Week 2 Homework: DNA Read, Write & Edit
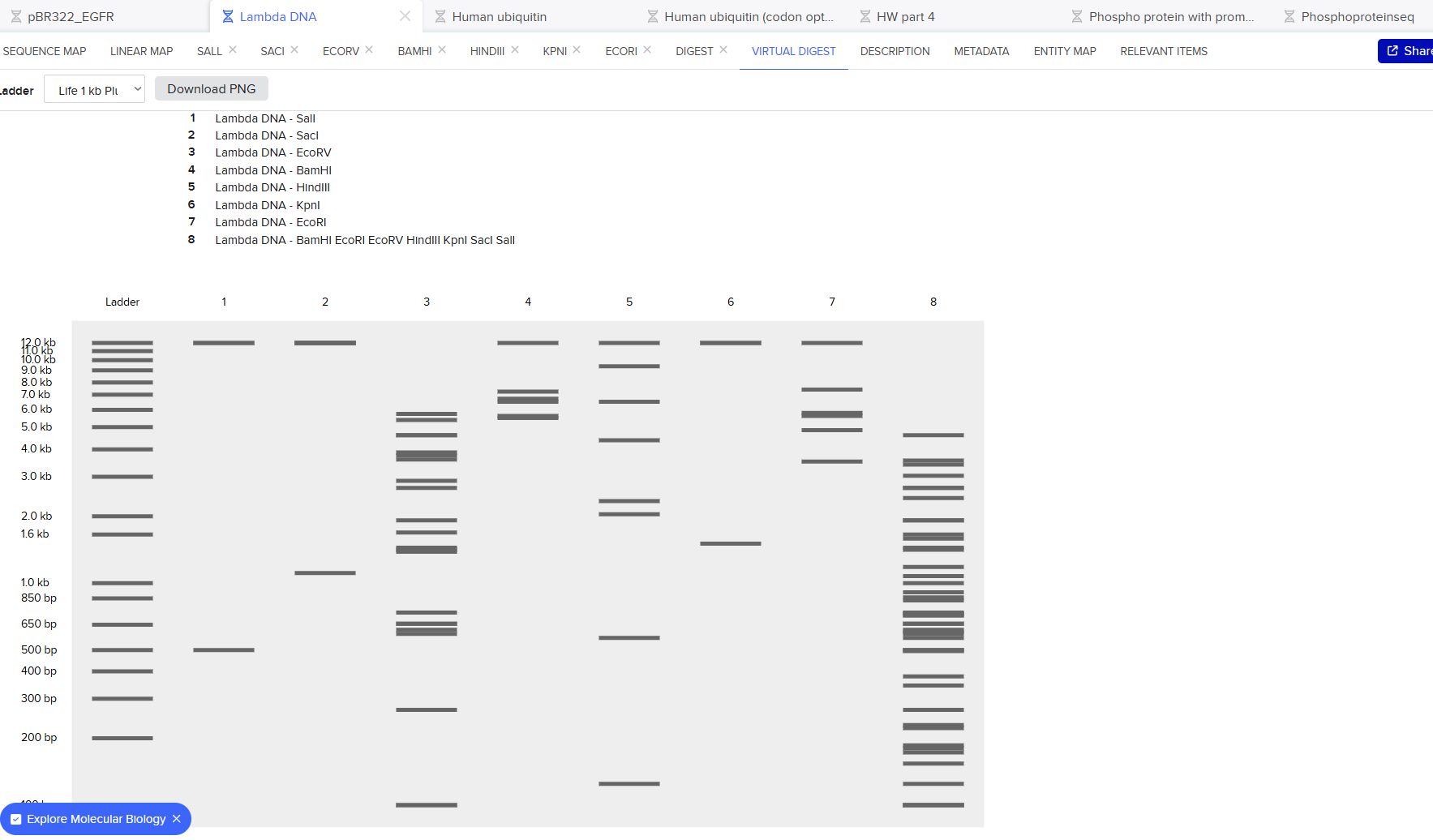
