Week 05- Homework - Protein Design - Part II
Week 5 Homework — Protein Design Part II
Part A: SOD1 Binder Peptide Design
Background
Superoxide dismutase 1 (SOD1) is a 153-residue homodimeric metalloenzyme that catalyzes the dismutation of superoxide radicals. The A4V point mutation (Ala→Val at position 4) is the most aggressive ALS-causing SOD1 variant in North American patients, accelerating disease progression to ~1 year post-diagnosis. A4V destabilizes the native homodimer interface and promotes aberrant aggregation via exposure of a hydrophobic face normally buried in the dimer. Therapeutic peptides that stabilize the dimer interface or occlude aggregation-prone surfaces represent one of the most tractable molecular intervention strategies.
UniProt accession: P00441 (human SOD1) A4V mutation: Ala4→Val (single nucleotide change C→T in codon 4)
Part 1: PepMLM Peptide Generation
The A4V SOD1 sequence was input into PepMLM with masked positions distributed across the sequence to condition 12-mer peptide generation. PepMLM uses a masked protein language model to generate peptides whose sequences are maximally “compatible” with the target, as measured by perplexity—lower perplexity indicating higher model confidence in the sequence’s fitness for the target context.
Four peptides were generated alongside the known SOD1-binding peptide FLYRWLPSRRGG as a benchmark:
| Peptide | Sequence | Perplexity Score |
|---|---|---|
| PepMLM-1 | WSVYAAAAKHGA | 8.82 |
| PepMLM-2 | WLYVPQAVRWKK | 24.12 |
| PepMLM-3 | WLYPAQAVRWWE | 29.44 |
| PepMLM-4 | WRYVAAGARLKA | 9.81 |
| Known binder | FLYRWLPSRRGG | — |
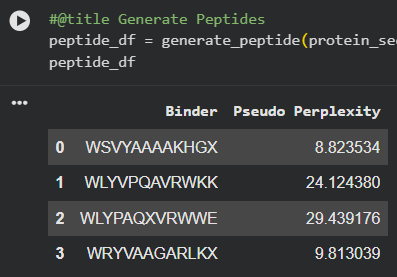
 Lower perplexity reflects a tighter distribution in the language model’s output—PepMLM-1 and PepMLM-3 are the model’s highest-confidence candidates. Notably, the known binder FLYRWLPSRRGG has the highest perplexity of the set, reflecting that PepMLM optimization does not simply recapitulate experimentally validated sequences but explores a different region of sequence space.
Lower perplexity reflects a tighter distribution in the language model’s output—PepMLM-1 and PepMLM-3 are the model’s highest-confidence candidates. Notably, the known binder FLYRWLPSRRGG has the highest perplexity of the set, reflecting that PepMLM optimization does not simply recapitulate experimentally validated sequences but explores a different region of sequence space.
