Week 1 HW: Principles and Practices
Class Assignment


Figure 1: The medicine ward in a typical government Indian hospital. The one shown is the medicine ward at my medical college.
First, describe a biological engineering application or tool you want to develop and why. This could be inspired by an idea for your HTGAA class project and/or something for which you are already doing in your research, or something you are just curious about.
Although I started off with imparting neuron-like properties in mycelial networks I decided to pivot to solving nosocomial infections with synthetic biology as the immediate benefits outweigh that of the former. In the wards , especially in under-resourced settings where I am from, maintenance of sterility and aseptic conditions in the post surgery wards and medicine wards is an under addressed issue. Under staffed , under resourced settings like mine, do not prioritise it leading to hospital acquired infections in the patients, often resulting in severe complications than which they came to the hospital for. With living colonisers that produce bactericidal and bacteriostatic chemicals (including fungicidal and antiviral) that generally cause secondary infections and HAIs(hospital acquired infections),synthetic biology has the scope to solve this problem saving over 100,000 lives in India alone, where people die of these HAIs. Low maintenance systems like these could change the way a lot of people are recovering during treatment in the wards. I would like to explore and build something tangible with Bacillus subtilis as the chassis organism , with quorum sensing to produce antimicrobial peptides against the priority pathogens causing hospital acquired infections
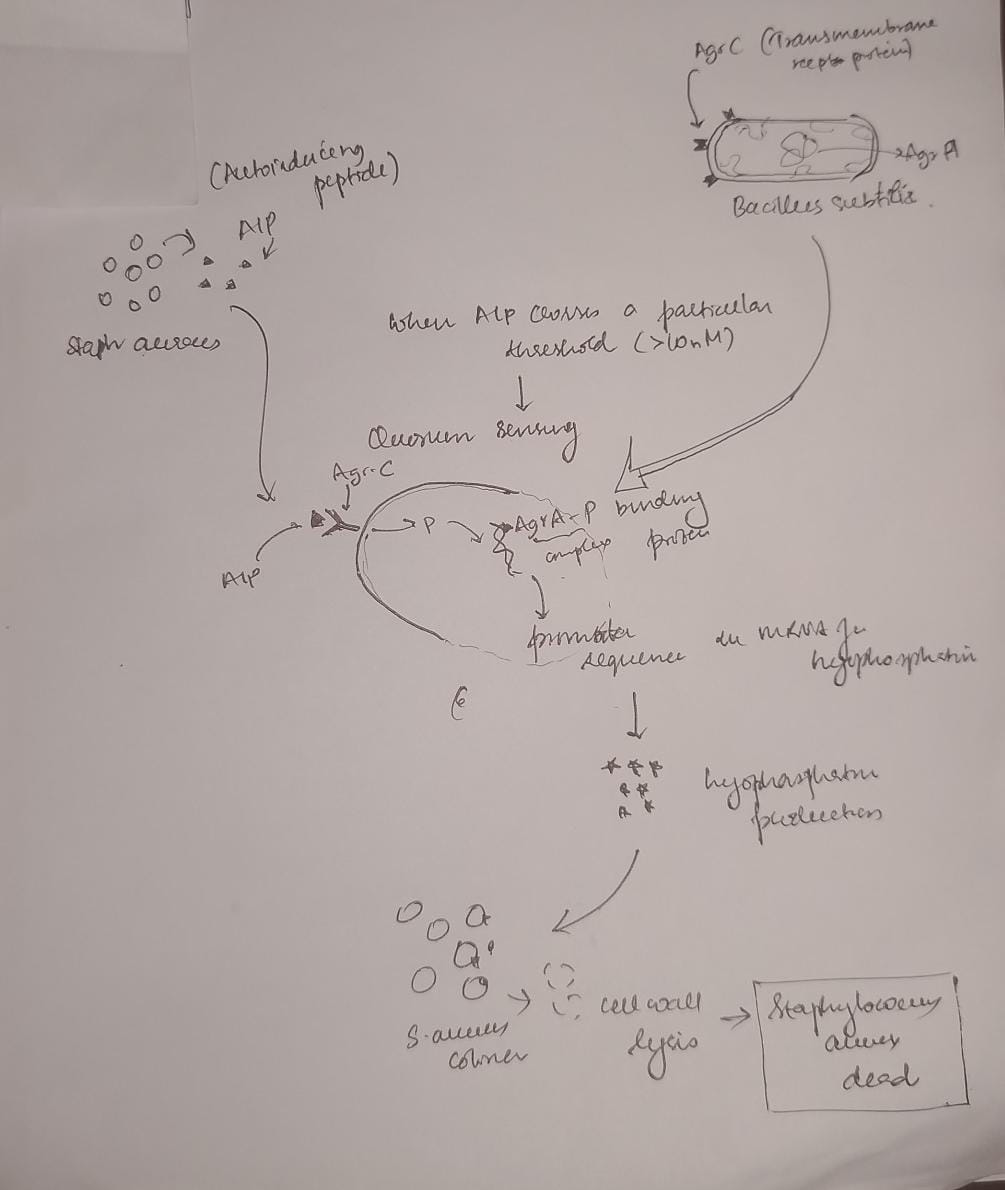

Figure 2: A diagram showing an example of the seek and kill mechanism in the engineered living surfaces. Here Bacillus subtilis acts as the chasis organism carrying the gene for Lysostaphin which is activated only on concentration of AIP produced by S. aureus crossing a certain threshold. This comprises of the sensor (Agr-C transmembrane protein in the genetically engineered B. subtilis), the logic gate(quorum) and the effector module (Lysostaphin produced by mRNA for the same activated by Agr-P protein)
Next, describe one or more governance/policy goals related to ensuring that this application or tool contributes to an “ethical” future, like ensuring non-malfeasance (preventing harm). Break big goals down into two or more specific sub-goals. Below is one example framework (developed in the context of synthetic genomics) you can choose to use or adapt, or you can develop your own. The example was developed to consider policy goals of ensuring safety and security, alongside other goals, like promoting constructive uses, but you could propose other goals for example, those relating to equity or autonomy.
Goal 1: Biosafety and Containment