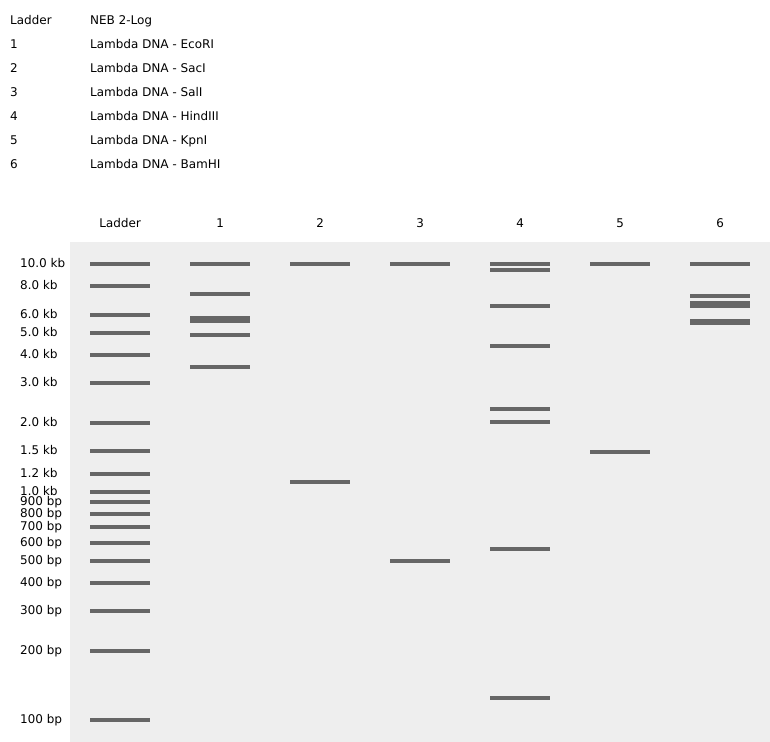
1. Gel Art:


vs


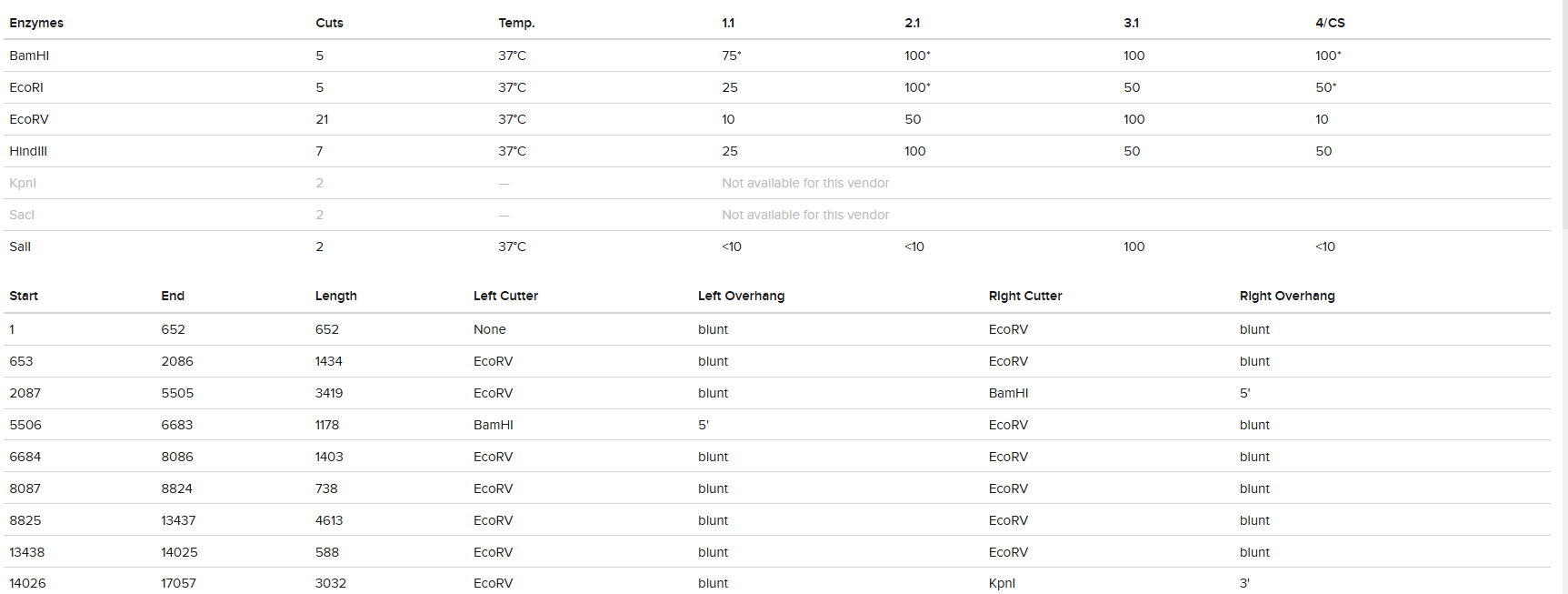