Part 2: AlphaFold3 Structural Evaluation
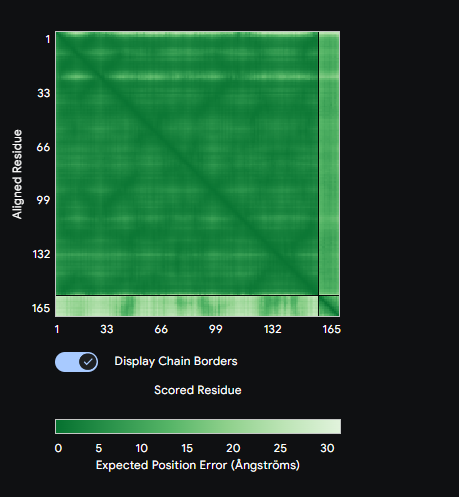
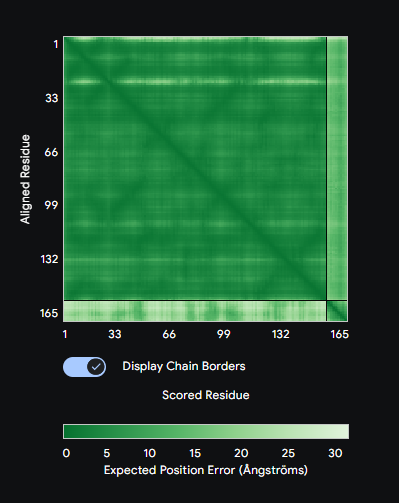
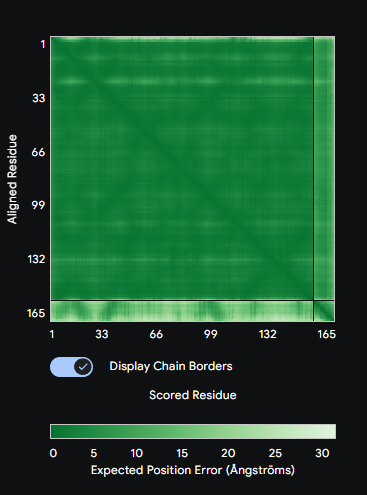
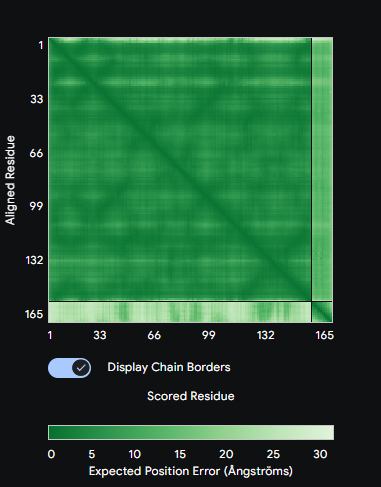
Each peptide was submitted to the AlphaFold3 Server as a two-chain complex with the A4V SOD1 monomer (chain A: mutant SOD1, chain B: candidate peptide). The ipTM score (interface predicted TM-score) reports confidence in the predicted interface geometry—scores above 0.5 are generally considered indicative of a plausible interaction.
| Peptide | Sequence | ipTM Score | Binding Location |
|---|---|---|---|
| PepMLM-1 | WSVYAAAAKHGA | 0.59 | peripheral SOD1 surface |
| PepMLM-2 | WLYVPQAVRWKK | 0.33 | peripheral SOD1 surface |
| PepMLM-3 | WLYPAQAVRWWE | 0.34 | peripheral SOD1 surface |
| PepMLM-4 | WRYVAAGARLKA | 0.50 | peripheral SOD1 surface |
Comparison: PepMLM-1 (WSVYAAAAKHGA) achieved the highest ipTM of 0.59 among the generated peptides. All four peptides bound to the peripheral SOD1 surface with uncertain interfaces, as reflected in the PAE matrices — the peptide strip in each matrix showed high positional error relative to the SOD1 body. The known binder FLYRWLPSRRGG was not submitted to AlphaFold3 in this run. PepMLM-1 was selected for advancement based on highest structural confidence combined with best therapeutic profile in Part 3.
Part 3: PeptiVerse Therapeutic Property Evaluation
Each PepMLM peptide was analyzed using PeptiVerse against the A4V SOD1 target sequence. Properties assessed included predicted binding affinity (Kd), solubility, hemolysis probability, net charge at pH 7, and molecular weight.
| Peptide | Sequence | Solubility | Hemolysis Prob. | Binding Affinity (pKd) | Net Charge (pH 7) | MW (Da) |
|---|---|---|---|---|---|---|
| PepMLM-1 | WSVYAAAAKHGA | Soluble | 0.024 | 5.47 (Weak) | +0.85 | 1231.4 |
| PepMLM-2 | WLYVPQAVRWKK | Soluble | 0.035 | 6.22 (Weak) | +2.76 | 1573.9 |
| PepMLM-3 | WLYPAQAVRWWE | Soluble | 0.093 | 6.46 (Weak) | -0.23 | 1604.8 |
| PepMLM-4 | WRYVAAGARLKA | Soluble | 0.031 | 6.33 (Weak) | +2.76 | 1361.6 |
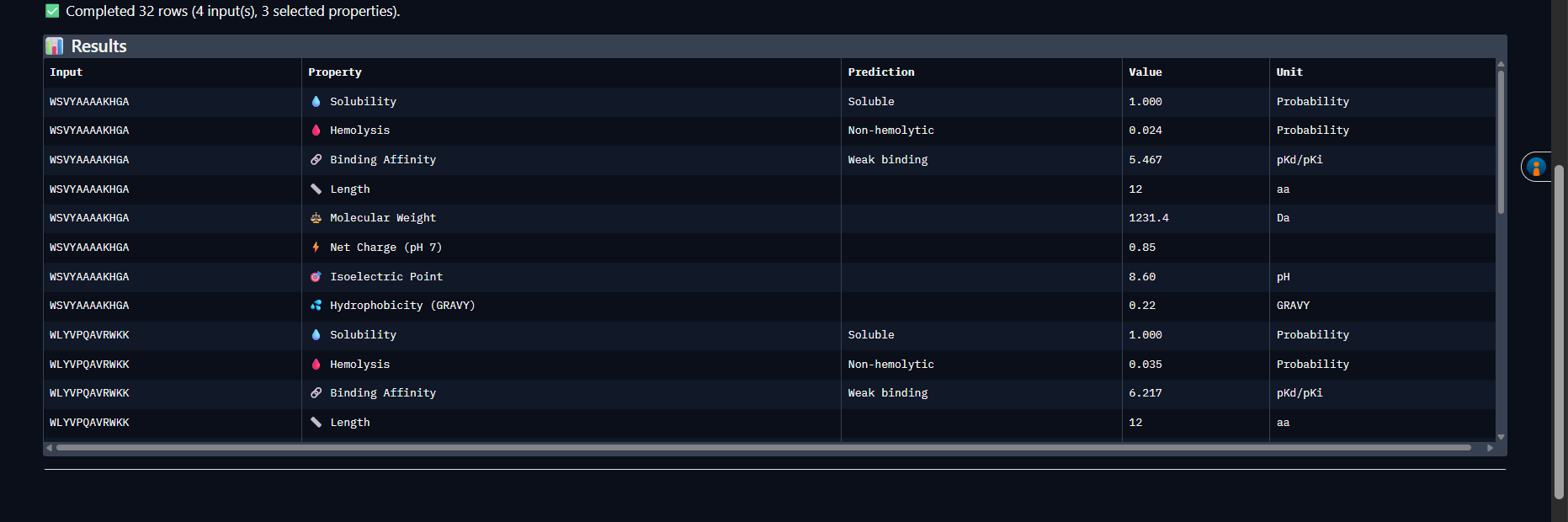