- Prevent uncontrolled spread of engineered colonizers beyond designated hospital surfaces.
- Prevent horizontal gene transfer of engineered antimicrobial genes to environmental organisms.
- Prevent environmental contamination by these engineered organisms.
Goal 2: Patient Safety
- Ensure the colonizer produces no toxins or allergens harmful to immunocompromised patients.
- Ensure colonizers cannot cause opportunistic infections in vulnerable populations.
Goal 3: Equitable Access
- Keep unit cost below existing chemical disinfectant protocols.
- Ensure technology is deployable without specialized infrastructure (cold chain, trained personnel).
- Open-source genetic constructs and protocols to prevent IP monopolization.
Goal 4: Environmental Safety
- Prevent the colonizer from establishing in non-target environments (soil, water systems, community surfaces outside hospitals).
- Ensure engineered antimicrobial peptide genes do not contribute to resistance development in wild microbial populations.
Next, describe at least three different potential governance “actions” by considering the four aspects below (Purpose, Design, Assumptions, Risks of Failure & “Success”
Governance Actions
Action 1 - Regulatory: Establishing a Regulatory Authority for Engineered Living Surfaces in Healthcare
- Purpose: Currently, India’s Genetic Engineering Appraisal Committee (GEAC) regulates environmental release of GMOs under the Environment Protection Act 1986, but no framework specifically addresses engineered microorganisms deployed on surfaces within healthcare infrastructure. Hospital surfaces occupy a grey zone-they are neither open environmental release nor contained laboratory use-and no existing regulatory body in India is equipped to evaluate the safety, efficacy, or deployment protocols of living antimicrobial surface systems in clinical settings.
- Design: I propose that ICMR (Indian Council of Medical Research) and CDSCO (Central Drugs Standard Control Organisation), along with GEAC, establish a joint Advisory Board specifically to evaluate proposals for engineered microorganisms intended for deployment within healthcare infrastructure. This board would scrutinize applications for biosafety, patient safety, and environmental containment, and approve relevant proposals. Funding for development and deployment could be channeled through existing government mechanisms such as BIRAC (Biotechnology Industry Research Assistance Council). International precedent exists: the EU’s Contained Use Directive for GMOs could serve as a template adaptable to India’s regulatory landscape.
- Assumptions: This assumes ICMR, CDSCO, and GEAC have the institutional capacity, funding, and political will to implement a new advisory framework. It also assumes that a centralized national framework can accommodate the enormous diversity of hospital settings across India-from tertiary urban centers to rural primary health centers-without becoming either too rigid or too permissive.
