Week-07-Homework-Genetic-Circuits-Part-II
Part 1: Intracellular Artificial Neural Networks (IANNs)
Q1: Advantages of IANNs over Traditional Genetic Circuits
Traditional genetic circuits operate as Boolean logic gates—each node is either “on” or “off,” with behavior determined by discrete threshold crossings of transcription factor concentrations. While powerful for implementing deterministic logic (AND, OR, NOT, toggle switches), this architecture has fundamental limitations when the biological task demands graded, context-dependent, or multi-signal integration.
IANNs—genetic circuits whose topology and weight-encoding logic mirrors the multilayer perceptron—address several of these constraints simultaneously:
Continuous signal representation. Traditional circuits collapse analog transcription factor concentrations into binary outputs at threshold crossings. IANNs retain continuous signal values through each layer, preserving quantitative information about input strength. For a biosensor tracking a continuously varying metabolite, this matters enormously—a Boolean circuit can only report “above or below threshold,” while an IANN can encode graded responses proportional to input magnitude.
Multi-signal integration with arbitrary weighting. A classical N-input AND gate requires all N inputs to exceed threshold simultaneously. An IANN neuron computes a weighted sum of inputs before applying a nonlinear activation function—meaning inputs contribute with variable importance, and the threshold for activation depends on the combined weighted signal. This enables far richer input-output relationships with fewer genetic components than equivalent Boolean implementations.
Robustness to parameter variation. Boolean circuits are brittle near threshold—small changes in promoter strength, RBS efficiency, or plasmid copy number shift the switching threshold and can flip circuit state. IANN nodes, by operating in a saturation regime far from their individual thresholds, are intrinsically more robust: perturbations shift the activation curve but leave the qualitative output largely intact.
Trainability and adaptability. In principle, IANN weights (encoded in repressor binding affinities, RBS strengths, or protein-protein interaction Kds) can be tuned—either computationally before construction or, in more advanced designs, via directed evolution. Traditional Boolean circuits have no analogous training paradigm; each redesign is a manual re-engineering effort.
Handling of combinatorially complex inputs. For tasks requiring integration of many simultaneous signals (e.g., a cell-state classifier distinguishing cancer subtypes based on 10+ miRNA inputs), Boolean circuits require exponentially many components; IANNs scale linearly with the number of inputs per layer.
Q2: Proposed IANN Application — Multi-Pathogen Antimicrobial Surface Classifier
Application context: My final project engineers Bacillus subtilis as a living antimicrobial surface for hospital environments in resource-constrained settings. The current design uses a single-input toehold switch sensing mecA mRNA (the methicillin resistance determinant) to trigger lysostaphin secretion against MRSA. This is functionally a single-input Boolean gate — pathogen present → kill. The critical limitation: hospital surfaces harbour multiple co-occurring pathogens (S. aureus, P. aeruginosa, K. pneumoniae, E. faecium), and a single-target response leaves the surface vulnerable to the remaining species. Worse, indiscriminate broad-spectrum antimicrobial secretion would disrupt commensal biofilms and accelerate resistance evolution.
An IANN architecture solves this by enabling the engineered B. subtilis to classify the pathogen environment from multiple simultaneous RNA signals and mount a proportional, pathogen-specific response — secreting the right antimicrobial at the right concentration, only when the threat profile warrants it.
Input/output behavior:
Layer 1 — Pathogen sensing (3 input nodes): Three orthogonal toehold switch sensors detect species-specific mRNA transcripts leaked from lysed or metabolically active pathogens on the surface: (1) mecA mRNA → MRSA detection, (2) lasR mRNA → P. aeruginosa quorum sensing signal, and (3) blaKPC mRNA → carbapenem-resistant K. pneumoniae. Each toehold switch output is an intermediate transcription factor produced at a level proportional to the cognate mRNA concentration — not a binary on/off, but a graded analog signal encoding pathogen burden.