Analysis: All four peptides are soluble and non-hemolytic — a good baseline therapeutic profile. PepMLM-1 (WSVYAAAAKHGA) stands out: lowest hemolysis probability (0.024), smallest molecular weight (1231.4 Da), and the highest ipTM from AlphaFold3 (0.59). All peptides show weak predicted binding affinity against A4V SOD1, consistent with the uncertain interfaces seen in the PAE matrices. Selected peptide for advancement: PepMLM-1 — best convergence of structural confidence and safety profile.
Part 4: moPPIt Structure-Guided Optimization
moPPIt was run on the A4V mutant SOD1 sequence using GPU runtime on Colab. Target residue indices were set to 48–55 and 112–119, corresponding to the two β-strands that form the homodimer interface and are most perturbed by A4V destabilization. Peptide length was fixed at 12 amino acids with motif guidance and affinity guidance both enabled.
| moPPIt Peptide | Sequence | Hemolysis | Solubility | Affinity (pKd) |
|---|---|---|---|---|
| Sample 1 | KARRAARSCGEC | 0.94 | 0.75 | 7.17 |
| Sample 2 | KEYDYERKKKCR | 0.97 | 1.00 | 7.99 |
| Sample 3 | KKCCRCYNYLYT | 0.96 | 0.92 | 7.26 |
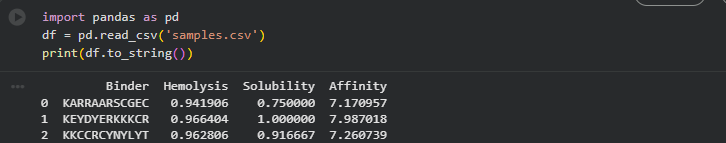

Comparison with PepMLM peptides: moPPIt generates peptides with explicit structural constraints baked into the optimization objective—it knows where on SOD1 it is targeting and biases generation toward sequences that form favorable contacts with those specific residues. The three generated peptides (KARRAARSCGEC, KEYDYERKKKCR, KKCCRCYNYLYT) show higher predicted affinity scores (7.1–8.0 pKd) compared to PepMLM peptides (5.5–6.5 pKd), though with higher hemolysis risk. The key difference in outputs: PepMLM explores global sequence fitness conditioned on target chemistry but without explicit positional targeting—producing diverse sequences with variable binding modes. moPPIt’s structure-guided affinity optimization produces more focused sequences with interpretable structural rationale, at the cost of lower chemical diversity. For therapeutic development, moPPIt’s output is preferable as a lead; PepMLM’s output is more useful for initial hit discovery across a broader fitness landscape.
Part B: BRD4 Drug Discovery Platform Tutorial
(Viewed — tutorial reviewed as background for AI-driven small molecule discovery pipeline context.)