- Risks of Failure & “Success”:
- Failure: The approval process takes 5–10 years, during which no legal deployment pathway exists. This pushes hospitals toward unregulated, informal use of engineered organisms with no safety oversight-paradoxically increasing risk. Meanwhile, over 1,000,000 people continue to die each year from HAIs in India alone.
- Success: The regulatory pathway, once established, becomes a barrier to entry that only large, well-resourced biotech companies can navigate. These companies gain outsized influence over the advisory committee’s decisions, creating a commercial monopoly over engineered living surfaces and undermining equitable access.
Action 2 - Technical: Engineering Auxotrophic Dependencies for Biocontainment
- Purpose: Currently, commercially available probiotic cleaning products use wild-type organisms with no engineered containment mechanisms. If these organisms spread beyond their intended surfaces, there are no built-in safeguards. I propose building auxotrophic dependencies into the B. subtilis chassis-engineering the organism to require a synthetic amino acid or nutrient not found in nature-so the colonizer cannot survive outside the hospital environment where that nutrient is actively supplied.
- Design: Academic synthetic biologists would develop and characterize the auxotrophic strains, validated by Institutional Biosafety Committees (IBCs). Standardized, well-characterized constructs would be deposited in repositories like Addgene or the iGEM Registry of Standard Biological Parts. Hospitals deploying the system would need to maintain a supply of the synthetic nutrient, which could be incorporated into routine surface treatment protocols. Redundant auxotrophies (requiring multiple synthetic nutrients simultaneously) would reduce the probability of reversion to environmental viability.
- Assumptions: This assumes we can effectively engineer genetic circuits with auxotrophic dependencies that reliably prevent survival outside the defined environment. It assumes reversion rates through mutation remain low enough to be safe-though published data on single auxotrophies in B. subtilis suggest reversion is non-trivial, making redundant safeguards essential. It also assumes that the hospital environment can be consistently maintained to support the colonizer’s engineered nutritional requirements.
- Risks of Failure & “Success”:
- Failure: Mutations in the engineered organisms bypass the installed auxotrophic dependencies, allowing survival outside the intended environment. A containment breach at scale could result in mass environmental contamination by antimicrobial-peptide-producing organisms, which becomes extremely difficult to remediate.
- Success: The auxotrophic dependency creates a new commercial bottleneck. Companies that hold patents on the specific synthetic nutrient required by the organism could monopolize the supply chain, establishing a cycle of dependence-you get the open-source organism for free, but you cannot use it without buying the proprietary nutrient. This undermines democratized access despite open-source genetic designs.
Action 3 - Incentive: Open-Source Consortium for Engineered Living Surfaces
- Purpose: Currently, engineered antimicrobial surface organisms are being developed in isolated academic labs (ETH Zurich, MIT Media Lab) and private companies, with no standardized sharing of constructs, safety data, or deployment protocols. Patents and IP held exclusively by these institutions greatly hinder the development and distribution of this technology in under-resourced settings where it is needed most. Akin to how the Linux operating system emerged from the collective efforts of thousands of developers and disrupted proprietary software, I propose an international open-access repository modeled on iGEM’s Registry of Standard Biological Parts-specifically for engineered surface colonizers-where anyone can contribute designs, genetic circuits, safety profiles, and deployment protocols.
- Design: Requires seed funding from organizations such as the WHO, Wellcome Trust, or Gates Foundation. Participating labs deposit genetic constructs, characterized safety profiles, and validated deployment protocols into the repository. In exchange, they receive access to the full library and structured co-authorship frameworks that incentivize contribution. WHO prequalifies validated strains for deployment in low-resource countries, creating a fast-track regulatory pathway analogous to its Essential Medicines prequalification programme. This decentralized model would accelerate development and acceptance of engineered living surfaces across diverse healthcare settings globally.
- Assumptions: This assumes academic labs and companies will share constructs and protocols openly, which conflicts with current publication-priority and profit-driven incentive structures in science and industry. It assumes WHO has the institutional capacity to manage a prequalification programme for engineered organisms, which it currently does not. It also assumes that sufficient contributors will participate to reach a critical mass of useful, well-characterized parts.
- Risks of Failure & “Success”:
- Failure: No one contributes, and the repository remains empty-a common fate of open-source initiatives without strong network effects or institutional mandates.
- Success: Open access enables actors with malicious intent to access engineered organisms and their protocols. However, the constructs in question are surface colonizers, not pathogens, which limits their weaponization potential compared to other synthetic biology tools. A more realistic success risk is that unvetted, poorly characterized constructs get deployed in hospitals by groups without adequate biosafety expertise, causing harm that discredits the entire approach.
