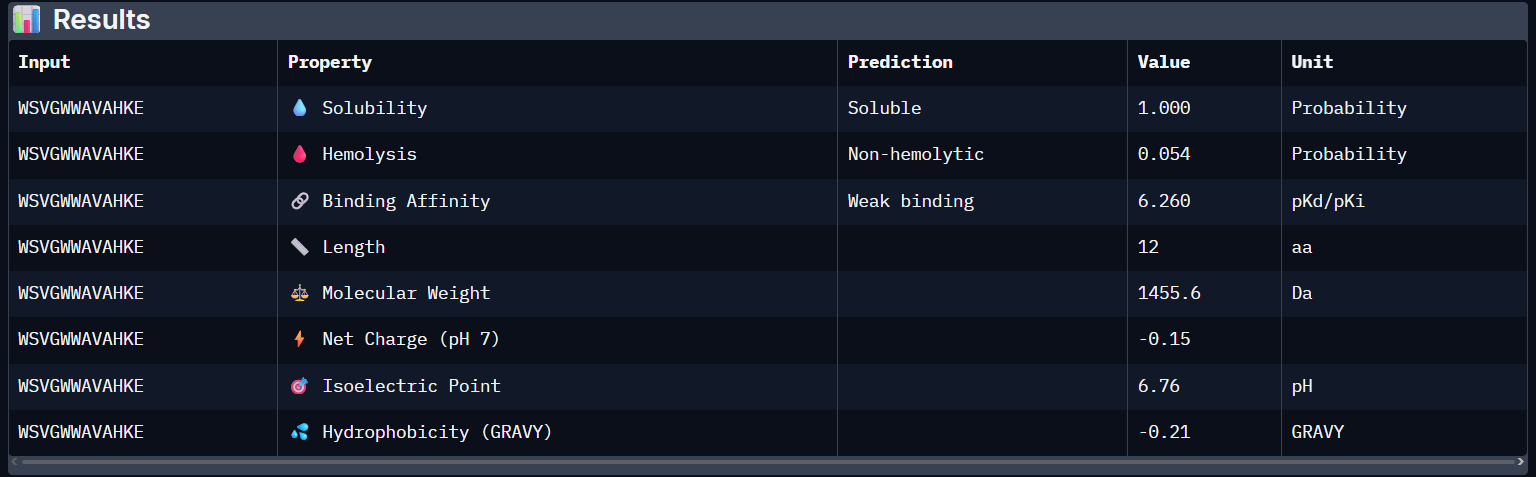
 FLYRWLPSRRGG:
FLYRWLPSRRGG:
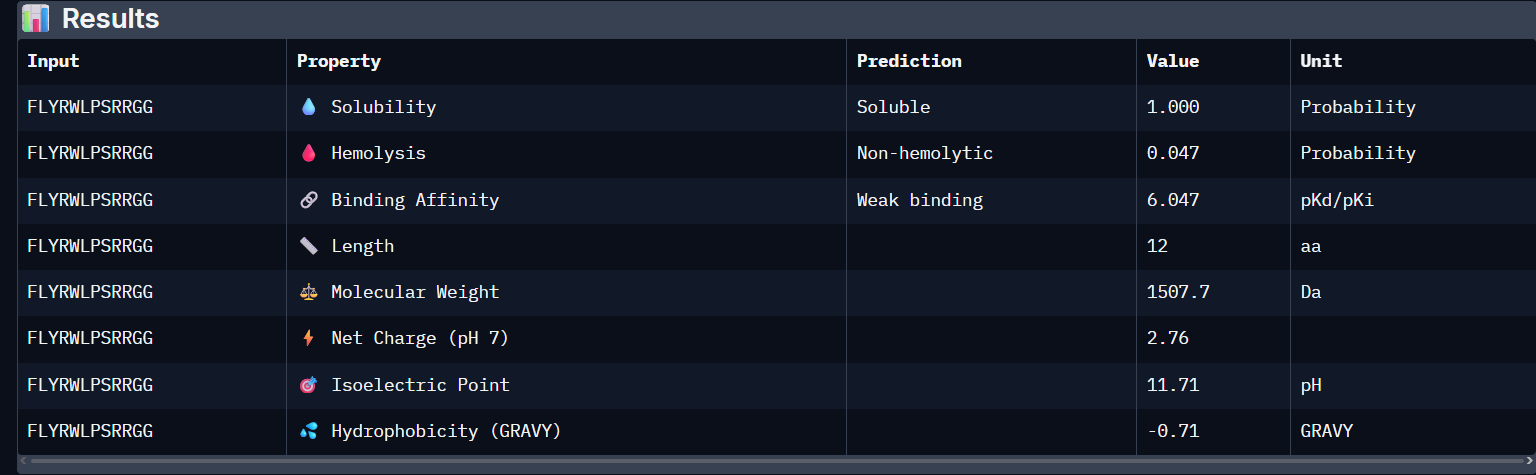

- Unfortunately, all the peptides showed above are soluble, weak binding affinity (which corresponds with the low ipTM score) and are non-hemolytic, thus there is no much to say about them (( .