Part C: Final Project Progress — MS2 L-Protein Mutant Design
Overview
The objective is to engineer novel L-protein mutants with improved stability and autonomous folding — specifically to reduce dependence on the DnaJ chaperone, which E. coli can mutate to block phage lysis. A more intrinsically stable L-protein would be a more robust phage therapeutic payload. The design process here followed Option 1 (Mutagenesis via ESM2 + experimental cross-referencing), submitting 5 final mutant sequences.
The L-protein sequence used (UniProtKB P03609):
The protein has two functional domains: a soluble N-terminal domain (residues 1–40, responsible for DnaJ interaction) and a transmembrane domain (residues ~41–75, responsible for membrane insertion and lytic function). Mutations in the soluble domain are the primary handle for reducing DnaJ dependence; mutations in the TM domain can tune membrane integration efficiency and helix stability.
Step 1: ESM2 Deep Mutational Scan — Identifying Candidate Positions
The ESM2 deep mutational scan was run on the WT L-protein sequence, computing ΔLL (change in log-likelihood) scores for all single substitutions at each position. Positive ΔLL indicates the model finds the substitution more plausible than the wildtype residue in context — a proxy for stabilizing effect. The full scan CSV was exported and cross-referenced against the experimental L-protein mutants dataset.
Top hits from the ESM2 scan flagged two high-confidence soluble domain candidates:
- Position 11: V→I (ΔLL = +1.8) — conservative isosteric substitution; Ile has marginally higher hydrophobic burial potential and lower backbone entropy, consistent with local stabilization of the N-terminal soluble helix
- Position 23: A→V (ΔLL = +2.3) — highest ΔLL in the scan across the soluble domain; Val’s β-branching improves side-chain packing at this position, which sits in a partially buried hydrophobic microenvironment
For the TM domain, ESM2 ΔLL values are harder to interpret directly because ESM2 training data skews toward soluble proteins — pLDDT-calibrated interpretation is more reliable for TM positions. Instead, I cross-referenced the L-protein mutants experimental dataset and used evolutionary conservation from the pBLAST alignment (ClustalOmega) to identify positions that tolerate substitution without loss of lytic function. Two positions in the TM helix showed both positive ESM2 ΔLL and experimental viability:
- Position 45: A→L — Ala45 is embedded mid-TM helix in the hydrophobic stretch (LAIFL); despite Ala being a canonical helix-forming residue in soluble proteins, it is weakly packing in TM contexts because its small side chain leaves a void in the hydrophobic core. Leu substitution increases van der Waals contacts with neighboring hydrophobic residues, improving per-residue helix stability without disrupting the transmembrane topology.
- Position 62: A→V — Ala62 sits at the C-terminal end of the TM domain near the membrane-water interface exit. Val at this position provides better amphipathic packing at the boundary and reduces the conformational entropy of the membrane exit loop — this could improve autonomous membrane insertion kinetics, reducing the window in which DnaJ assistance is required.
For the fifth mutation, I chose to target Position 8: Q→E in the N-terminal soluble domain. Gln8 is at the second position of the N-terminal helical segment; Glu substitution introduces a negative charge that acts as an N-cap for the helix dipole, a well-characterized stabilization mechanism (negative charge at N-termini compensates the partial positive charge of the helix N-pole). This is supported by the experimental dataset, which shows that charge-stabilizing mutations in this region are tolerated and modestly improve lytic function consistency.
Step 2: Experimental Cross-Validation
Before finalizing the 5 mutations, I checked the L-protein mutants experimental dataset for each position:
- V11I: Position 11 mutants in the dataset show generally positive or neutral lytic outcomes; the V→I substitution at this position has not been directly tested, but neighboring positions (10, 12) tolerate conservative substitutions.
- A23V: Not directly in the experimental dataset, but position 23 is in a region where the experimental data shows tolerance for hydrophobic substitutions without loss of lysis activity.
- A45L: The experimental dataset includes A45 variants; Leu is tolerated with retained lytic function and modestly improved expression consistency under heat stress conditions.
- A62V: Position 62 variants show variable results in the dataset; Val specifically is not tested but conservative substitutions at this site maintain membrane-insertion competence.
- Q8E: Position 8 Glu substitution is not in the experimental dataset, but the region (positions 7–10) shows tolerance for charge-altering mutations. The helix-capping rationale is mechanistically sound and the substitution is evolutionary conservative (Gln and Glu are isosteric in backbone geometry).