- Next, score (from 1-3 with, 1 as the best, or n/a) each of your governance actions against your rubric of policy goals. Scoring Matrix *Scored 1–3 (1 = best/strongest contribution, 3 = weakest/least contribution, n/a = not applicable)
| Does the option: | Action 1: Regulatory Authority | Action 2: Auxotrophic Dependencies | Action 3: Open-Source Consortium |
|---|---|---|---|
| 1. Biosafety/Containment | |||
| a)Prevent uncontrolled spread | 1 | 1 | 3 |
| b)Prevent horizontal gene transfer | 2 | 1 | 3 |
| c)Prevent environmental contamination | 2 | 1 | 3 |
| 2. Patient Safety | |||
| a)No toxins/allergens to immunocompromised | 1 | 2 | 3 |
| b)Cannot cause opportunistic infection | 1 | 2 | 3 |
| 3. Equitable Access | |||
| a)Cost below chemical disinfectants | 3 | 2 | 1 |
| b)Deployable without specialized infrastructure | 3 | 2 | 1 |
| c)Open-source constructs | 3 | 2 | 1 |
| 3. Environmental Safety | |||
| a)No establishment in non-target environments | 2 | 1 | 3 |
| b)No contribution to resistance development | 2 | 1 | 3 |
| 4. Other Considerations | |||
| a)Minimizing costs and burdens to stakeholders | 3 | 2 | 1 |
| b)Feasibility | 2 | 2 | 2 |
| c)Not impede research | 3 | 2 | 1 |
| d)Promote constructive applications | 2 | 2 | 1 |
| Total | 30 | 23 | 30 |
| Action 2 (Technical) scores best overall (loweest score is the best) as it directly addresses the safety and containment goals. |
Last, drawing upon this scoring, describe which governance option, or combination of options, you would prioritize, and why. Outline any trade-offs you considered as well as assumptions and uncertainties.
Audience: Indian Council of Medical Research (ICMR)
I recommend a combination of all three governance actions, implemented in a phased sequence, because no single action adequately addresses all four policy goals.
Phase 1 : Technical safeguards first. The auxotrophic dependencies (Action 2) should be engineered and validated before any deployment is considered. This is non-negotiable. Without built-in biocontainment, no regulatory framework or open-source initiative can compensate for an organism that escapes its intended environment. Redundant auxotrophies should be the minimum standard.
Phase 2 : Open-source consortium in parallel. The open-source repository (Action 3) should be established concurrently with technical development, ensuring that safety data, construct characterization, and deployment protocols are shared from the beginning rather than locked behind institutional IP. This directly addresses the equitable access goals and prevents the commercial monopoly risks identified in Actions 1 and 2.
Phase 3 : Regulatory framework informed by real data. The regulatory authority (Action 1) should be established after sufficient technical and safety data exists from the first two phases. This prevents the common failure mode of regulating a technology before it is understood, which leads to either overly restrictive or dangerously permissive frameworks.
Key trade-offs: Speed versus safety remains the central tension. Moving too fast with deployment risks a containment failure that could set back the entire field of engineered living surfaces by a decade—eroding public trust in synthetic biology. Moving too slowly means people continue to die from preventable infections in hospitals that lack the resources for conventional sterility maintenance. The phased approach attempts to balance this by building safety into the organism first, sharing knowledge openly second, and formalizing regulation third.
Uncertainties: It remains unclear whether auxotrophic containment can be made robust enough at scale, whether open-source incentives can overcome academic publishing pressures, and whether Indian regulatory bodies can develop new frameworks within a reasonable timeframe. These uncertainties should be revisited as the technology matures.
Reflecting on what you learned and did in class this week, outline any ethical concerns that arose, especially any that were new to you. Then propose any governance actions you think might be appropriate to address those issues. This should be included on your class page for this week. This week’s exploration of governance frameworks surfaced several ethical concerns that were new to me.
The first is the tension between speed and safety in resource-constrained settings. In well-funded Western hospitals, the cost of waiting for perfect regulation is inconvenience. In under-resourced Indian hospitals where I am training, the cost of waiting is measured in lives. This creates an ethical pressure to deploy before full safety validation ,a pressure I now recognize must be resisted, because a single high-profile failure would destroy trust in this entire approach and harm far more people long-term.