“Choose one peptide you would advance and justify your decision briefly”:
- WVSYAVAAGWKL has the highest affinity ~6.8 and appears to be the best choice;
Part 4:
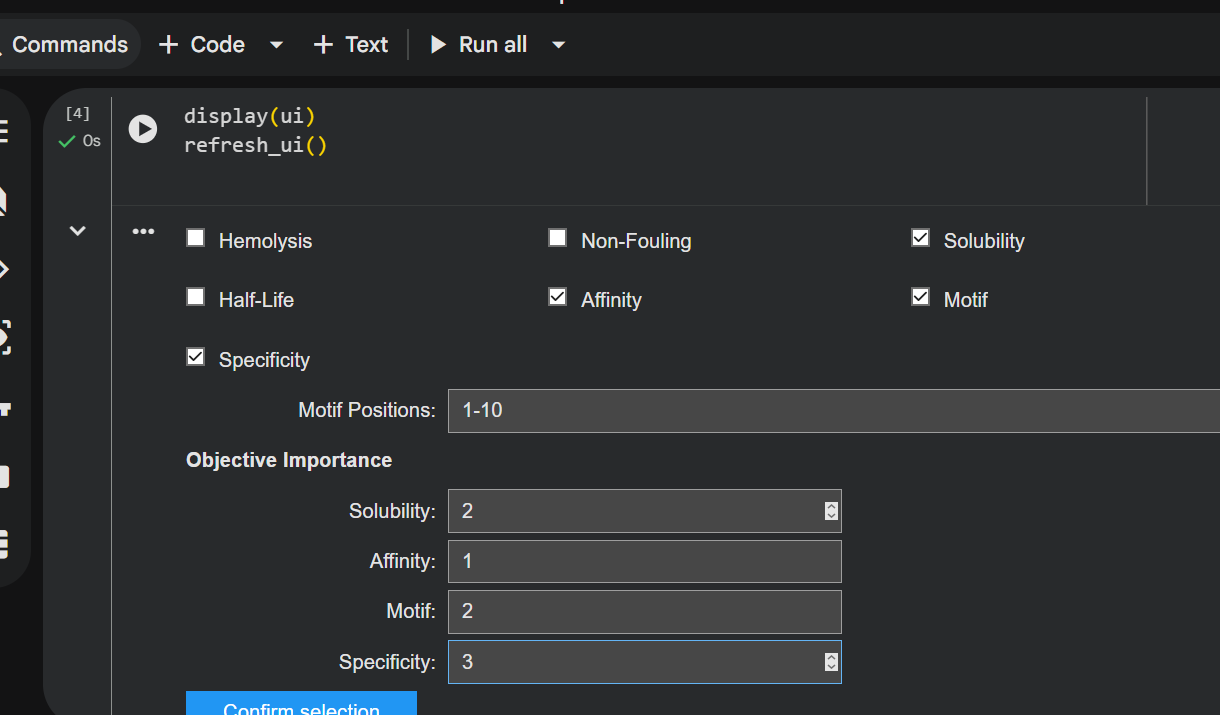

Generated peptide: CQGVFWTMCSHD
- nr. of amino acids: 12
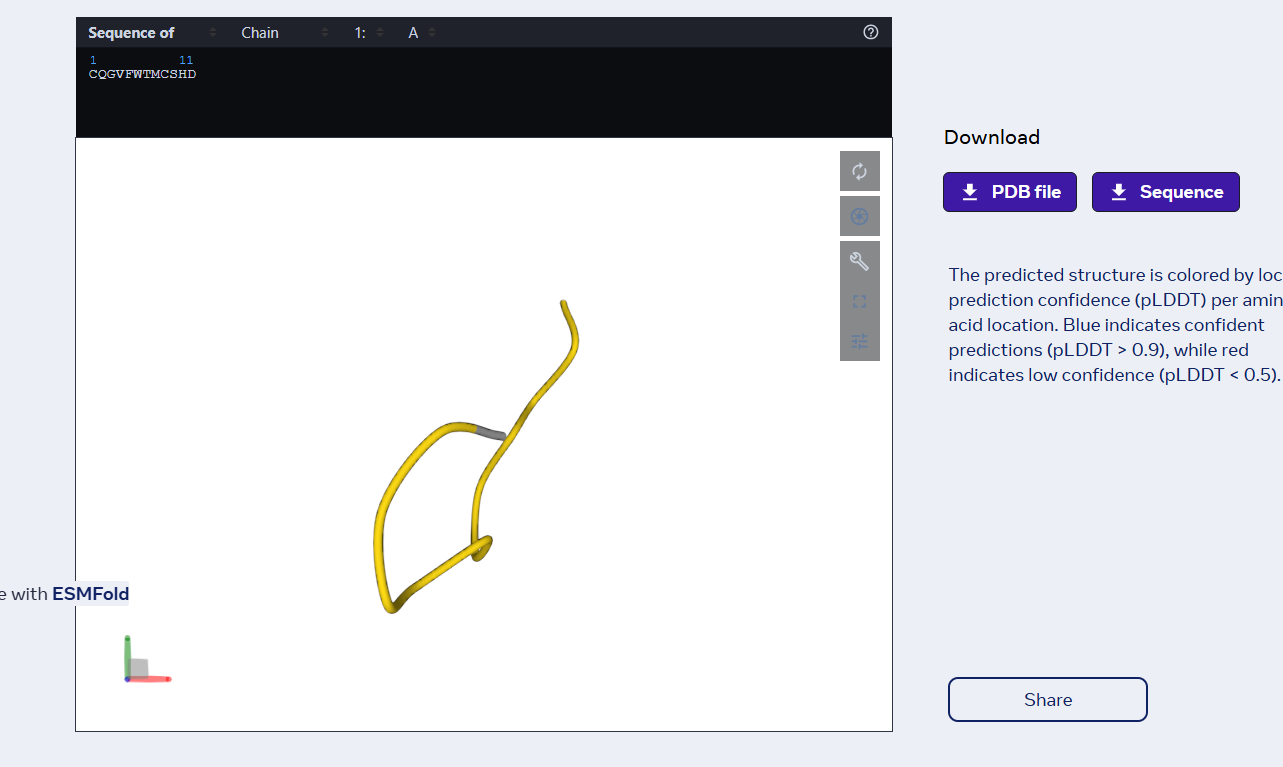

- the peptide has the same number of AAs as the previous ones
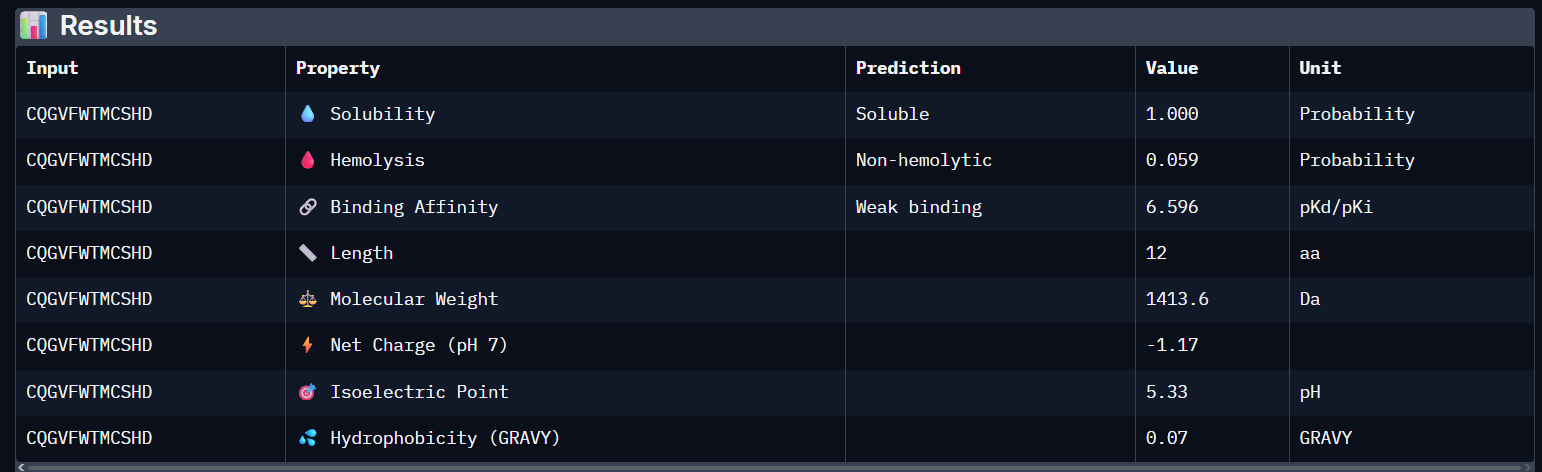

- it still has a weak binding affinity..
Part C:
I read the documentation resources about the MS2-L its similarity with the other amurines. List with a few important things for this project:
- MS2 is a bacteriophage that infects E.coli and has a genome containing only 4 proteins: the lysis protein (MS2-L/L-protein), the capsid protein, the maturation protein (initiates infection) and replicase;
- MS2-L is a protein found in the MS2 bacteriophage that produces the lysis of the infected bacteria using amuralytic ways, meaning that it doesn’t directly break down the peptoglycan wall (as the majority of bacteriophage lysis proteins do), but it most likely inserts into the cytoplasmatic membrane of E.coli;
- MS2-L is often compared with ΦX174 (also an amurine). The difference is that ΦX174 seems to alter the path of PG wall synthesis (inhibits the MraY enzyme), eventually leading to cell lysis.
- MS2-L is chaperone dependent. Without the DnaJ chaperone found in E.coli, the protein remain self-inhibited. The removal of the first domain of the lysis protein made the protein become chaperone-independent without altering the lysis function and increasing the speed of induced lysis with 20 minutes compared to the normal protein.
- the soluble domain of MS2-L –> first 40 AAs, the Transmembrane domain –> the last 35 AAs;