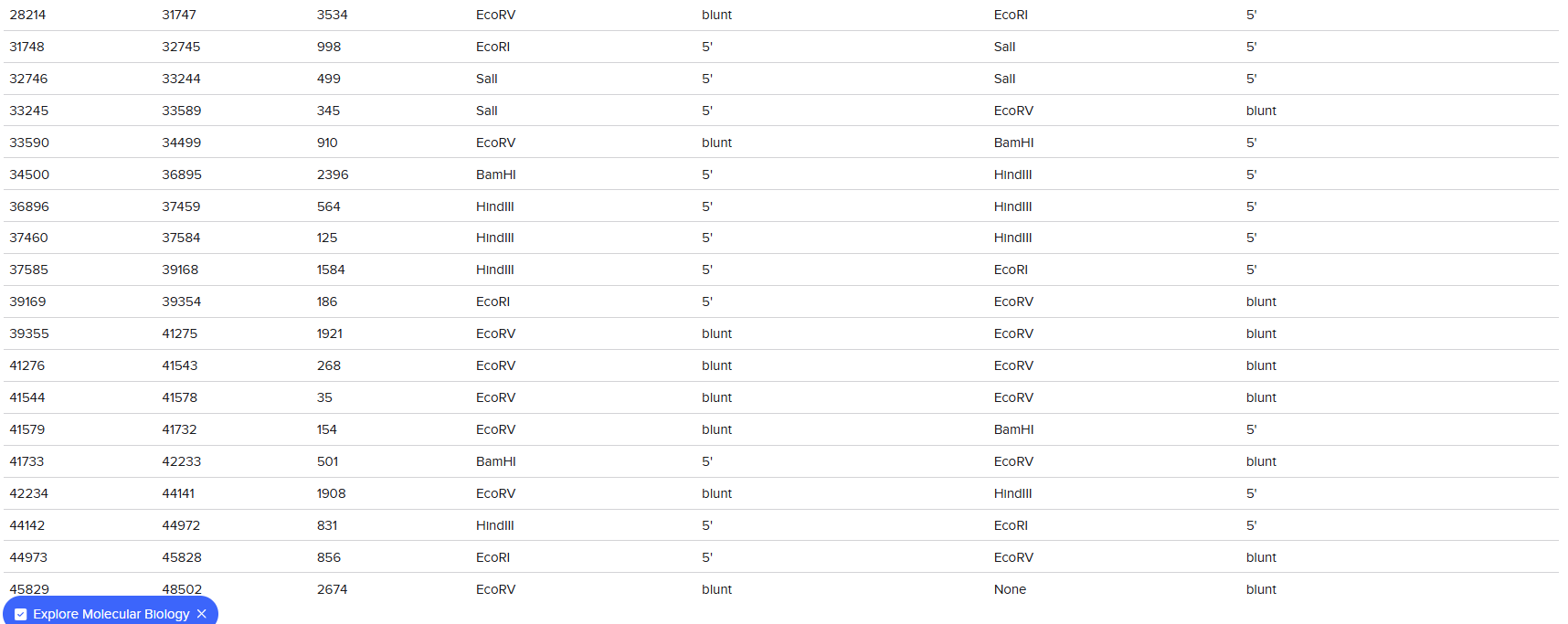
The digestion results:
3.1. Choose your protein.
Human ubiquitin: one of the most highly conserved proteins across eukaryotic species, with an identical amino acid sequence in organisms ranging from yeast to humans. It marks misfolded proteins for degradation. Protein sequence - 76 amino acids sp|P0CG48|UBC_HUMAN Polyubiquitin-C OS=Homo sapiens OX=9606 GN=UBC PE=1 SV=2 MQIFVKTLTGKTITLEVEPSDTIENVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLSDYNIQKESTLHLVLRLRGG
3.2. Reverse Translation:
ATGCAGATCTTCGTGAAGACCCTGACCGGCAAGACCATCACTCTGGAGGTGGAGCCCAGTGACACCATCGAGAACGTGAAGGCCAAGATCCAGGACAAGGAGGGCATCCCTCCTGACCAGCAGAGGCTGATCTTTGCCGGCAAGCAGCTGGAAGATGGCCGCACCCTGTCTGACTACAACATCCAGAAGGAGTCCACCCTGCACCTGGTGCTGCGTCTGCGTGGCGGC
3.3. Codon optimization:
Different organisms have distinct preferences for which codons they use for each amino acid, due to differences in tRNA abundance and translation efficiency. The native human sequence may contain codons that are rare in other hosts, leading to slower translation and ribosomal stalling, which increase the risk of misfolding. I decided to optimize for Escherichia coli K-12, which is fast-growing, inexpensive, well-characterized, and widely used for high-yield protein expression in research and industry. Using Benchling codon optimization tool, I obtained:
ATGCAGATCTTTGTGAAAACCCTGACCGGTAAAACCATTACCCTGGAAGTGGAGCCGAGCGATACCATTGAAAACGTGAAAGCGAAAATTCAGGATAAAGAAGGCATTCCGCCGGATCAGCAGCGCCTGATTTTTGCCGGCAAACAGCTGGAAGATGGTCGTACCCTGAGCGACTATAACATTCAGAAAGAAAGCACCTTACATCTGGTGCTGCGTCTGCGTGGTGGT
3.4. You have a sequence! Now what?
The optimized DNA sequence can be synthesized and cloned into a vector for protein production in E.coli K-12:
The DNA is inserted into a plasmid and transported into E. coli cells. Host RNA polymerase transcribes the DNA into mRNA. Ribosomes then translate the mRNA into the polypeptide chain using tRNAs, amino acids, and energy from GTP/ATP. The protein folds , and cells are lysed to purify it.
3.5. How does it work in nature/biological systems?
How a single gene codes for multiple proteins at the transcriptional level: One common mechanism is splicing, where a single pre-mRNA transcript is processed in different ways to include or exclude exons, producing multiple mature mRNA isoforms from the same gene. These isoforms are translated into distinct proteins with different functions. (Ubiquitin genes themselves primarily produce identical monomers via polyubiquitin precursors or fusions, with cleavage occurring post-translationally; alternative splicing is more prominent in other human genes, e.g., for diversity in receptors or enzymes.) Alignment of DNA, transcribed RNA, and translated Protein:
Important:
- in the genome there are multiple genes that code for ubiquitin (ribosomal fusion genes and polyubiquitin genes), due to the necessity of upregulating the production during periods of increased metabolic stress;
- in general, ubiquitin genes do not contain introns, most likely due to the need of quick production and low change for errors to occur.