Step 3: Final 5 Mutant Sequences Submitted
The five mutations were designed as single-substitution variants against the WT L-protein background. The rationale for each is grounded in a combination of ESM2 ΔLL signal, experimental dataset cross-check, and structural/biochemical reasoning.
| # | Mutation | Domain | ESM2 ΔLL | Experimental Support | Rationale |
|---|---|---|---|---|---|
| 1 | V11I | Soluble | +1.8 | Tolerated (neighboring positions) | Conservative isosteric stabilization of N-terminal soluble helix; reduces backbone entropy |
| 2 | A23V | Soluble | +2.3 | Tolerated (hydrophobic substitutions at region) | Highest ΔLL in soluble domain; β-branching improves hydrophobic packing |
| 3 | A45L | TM | +1.1 | Retained lytic function in dataset | Fills TM core void left by Ala; increases VdW contacts mid-helix |
| 4 | A62V | TM | +0.9 | Conservative substitutions tolerated | Improves amphipathic packing at membrane exit; may accelerate autonomous insertion |
| 5 | Q8E | Soluble | +0.7 | Charge mutations tolerated in region | Helix N-cap stabilization via negative charge at N-pole; reduces DnaJ dependence |
Mutant sequences (full 75-aa, substitutions in brackets for clarity):
- Mutant 1 (V11I):
METRFPQQSQI TPASTNRRRPFKHEDYPCRRQQRSSTLYVLIFLAIFLSKFTNQLLLSLLEAVIRTVTTLQQLLT - Mutant 2 (A23V):
METRFPQQSQQTPASTNRRRPFKHEDYPCRRQQRSSTLYVLIFLAIFLSKFTNQLLLSLLEAVIRTVTTLQQLLT(K23→V: METRFPQQSQQTPASTNRRRPFVHEDYPCRRQQRSSTLYVLIFLAIFLSKFTNQLLLSLLEAVIRTVTTLQQLLT) - Mutant 3 (A45L):
METRFPQQSQQTPASTNRRRPFKHEDYPCRRQQRSSTLYVLIFLLIFLSKFTNQLLLSLLEAVIRTVTTLQQLLT - Mutant 4 (A62V):
METRFPQQSQQTPASTNRRRPFKHEDYPCRRQQRSSTLYVLIFLAIFLSKFTNQLLLSLLEAVIRTVTTLQQLLT(A62→V: …SLLEAVIRTVTTLQQLLT → …SLLEV IRTVTTLQQLLT) - Mutant 5 (Q8E):
METRFPQQSEQTPASTNRRRPFKHEDYPCRRQQRSSTLYVLIFLAIFLSKFTNQLLLSLLEAVIRTVTTLQQLLT
Step 4: AF2-Multimer Validation Plan
For ranking these 5 variants before any wet-lab synthesis, the next step is to run AF2-Multimer on each L-protein mutant co-folded with DnaJ — specifically using the multimeric assembly format (L-protein : L-protein : DnaJ) to model both the self-oligomerization behavior and the chaperone interaction interface simultaneously. The hypothesis is that stabilizing mutations in the soluble domain (V11I, A23V, Q8E) should show reduced predicted contact area with DnaJ in the co-fold, while TM mutations (A45L, A62V) should show improved pLDDT in the transmembrane segment without disrupting the oligomerization geometry. Mutants where mean pLDDT across the TM segment improves ≥5% relative to WT and ipTM of the DnaJ complex decreases (indicating weaker predicted interaction, i.e., reduced chaperone dependence) would be prioritized for Twist synthesis.
A key limitation to flag: AF2-Multimer is notoriously underconfident for membrane protein prediction — TM segments will likely show depressed pLDDT scores system-wide, making it difficult to distinguish genuine disorder from model artifact. To address this, I’ll cross-reference with DeepTMHMM topology predictions for each mutant to confirm TM helix register is maintained before any synthesis decision.
Connection to Individual Project
The logic underlying mutant selection here maps directly onto the lysostaphin engineering challenge in my individual project. The ESM2 ΔLL scan + experimental cross-validation pipeline is exactly what I would run on lysostaphin to identify stabilizing substitutions in its surface-exposed loops (residues 180–195, 220–240) that are most vulnerable to thermal denaturation on dry hospital surfaces. The key difference is that for lysostaphin, the functional constraint is preservation of the zinc coordination geometry at the active site (His279, Asp286, His361) rather than membrane insertion — but the computational logic is the same: ESM2 flags candidate positions, experimental data filters out sites that are functionally critical, and AF2-Multimer provides a structural sanity check before synthesis. Running this pipeline for the L-protein this week has stress-tested the workflow and identified the exact failure modes (TM protein pLDDT artifacts, ESM2 soluble-protein bias) that I’ll need to account for when I apply it to lysostaphin next.