Then I ran the colab linked at option one of the protein engineering part and analyzed the results:
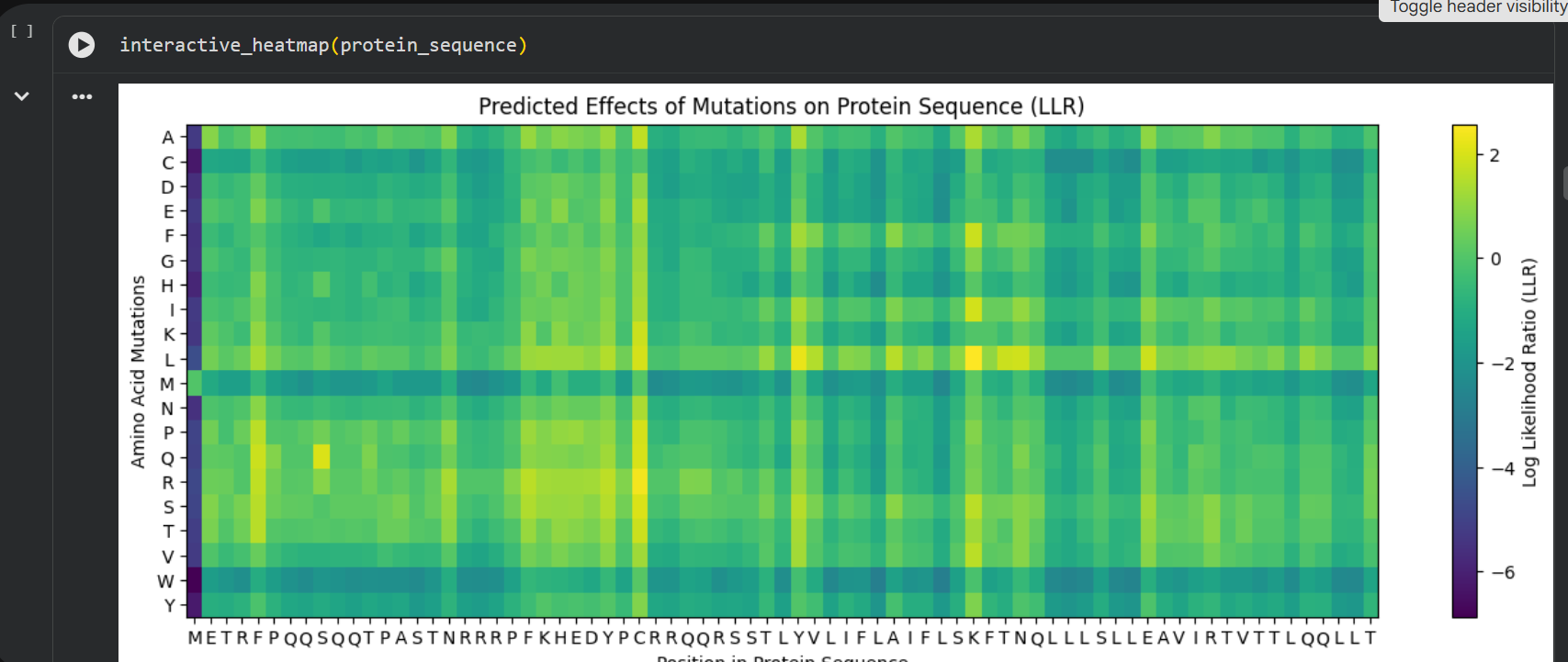

log likelihood ratio: < 0 - the original sequence is favoured; > 0 - the engineered protein is favoured The obvious observations that can be made by looking at the graph are:
the first resiude of the protein (M) is irreplaceable;
Mutations of a residue with: W - Tryptophan, M - Methionine, C - Cysteine , are not favoured
almost any mutation of the 29th residue (C) is favoured (Also generated using the colab code:) Position Wild_Type_AA Mutation_AA LLR_Score 989 50 K L 2.561464 574 29 C R 2.395427 769 39 Y L 2.241778 575 29 C S 2.043150 173 9 S Q 2.014323 573 29 C Q 1.997049 572 29 C P 1.971028 569 29 C L 1.960646 987 50 K I 1.928798 1049 53 N L 1.864930 1209 61 E L 1.818096 1029 52 T L 1.813965 984 50 K F 1.802066 576 29 C T 1.797247 568 29 C K 1.795877 93 5 F Q 1.795244 94 5 F R 1.659716 560 29 C A 1.648655 534 27 Y R 1.628060 434 22 F R 1.602028 92 5 F P 1.596889 997 50 K V 1.594573 995 50 K S 1.574555 96 5 F T 1.559023 95 5 F S 1.556416 889 45 A L 1.539248 775 39 Y S 1.517457 535 27 Y S 1.497052 789 40 V L 1.477630 529 27 Y L 1.474638 435 22 F S 1.423357 563 29 C E 1.383282 760 39 Y A 1.364997 571 29 C N 1.362601 980 50 K A 1.357792 567 29 C I 1.344121 89 5 F L 1.332615 334 17 N R 1.323652 767 39 Y I 1.320101 776 39 Y T 1.302803 514 26 D R 1.268762 566 29 C H 1.246106 764 39 Y F 1.245850 777 39 Y V 1.244389 454 23 K R 1.236555 494 25 E R 1.229349 474 24 H R 1.227778 996 50 K T 1.222128 533 27 Y Q 1.218850 536 27 Y T 1.215567
Choosing the most favoured mutations:
- I compared the most favoured mutations given by the colab with the results from the experimental data provided in the Excel and I chose a combination of the most favoured mutations to generate the final engineered MS2-L lysis protein:
- Mutations: S9Q, C29R, K50L Sequence: METRFPQQQQQTPASTNRRRPFKHEDYPRRRQQRSSTLYVLIFLAIFLSLFTNQLLLSLLEAVIRTVTTLQQLLT
- Mutations: S9Q, C29S, K50I Sequence: METRFPQQQQQTPASTNRRRPFKHEDYPSRRQQRSSTLYVLIFLAIFLSIFTNQLLLSLLEAVIRTVTTLQQLLT
- Mutations 3: S9Q, C29P, K50F Sequence: METRFPQQQQQTPASTNRRRPFKHEDYPPRRQQRSSTLYVLIFLAIFLSFFTNQLLLSLLEAVIRTVTTLQQLLT
- Mutation P13L Sequence: METRFPQQSQQTLASTNRRRPFKHEDYPCRRQQRSSTLYVLIFLAIFLSKFTNQLLLSLLEAVIRTVTTLQQLLT
- Mutation S15A Sequence: METRFPQQSQQTPAATNRRRPFKHEDYPCRRQQRSSTLYVLIFLAIFLSKFTNQLLLSLLEAVIRTVTTLQQLLT
- nr. of amino acids: 12
Genetic Circuits Part 1
- What are some components in the Phusion High-Fidelity PCR Master Mix and what is their purpose?
Components: Phusion DNA Polymerase (low error rate), nucleotides (A, T, C, G without U), reaction buffer (ensure maximum enzyme activity);
- What are some factors that determine primer annealing temperature during PCR?
Each primer has a different annealing temperature determined by the environment and by the nucleotides that make up the primer
- There are two methods from this class that create linear fragments of DNA: PCR, and restriction enzyme digests. Compare and contrast these two methods, both in terms of protocol as well as when one may be preferable to use over the other.
PCR makes strands by attaching primers with tails that hang freely. The result is a DNA sequence made of the original one + the tails of the primers that were added. Restriction enymes cut DNA and the resulted sequence part of the original sequence.