- ubiquitin suffers post translation modifications
- ubiquitination is the process by which E1, E2 and E3 enzymes mark misfolded proteins or danaturated ones for protein degradation by attaching the damaged protein to ubiquitin as a substrate
- E1 activates ubiquitin
- E2 transports ubiquitin to the target protein
- E3 facilitates ubiquity-substrate binding
4.2
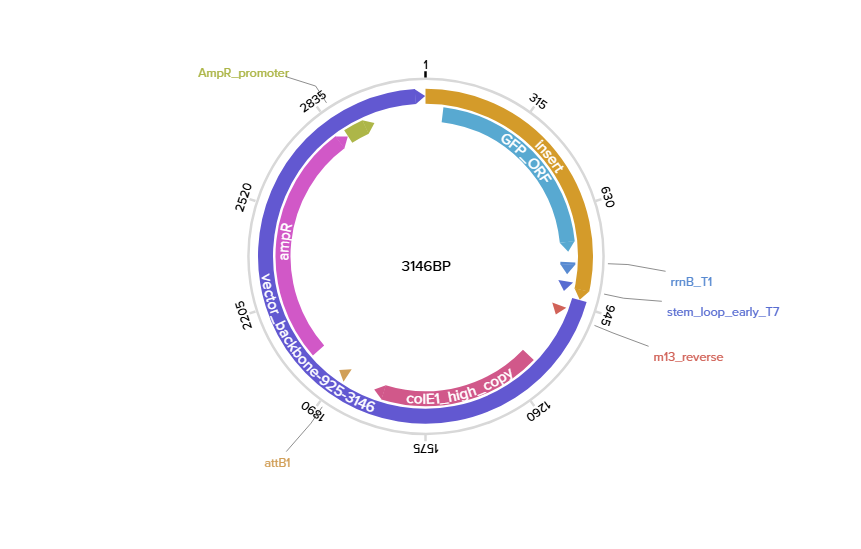
Complete sequence (promoter, RBS, Start codon, coding sequence, 7xHis Tag, end codon, terminator)
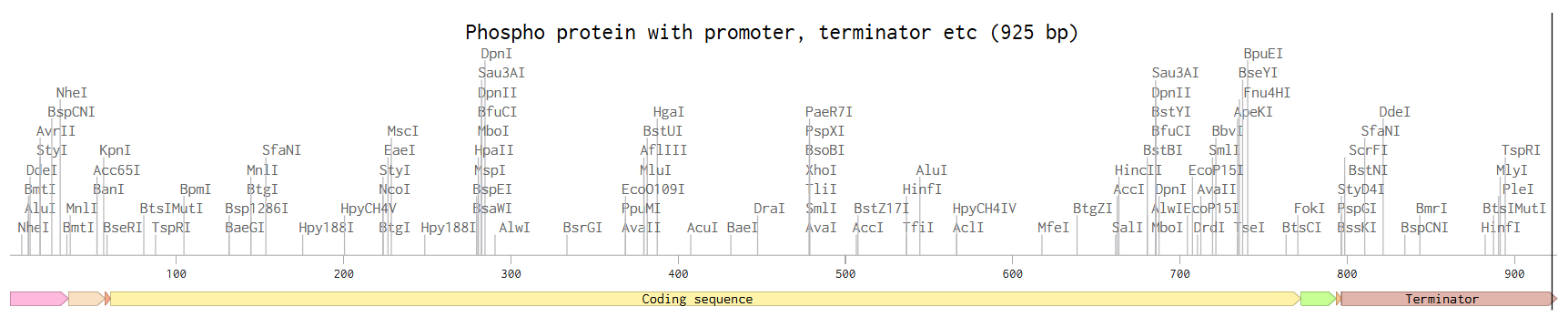


4.3, 4.4, 4,5, 4,6 Results:
5.1 What DNA would you want to sequence (e.g., read) and why? DNA READ
i)I would be enthralled to sequence the genes responsible for ribonucleic vaults. A more profound understanding of how the MVPs (major vault proteins) create literal containers used for cellular transportation would help us better understand how to create intricate structures far more complex than what evolution could achieve in the coming millions of years.
ii) I would use Third-Generation sequencing. Why? Unlike second-generation, which requires plenty of repetitions on amplified clusters, third generation sequencing performs single-molecule, real-time sequencing, hence being a lot more efficient. & It does not require PCR amplification, meaning it bypasses the cycle of amplification that can lead to errors.