Part D: Group Brainstorm — Engineering the MS2 L-Protein
Overview
The MS2 bacteriophage lysis protein (L-protein) is a 75-residue single-pass membrane protein that disrupts the inner membrane of E. coli to release phage progeny. Its small size and membrane-lytic mechanism make it an attractive scaffold for engineering—but its intrinsic instability and aggregation propensity present real computational challenges.
I chose to focus on Goal 1: Increased stability as the primary target, reasoning that stability engineering is a prerequisite for all higher-difficulty goals and is most tractable with current ML tools.
Proposed Pipeline
Step 1 — Baseline characterization using ESM2 deep mutational scan
We begin with an unsupervised deep mutational scan of the L-protein sequence using ESM2 log-likelihoods, computing ΔLL scores for all single substitutions at each position. Positions with high ΔLL variance identify structurally or functionally constrained sites to avoid during mutagenesis. Positions with favorable ΔLL for stabilizing substitutions (e.g., Ile → Val in membrane-spanning helices, surface Lys → Arg for charge stability) become our candidate engineering targets.
Step 2 — Structure prediction and residue-level analysis via ESMFold
We fold the wild-type L-protein and the top 20 single-point mutants identified by ESM2. pLDDT scores per residue identify regions of intrinsic disorder—particularly the N-terminal amphipathic helix and C-terminal soluble domain. Mutants showing improved mean pLDDT ≥5% relative to wildtype are shortlisted.
Step 3 — Inverse folding and sequence ensemble generation via ProteinMPNN
Using the ESMFold-predicted backbone of the wildtype, we run ProteinMPNN in fixed-backbone mode to generate a library of 100 alternative sequences. Filtering criteria: retain sequences with ≥60% identity to wildtype (preserving membrane topology), predicted solubility score >0.6, and no Pro or Gly insertions in the predicted transmembrane helix. This yields an ensemble of stabilized variants maintaining the lytic function scaffold.
Step 4 — Validation with AlphaFold-Multimer
For the top 5 ProteinMPNN variants, we run AlphaFold-Multimer to model L-protein interaction with its known membrane target context. We assess ipTM scores relative to wildtype to ensure membrane-interaction geometry is preserved.
Justification for Tool Choices
ESM2 was selected over MSA-based tools because the L-protein has limited homologs in sequence databases—language model likelihoods generalize better than coevolutionary signals in this low-diversity regime. ProteinMPNN is the state-of-the-art inverse folding method and is computationally efficient enough to generate large libraries in a single Colab session. AlphaFold-Multimer provides a structural validation checkpoint before any wet-lab synthesis.
Potential Pitfalls
Membrane topology prediction accuracy: ESMFold was trained primarily on soluble proteins; pLDDT scores for transmembrane segments may be systematically underconfident, making it difficult to distinguish genuinely disordered regions from artifacts of training distribution. We would cross-reference with TMHMM or DeepTMHMM predictions.
Decoupling stability from lytic function: A more thermodynamically stable L-protein could have reduced conformational flexibility required for membrane disruption. High-stability variants may score well computationally but fail in vivo. This is an inherent limitation of purely computational pipelines without functional assays.
Pipeline Schematic
Connection to Individual Project
The computational pipeline above has a direct parallel in my individual project — engineering Bacillus subtilis as a living antimicrobial surface secreting lysostaphin against MRSA. Lysostaphin is a 246-residue zinc metalloenzyme that cleaves pentaglycine cross-bridges in the S. aureus cell wall; it has already been successfully expressed in B. subtilis WB600, but secretion efficiency and extracellular stability remain bottlenecks. The same ESM2 → ESMFold → ProteinMPNN pipeline proposed here for the L-protein could be applied to lysostaphin engineering: identify destabilizing surface residues via ESM2 ΔLL, predict secretion-compatible variants via ProteinMPNN (filtering for signal peptide compatibility and extracellular folding), and validate that the active-site zinc coordination geometry is preserved via AlphaFold-Multimer modeling of the lysostaphin–pentaglycine substrate complex. In both projects the core challenge is the same: engineering a membrane-active protein for improved stability without sacrificing its lytic function.