- How can you ensure that the DNA sequences that you have digested and PCR-ed will be appropriate for Gibson cloning?
The sequences need to have identical ends in order to be cut and ligated.
- How does the plasmid DNA enter the E. coli cells during transformation?
The E.coli is thermally or electrically stimulated to accept the vector by creating holes in the membrane. This often kills the bacteria..There are newer methods that use really small tubes to create holes that do not damage the bacteria as much as the two previous methods.
- Describe another assembly method in detail (such as Golden Gate Assembly)
- Explain the other method in 5 - 7 sentences plus diagrams (either handmade or online).
- Model this assembly method with Benchling or Asimov Kernel!
Golden Gate assembly is a highly efficient DNA cloning method. It uses type II enzymes that “cut” DNA consequent to recognized nucleotide sequence (usually palindromic), mostly after 4-5 nucleotides-post recognition site; and are dependent of Mg2+ as a cofactor. They act on both the plasmid and fragment pretty much the same; thus, the digested insertion fragment covalently binds to plasmid overhang complementary DNA strand.
Compared to Gibson a. it doesn’t require polymerases to build up lacking DNA strand fragments, but it does usually include ligase enzymes to ensure vector stability and successful annealing. G.G. assembly is preferred when working with many insertion fragments (30-50) because of its simultaneous activity of ligases and restriction enzymes.
Week 7 Homework
Part A:
What advantages do IANNs have over traditional genetic circuits, whose input/output behaviors are Boolean functions?
Intracellular artificial networks offer precise control over the output of an application, thus opening the gates for complex circuits where it is not only important that certain parameters exist, but also their quantity. Using IANNs it is possible to design a circuit whose output varies based on both the presence of x different molecules and their concentration.
Describe a useful application for an IANN; include a detailed description of input/output behavior, as well as any limitations an IANN might face to achieve your goal.
Creating a system that expresses 2 different protein based on the stimuli received by 3 other molecules. Input: molecule A (upragulates the expression of protein X & downregulates the expression of protein Y), molecule B upragulates the expression of protein Y & downregulates the expression of protein X, molecule C (upragulates the expression of protein X & upregulates the expression of protein Y - positive feedbak loop); Output: protein X, protein Y; Description of the circuit: The promoters of each gene are upregulated of downregulated by the 3 input molecules. We will consder that molecule A has an effect equal to the value 2 for upregulation and downregulation, molecule B has an effect of 1.5 on upregulation and 2.5 on downregulation and moelcule Chas an effect of 7.0 on upregulation. More molecule B willl be needed for the upregulation of molecule B when molecule A is present and more molecule A will be needed for upregulation in the presence of moelcule B, while moelcule C acts as a general upregulator that overwrites the other 2 molecules and amking sure that both genes will be expressed even though not in equal amounts.
Below is a diagram depicting an intracellular single-layer perceptron where the X1 input is DNA encoding for the Csy4 endoribonuclease and the X2 input is DNA encoding for a fluorescent protein output whose mRNA is regulated by Csy4. Tx: transcription; Tl: translation.
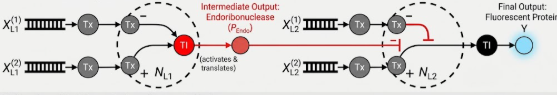

(Gemini Ai generated image by following a set of steps)
Part B:
What are some examples of existing fungal materials and what are they used for? What are their advantages and disadvantages over traditional counterparts?
Fungi have moved from the forest floor to the factory, particularly through the use of mycelium to create sustainable alternatives to traditional materials. Myco-leather offers an ethical substitute for animal hides, while mushroom packaging provides a biodegradable shield for electronics that competes with plastic foams. In architecture, myco-bricks utilize fungal growth to bond organic waste into fire-resistant structural elements. These biological materials are carbon-negative and non-toxic, providing a stark contrast to the high-polluting industries they aim to replace. However, the sheer scale of global demand remains a challenge, while millions of trees are felled every year to satisfy the timber and paper markets, the fungal industry is still scaling its production methods to reach that level of output. Furthermore, the porous nature of untreated mycelium means it currently lacks the inherent water-resistance and long-term durability found in chemically treated plastics or leathers, requiring further innovation in bio-coatings to match the lifespan of traditional goods.
What might you want to genetically engineer fungi to do and why? What are the advantages of doing synthetic biology in fungi as opposed to bacteria?
Designing strains for bioremediation that digest microplastics or sequester toxins from the soil, as well as infrastructure where fungal spores dormant in a wall grow to seal cracks autonomously when exposed to air. In medicine, the ability of fungi to produce complex human proteins could enable localized drug manufacturing in crisis zones; for instance, a portable bioreactor could synthesize a vital treatment for a patient before they are medevaced to a higher-level trauma center. Fungi hold a significant edge over bacteria in these tasks because, as eukaryotes, they possess specialized cellular machinery like the Golgi apparatus required for proper protein folding and modifications. They are also natural secretors, making it easier to harvest products compared to the destructive extraction methods often needed for bacteria. Research into these eukaryotic workhorses remains buoyed by the fact that they offer the structural strength of a multicellular network alongside the sophisticated chemical processing of higher life forms.
Homework Week 9
Part A
Explain the main advantages of cell-free protein synthesis over traditional in vivo methods, specifically in terms of flexibility and control over experimental variables. Name at least two cases where cell-free expression is more beneficial than cell production.
The main advantage of using cell free systems of in vivo methods are reduced system complexity. A system (not just in synbio) should achieve the desired output with a minimum level of complexity. Using a living organsism to express a gene or to trigger a response from another organism unnecessarily elevates the complexity by adding elements that do not serve the system purposea, thus increasing the risk of failure. Any element added to a system can potentially interact with other elements in unforeseen scenerios and cause an incorrect output.