5.2 DNA Write:
i) A genetic circuit designed for microplastic detection that pollutes aquatic environments.
The sheer volume of plastic entering oceans has created a vicious cycle where macroplastic breaks down into microplastic that ends up in our food and has toxicological effects on our health. The design would be a genetic circuit that responds to Terephthalic Acid (TPA), a primary product of PET plastic degradation.
- The Sensor: A specific transcription factor (TpaR) that remains inactive until it binds to TPA.
- The Reporter: Once TpaR is activated, it triggers the expression of a protein that can be seen with ease.
ii)
- Synthesis Technology: Silicon-based Synthesis I would choose this technology because it uses a silicon platform to act as a solid support for the chemical reactions. Traditional plastic plates are inefficient; by moving to silicon, we can rebuke the wastefulness of older methods, reducing reagent use by over 99%.
- Scalability: It can synthesize 9,600 genes on a single chip Length Constraints: The longer the DNA strand is, the more the efficiency of adding new bases drops. It would also be preferred to integrate a skill switch for keeping the bacteria population under control.
5.3 DNA Edit:
(i) The Edit: Enabling Nitrogen Fixation in Cereals Currently, only legumes can naturally convert atmospheric nitrogen into a usable form via a secretive symbiotic relationship with specialized bacteria. Most cereal crops lack this ability, forcing farmers into a vicious cycle of applying synthetic nitrogen fertilizers. The planet’s soil and waterways bear the brunt of this practice, leading to massive nutrient runoff and greenhouse gas emissions.
I would want to edit the Symbiosis Receptor-Like Kinase (SYMRK) and Nod Factor Receptor (NFR) genes in wheat. These edits would “re-tune” the plant’s root receptors to recognize the signaling molecules of nitrogen-fixing bacteria.
(ii) The Technology: Prime Editing To achieve these precise changes without causing accidental damage to the rest of the genome, I would use Prime Editing (PE).
While standard CRISPR-Cas9 is effective for cutting genes, Prime Editing is a search-and-replace tool that can rewrite specific DNA letters with manifold precision.
How it Edits DNA (simplified): Targeting: The Prime Editor is a protein consisting of a Cas9 nickase and a Reverse Transcriptase is guided to the specific root receptor gene. Nicking: Instead of cutting both strands of DNA, it creates a small nick in only one strand. Reverse Transcription: The pegRNA (prime editing guide RNA) provides a template. The RT enzyme reads this RNA and synthesizes a new strand of DNA that contains the desired edit. Flap Competition: The newly synthesized DNA flap competes with the original unedited DNA flap.
Incorporation: Through natural cellular repair, the old flap is removed, and the new, edited sequence is permanently integrated into the genome.
Inputs: The Prime Editor Construct: Usually delivered as a plasmid or mRNA encoding the Cas9-RT fusion protein. The pegRNA: The custom RNA sequence that searches and replaces. Delivery System: For plants, we often use Agrobacterium-mediated transformation to shoot these components into the plant tissue. Target Cells: Embryogenic callus cells from the wheat plant, which can be grown back into a whole, fertile plant. Limitations: Efficiency and Precision Although Prime Editing is highly precise, it is not yet perfect: Efficiency: In many plant species, the editing efficiency is still quite low (often <10%). Getting the “search-and-replace” to actually stick across millions of cells remains a bottleneck. Size Constraints: Because the Prime Editor protein is so large, it is difficult to deliver it. Target Range: The editor must be near a specific sequence called a PAM site. If the receptor gene we want to edit isn’t near a PAM, the tool is effectively blind to that location.