The second is the dual-use nature of open-source biology. I initially assumed open-sourcing was unambiguously good democratizing access, preventing monopoly. But the governance analysis forced me to consider that open access also means open access for actors without adequate biosafety training or with malicious intent. Akin to the quacks without formal medical education who practice in remote and under resourced areas causing the patients more damage and sometimes death. The realistic risk is not bioweapons (surface colonizers are poor candidates), but rather poorly characterized organisms deployed without adequate containment by well-meaning but under-equipped groups.
The third and most personally challenging is the realization that the success of this technology could create new forms of dependency even while solving the original problem. If the auxotrophic nutrient becomes a patented commercial product, under-resourced hospitals could end up dependent on supply chains they cannot control or afford. This mirrors the broader pattern I have seen in Indian healthcare, where solutions designed for equity are captured by commercial interests. Governance must anticipate this from the beginning, not as an afterthought.
Week 2 Lecture Prep
Homework Questions from Professor Jacobson
Nature’s machinery for copying DNA is called polymerase. What is the error rate of polymerase? How does this compare to the length of the human genome. How does biology deal with that discrepancy?
- Error rate: The raw “bit-flip” rate of a standard DNA Polymerase is roughly $10^{-5}$ (1 in 100,000). With intrinsic proofreading (3’ → 5’ exonuclease activity), it drops to ≈ $10^{-7}$.
- The comparison to the human genome : The human genome is ≈ $3.2 \times 10^9$ base pairs (3.2 Gigabytes of code).
- The way biology deals with this discrepancy: A post processing algorithm called MMR is run to fix these errors. This brings the final error rate to ≈ $10^{-9}$ , which is around 3 errors per cell division which is low enough to preserve genetic code but high enough to trigger evolution- an optimal signal to noise ratio.
How many different ways are there to code for an average human protein? Why don’t they all work?
- The Coding Space: Due to codon degeneracy (61 codons for 20 amino acids), there are roughly $3N$ combinations for a protein of length $N$. For an average 400-amino acid protein, this is $3{400}$ (approx $10^{190}$)
- Why most “codes” fail (Optimization constraints):
- Codon Bias: The cell has a limited “RAM” of specific tRNAs. Rare codons cause ribosomal stalling because the hardware isn’t available.
- mRNA Structure: Bad sequences can fold into tight hairpins, jamming the ribosome like a paper shredder.
- GC Content: If G/C is too high (>70%) or low, the DNA becomes structurally unstable or difficult to replicate
Homework Questions from Dr. LeProust
- What’s the most commonly used method for oligo synthesis currently?
- Phosphoramidite Chemistry: A solid-phase synthesis method that builds DNA strands nucleotide by nucleotide on a support matrix (like a silicon chip or column).
- Why is it difficult to make oligos longer than 200nt via direct synthesis?
- Yield Decay: Synthesis is an exponential decay function. With 99.5% efficiency per step, a 200nt strand has a yield of $0.995^{200} \approx 36%$. Beyond this, the “noise” (truncated sequences and side reactions) drowns out the signal, making purification inefficient.
- Why can’t you make a 2000bp gene via direct oligo synthesis?
- The Yield Cliff: The math makes it impossible: $0.995^{2000} \approx 0.004%$. You cannot “print” a gene this long atom-by-atom because the yield is effectively zero.
- The Solution: We switch from “printing” to “assembly.” We synthesize short tiles (e.g., 200bp) and stitch them together using methods like Gibson Assembly or Golden Gate Assembly.
Homework Question from George Church
- What are the 10 essential amino acids in all animals and how does this affect your view of the “Lysine Contingency”?
- The 10 Essentials: Phenylalanine, Valine, Threonine, Tryptophan, Isoleucine, Methionine, Histidine, Arginine, Leucine, Lysine.
- The Flaw: The Jurassic Park logic was security theater. Since all animals (including humans) are Lysine auxotrophs (we cannot produce it), a strictly Lysine-depeddent dinosaur would simply survive by eating chickens, soy, or humans—sources rich in Lysine.
- True containment requires engineering dependence on a Non-Canonical Amino Acid (ncAA)—a synthetic building block not found in nature. This ensures “Zero Leakage” because the organism cannot find the nutrient in the wild.
Citation: I used Gemini and Claude to stress test my ideas and refine them for the sections on auxotrophy All governance analysis, clinical observations, and project framing are my own work..