Describe the main components of a cell-free expression system and explain the role of each component.
A cell-free expression (CFE) system functions by utilizing a cellular extract containing the essential machinery for translation. During preparation, the host cell’s membrane is felled to release ribosomes, tRNAs, and enzymes into a controlled environment. This extract is then supplied with a genetic template (DNA or mRNA) that provides the blueprints for the target protein.To maintain synthesis, the reaction is buoyed by an energy regeneration system using ATP, GTP, and secondary energy sources like phosphoenolpyruvate. Combined with a supply of amino acids and essential ions such as $Mg^{2+}$ and $K^{+}$, these components allow for rapid protein production without the constraints of cell viability. Because these systems can be freeze-dried, they are easily medevaced to remote regions for urgent diagnostic or medical applications.
- Why is energy provision regeneration critical in cell-free systems? Describe a method you could use to ensure continuous ATP supply in your cell-free experiment.
Energy regeneration is essential because protein synthesis consumes high-energy phosphate bonds rapidly. Without a regeneration system, the initial $ATP$ supply is quickly felled, leading to a premature halt in translation as $ADP$ and inorganic phosphate accumulate. To prevent this, the reaction must be buoyed by an enzymatic system that recycles nucleotides, ensuring the translational machinery has a constant fuel source for amino acid polymerization.One effective method involves supplementing the reaction with high-energy phosphate donors like creatine phosphate. When paired with the enzyme creatine kinase, this system transfers phosphate groups to $ADP$, replenishing $ATP$ levels in real time. For even longer durations, a continuous-exchange setup can be used; here, a dialysis membrane allows fresh nutrients to enter while metabolic inhibitors are medevaced into a separate buffer reservoir, sustaining the reaction for days.
- Compare prokaryotic versus eukaryotic cell-free expression systems. Choose a protein to produce in each system and explain why.
Prokaryotic and eukaryotic cell-free systems differ primarily in their complexity and post-translational capabilities. Prokaryotic systems, such as those derived from E. coli, are highly efficient and cost-effective, making them ideal for the rapid production of small, cytoplasmic proteins. In this system, the cellular extract is felled from its original source to provide high-speed ribosomal activity. A suitable protein for this system would be Green Fluorescent Protein (GFP), as it requires no complex modifications and serves as a robust reporter. Conversely, eukaryotic systems like wheat germ or rabbit reticulocyte lysates are buoyed by more sophisticated folding machinery and are better suited for human proteins like Insulin. These systems can handle the specific disulfide bond formations and glycosylation patterns necessary for therapeutic proteins to function correctly, though they often yield lower total quantities than their bacterial counterparts.
- How would you design a cell-free experiment to optimize the expression of a membrane protein? Discuss the challenges and how you would address them in your setup.
Designing a cell-free experiment for membrane proteins requires addressing the inherent hydrophobicity of the target, which often leads to misfolding or aggregation. To optimize expression, the reaction must be supplemented with a hydrophobic environment, such as detergents, liposomes, or nanodiscs, to provide a scaffold for the protein’s transmembrane domains. The primary challenge is that membrane proteins often remain stuck in the extract’s machinery or precipitate out of solution; if the folding environment is insufficient, the protein’s functional potential is essentially felled before it can be characterized. My setup would utilize a “co-translational” approach where nanodiscs are present from the start, allowing the nascent protein to insert directly into a lipid bilayer as it exits the ribosome, thereby mimicking the natural cellular environment and preventing the formation of insoluble aggregates.
- Imagine you observe a low yield of your target protein in a cell-free system. Describe three possible reasons for this and suggest a troubleshooting strategy for each.
When troubleshooting a low protein yield, one must first investigate template degradation, as endogenous nucleases in the extract can destroy the genetic blueprint. To address this, I would incorporate nuclease inhibitors or use circular plasmid DNA, which is less susceptible to being felled by exonucleases compared to linear PCR products. A second possibility is energy depletion, where the reaction stalls because the $ATP$ supply is exhausted; here, the strategy involves optimizing the energy regeneration system or using a continuous-flow setup to ensure the machinery is constantly supplied with fresh fuel. Finally, the issue could stem from toxic byproduct accumulation, such as inorganic phosphate or local pH shifts. These inhibitory metabolites can be medevaced from the active reaction site through dialysis, allowing the synthesis to proceed in a cleaner, more favorable environment for an extended period.
Homework question from Kate Adamala
- The Frontier of Synthetic Life: Designing the Minimal Bioremediation CellThe quest to define the absolute requirements for life has transitioned from a philosophical inquiry to a precise engineering discipline. Through the development of Synthetic Minimal Cells (SMCs), we are no longer limited to modifying what exists; we are beginning to build functional biological units from the bottom up. One of the most promising applications for this technology lies in environmental nanomedicine—specifically, the creation of “sentinel” cells designed to detect and neutralize heavy metal toxins in aquatic ecosystems.I. The Architecture of a Synthetic SentinelA useful synthetic minimal cell must be more than a simple container; it must be a responsive machine. Our proposed design focuses on a Mercury-Sensing and Sequestration Cell. The “logic” of the cell is straightforward:Input: Divalent mercury ions ($Hg^{2+}$) permeating the lipid bilayer.Processing: A genetic circuit that triggers protein synthesis only in the presence of the metal.Output: The production of Metallothionein, a protein that binds and “sponges” the mercury, and Luciferase, which provides a bioluminescent signal.Unlike a standard cell-free system, encapsulation is a mechanical necessity here. Without a membrane, the sequestered toxins would remain in the bulk environment, and the delicate transcriptional machinery would be quickly felled by environmental proteases or chemical degradation.II. Engineering Constraints and ComponentsThe design of an SMC requires a careful selection of both hardware (lipids) and software (genes). For this environmental sentinel, the following specifications are required:The Membrane and ChassisThe membrane must be robust enough to withstand varying water temperatures while remaining permeable to small ions. A blend of POPC and DPPC phospholipids, stabilized with Cholesterol, creates a “stealth” exterior. The internal environment is powered by a bacterial Tx/Tl system (derived from E. coli), which is ideal because the mercury-responsive operons ($merR$) are natively optimized for bacterial machinery.The Genetic CircuitryTo achieve the desired outcome—a visible reduction in toxicity and a user-readable signal—we incorporate a specific genetic toolkit:merR & PmerT: The sensor-promoter complex that acts as the “on-switch.“bmtA (Metallothionein): The functional “sponge” that traps mercury.hlyA ($\alpha$-Hemolysin): A pore-forming protein that allows the cell to communicate its status to the environment.III. Measurement and OutcomesSuccess in synthetic biology is defined by measurable function. In a laboratory setting, the efficiency of this SMC is quantified through two primary metrics:Optical Output: The intensity of bioluminescence relative to mercury concentration.Sequestration Efficiency: Using ICP-MS to verify that the concentration of free mercury in the sample has been significantly lowered.IV. Conclusion: The Safety of Non-Living SolutionsWhile a genetically modified natural bacterium could perform similar tasks, the SMC offers a critical safety advantage: containment. Because these cells lack the genes for replication and metabolism, they cannot evolve or establish an invasive population. They are discrete tools that perform a job and then degrade. In scenarios where fragile ecosystems are threatened by industrial runoff, these synthetic units could be deployed to high-risk areas, ensuring that the health of the local flora and fauna is buoyed by precise, controlled intervention. Should an environmental crisis escalate, these bio-nanobots could be deployed rapidly—effectively medevaced into toxic zones to perform tasks where living organisms would perish.
Homework Questions from Peter Nguyen:
The RAD-Tex Biosensing Fabric is a textile-integrated BioBits® sensor array that provides real-time, visual fluorescence feedback to astronauts when exposed to critical levels of ionizing radiation or localized DNA-damaging environmental toxins.
How it Works The system utilizes BioBits® freeze-dried protein synthesis machinery (ribosomes, RNA polymerase, and energy sources) embedded within the hollow-core fibers of a spacesuit’s outer layer. Upon a high-energy radiation event or a chemical trigger, a microfluidic capillary system within the fabric releases a rehydration buffer. This “wakes up” the FDCF system, which immediately begins transcribing and translating a specific DNA template into Green Fluorescent Protein (GFP). The resulting glow is clearly visible through the P51 Molecular Fluorescence Viewer, effectively turning the suit into a biological dashboard.
Societal Challenge & Market Need Long-duration spaceflight exposes crews to invisible but lethal cosmic radiation and toxic lunar/Martian regolith. Current electronic sensors can fail or lack the granularity to show where a specific “leak” or exposure occurred on an astronaut’s body; this biological sensor provides localized, intuitive evidence of environmental threats.
Addressing Limitations Stability: The lyophilized (freeze-dried) state allows the BioBits® components to remain shelf-stable for months without refrigeration, which is essential for deep-space transit.
Activation: We use a “burst-valve” microfluidic system that only introduces water/nutrients when a certain threshold of environmental stress is detected or when the user manually initiates a check.
One-time Use: To mitigate this, the fabric is designed with modular, “rip-and-replace” patches. Once a patch has been “activated” and used, it can be swapped for a fresh one during habitat maintenance.
Homework question from Ally Huang:
Cosmic radiation and lunar dust toxicity pose existential threats to long-term lunar habitation. Unlike Earth, where the atmosphere protects us, astronauts are vulnerable to DNA-damaging particles that can cause rapid cellular degradation. If a crew member is felled by acute radiation syndrome during an EVA, the mission’s success is compromised. Because astronauts cannot be easily medevaced from the lunar surface to a terrestrial hospital, we need immediate, localized biological monitoring to detect environmental threats before physiological symptoms manifest. (86 words)
Molecular/Genetic Target The molecular target is the recA promoter fused to a Green Fluorescent Protein (GFP) reporter gene, designed to respond to DNA damage within the cell-free system. (28 words)
Relevance to Space Biology The recA promoter is a classic SOS response element. In this system, ionizing radiation causes DNA fragmentation within the FDCF matrix. As the system attempts to respond to the damage, the recA promoter triggers the BioBits® machinery to express GFP. This provides a direct, biological proxy for the radiation dose the astronaut is receiving. By correlating fluorescence intensity with damage, we can monitor the “biological age” or integrity of the suit’s shielding and the astronaut’s immediate exposure level in real-time. (93 words)
Hypothesis & Reasoning We hypothesize that FDCF BioBits® systems can be lyophilized into textile fibers and remain functional in a space environment to serve as a high-sensitivity radiation biosensor. We reason that since cell-free systems lack a cell wall, they allow for faster interaction between environmental stressors and the genetic reporter than traditional whole-cell biosensors. This immediacy, buoyed by the P51 viewer’s ability to detect low-level fluorescence, will allow for a “canary in the coal mine” system for deep-space missions, providing a reliable, low-mass safety layer for extravehicular activities. (104 words)
Experimental Plan We will test three samples: FDCF-embedded fabric exposed to varying doses of ionizing radiation, a non-exposed negative control, and a positive control activated by a chemical inducer. After exposure, samples will be rehydrated with the BioBits® activation buffer. We will measure the fluorescence output using the P51 Molecular Fluorescence Viewer at 15-minute intervals. The miniPCR® will be used to maintain a consistent incubation temperature of 37°C. Success is defined as a statistically significant correlation between radiation dosage and GFP fluorescence intensity compared to the non-irradiated control.
Advanced Imaging & Measurement Technology
Homework: Final Project
Please identify at least one (ideally many) aspect(s) of your project that you will measure. It could be the mass or sequence of a protein, the presence, absence, or quantity of a biomarker, etc. Please describe all of the elements you would like to measure, and furthermore describe how you will perform these measurements. What are the technologies you will use (e.g., gel electrophoresis, DNA sequencing, mass spectrometry, etc.)? Describe in detail.
- Sanger Sequencing will be utilized to confirm the primary nucleotide sequence of the plasmids. Quantitative Real-Time PCR (qPCR) will be employed to measure mRNA expression levels, ensuring that the genetic instructions are being transcribed at the predicted rates within the cell-free environment.
- Technology: Cryo-Electron Microscopy (Cryo-EM) will be used to visualize the morphology and assembly of the bacterial microcompartments. Additionally, Dynamic Light Scattering (DLS) will measure the hydrodynamic radius and polydispersity index to ensure a uniform population of nanocages, while Liquid Chromatography-Mass Spectrometry an be used to quantify the ratio of encapsulated versus non-encapsulated proteins.
- Technology: Fluorimetry and Surface Plasmon Resonance (SPR) will be employed to measure the binding affinity of the activation protein to the circuit’s sensor. By using fluorescent reporter proteins for the therapeutic payload during testing, the “transfer function” or the logic curve of the IANN can be mapped to ensure that the dose-response remains calibrated and autonomous.
- Technology: Flow Cytometry will be utilized to assess cell viability and the activation state of surrounding immune cells (T cells and NK cells). The LDH (Lactate Dehydrogenase) Release Assay will serve as a primary metric for measuring the killing capacity (cytotoxicity) of the engineered system when exposed to cancer cell lines in vitro.
Homework: Waters Part I — Molecular Weight
- Calculated Theoretical Molecular WeightBased on the provided 246 amino acid sequence –> MW: ~27,949.2 Da.
- Experimental Molecular Weight Calculation: 27,981.9 Da
- Accuracy: 0.117%
- z ~= 23
Homework: Waters Part II — Secondary/Tertiary structure
- Native vs. Denatured Protein ConformationsThe primary difference between native and denatured protein states in mass spectrometry lies in the surface area and solvent accessibility of the molecule.Native State (Folded): In its natural environment, a protein like eGFP is tightly folded into a compact 3D structure. Most of its basic amino acid residues (like Lysine and Arginine), which accept protons during Electrospray Ionization (ESI), are buried within the protein’s core. Consequently, the protein can only pick up a few charges. This results in:Higher $m/z$ values: Because the charge is low, the mass-to-charge ratio is high.Narrow Distribution: Only a few charge states (peaks) are visible because the folded structure is rigid.Denatured State (Unfolded): When a protein is denatured, it unfolds into a “random coil” or extended string. This exposes nearly all basic sites to the solvent, allowing for significant protonation .Broad Distribution: A wide “envelope” of many charge states is observed because the unfolded protein can adopt various degrees of protonation.
## Homework: Waters Part III
- Theoretical Molecular Weight CalculationBased on the provided eGFP sequence (247 amino acids including the LE linker and HHHHHH His-tag):Amino Acid Counts: 20 Lysines (K) and 6 Arginines (R).Average Molecular Weight: Approximately $28,006.3$ Da (unmodified).Matured eGFP (with chromophore): Accounting for the $-20.03$ Da loss during chromophore formation (T66-Y67-G68), the corrected theoretical weight is approximately $27,986.3$ Da.
- MW = 27,981.9 Da
- 157ppm
- Charg =e state ~= 19
- Cleavage Sites: 20 Lysines (K) + 6 Arginines (R) = 26 sites.Predicted Peptides: Assuming no missed cleavages, trypsin will generate 27 peptides.Observed Chromatographic Peaks: Between 0.5 and 6 minutes, there are 18 peaks with $>10%$ relative abundance (labeled peaks at 0.43, 0.61, 1.43, 1.80, 1.85, 1.93, 2.17, 2.26, 2.54, 2.78, 3.53, 3.59, 3.70, 4.30, 4.48, 4.64, 4.87, and 5.06).
- To identify the peptide eluting at 2.78 minutes, the experimental data from Figure 5b is compared against the theoretical tryptic peptides predicted by the PeptideMass tool. The observed peak has a mass-to-charge ratio of 525.76712. By examining the isotopic spacing in the zoom-in inset, where the difference between peaks is approximately $0.492$ Da, the charge state is determined to be $z = 2$. Using the formula MW = z* (m/z - 1.0078) + 1.0078, the singly charged mass is calculated to be 1050.52644 Da. This matches the theoretical mass of the eGFP tryptic peptide FEGDTLVNR ($1050.514$ Da). The mass accuracy of this measurement is approximately $11.8$ ppm
Homework Part IV:
To find these species, multiply the subunit weight by the number of units and match the total to the labels on the graph’s horizontal axis.
7FU Decamer: This matches the peak labeled 3.4 Megadaltons.
8FU Didecamer: This aligns with the tallest peak on the spectrum, labeled 8.33 Megadaltons.
8FU 3-Decamer: This corresponds to the peak at 12.67 Megadaltons.
8FU 4-Decamer: This is represented by the small cluster of activity visible around the 17 Megadaltons mark.
The reason the experimental peaks (like 8.33) are slightly higher than the calculated weights (like 8.0) is due to “extra” mass from attached sugars or small linker proteins common in these large biological structures.