Layer 1 → Layer 2 integration (hidden layer): The three toehold switch outputs converge on two Layer 2 integration nodes, each controlled by a synthetic promoter with multiple operator sites. Integration node H₁ computes a weighted sum biased toward Gram-positive signals (mecA weight w₁ = high, lasR weight w₂ = low, blaKPC weight w₃ = medium). Integration node H₂ computes a weighted sum biased toward Gram-negative signals (w₁ = low, w₂ = high, w₃ = high). Weights are physically encoded by RBS strength on each transcription factor input feeding that node. The nonlinear activation function at each node is the inherent Hill-function response of the synthetic promoter (~n = 2–4 depending on operator architecture).
Layer 2 → Output layer (effector selection): H₁ drives lysostaphin secretion (Gram-positive killing — cleaves pentaglycine cross-bridges in S. aureus cell wall). H₂ drives Art-175 secretion (engineered endolysin–SMAP29 fusion effective against Gram-negative outer membranes). A third output node, activated only when H₁ AND H₂ both exceed threshold, drives dispersin B (biofilm-degrading enzyme targeting poly-β-1,6-N-acetylglucosamine in mixed-species biofilms).
Why IANN architecture matters here over Boolean logic: A Boolean implementation would require 2³ = 8 separate circuit branches to cover all pathogen combinations — and still produce only binary (full-dose or zero) outputs. The IANN achieves graded, proportional antimicrobial secretion from a 3-node → 2-node → 3-output architecture (8 genetic parts vs. ~24 for equivalent Boolean coverage), and critically, it responds to partial pathogen signatures with partial responses, avoiding wasteful over-secretion that accelerates resistance evolution.
Connection to my project: This IANN design is a direct architectural upgrade of the single-input toehold-switch → T7 RNAP → lysostaphin pipeline I described in Week 3. The toehold switch sensing layer is already validated computationally (30 high-confidence mecA designs from my screening of 89 candidates, top candidate at position 234 with 9.1× ON/OFF ratio). Extending this to lasR and blaKPC targets requires the same toehold switch design workflow. The T7 RNAP amplification cascade from my current design maps naturally onto the hidden layer integration nodes.
Potential limitations:
- Cross-reactivity between toehold switches: At high mRNA concentrations, partial complementarity between toehold switch sensors and non-target transcripts could produce false-positive signals. Computational screening (NUPACK) must verify orthogonality across all three sensor-target pairs — my Week 3 screening pipeline would need to be extended to a 3×3 cross-reactivity matrix.
- Temporal mismatch between sensing and effector secretion: Toehold switches respond within minutes of mRNA exposure, but lysostaphin and Art-175 secretion requires protein synthesis and Sec-pathway translocation (~30–60 minutes). Fast-growing pathogens like K. pneumoniae (doubling time ~20 min) may establish dangerous colony densities before the IANN output layer reaches effective antimicrobial concentrations.
- Weight tuning in vivo: Unlike digital IANNs where weights are adjusted by gradient descent, genetic IANN weights (RBS strengths) are fixed at construction time. Mis-calibrated weights would require re-engineering the construct — there is no in-situ learning. Directed evolution of the RBS library could partially address this, but at significant experimental cost.
Q3: Multilayer Perceptron Diagram — Endoribonuclease → Fluorescent Protein
Design rationale: Using an endoribonuclease as the Layer 1 → Layer 2 signal carrier is architecturally elegant — it operates post-transcriptionally, decoupling Layer 1 and Layer 2 on different molecular substrates (protein vs. RNA), and avoids transcriptional interference between layers. CasRx (a Type VI-D CRISPR effector) is orthogonal to most bacterial and mammalian transcriptional regulators, making it particularly suitable for implementing clean two-layer IANNs without cross-talk.
Part 2: Fungal Materials
Q1: Existing Fungal Material Examples, Applications, and Trade-offs
Fungal materials—primarily mycelium composites formed from the dense hyphal networks of filamentous fungi—have emerged as a serious alternative to petroleum-derived foams, plastics, and leather in the last decade.
Mycelium composite packaging (Ecovative Design): The flagship application: agricultural waste (corn husks, hemp hurds) is colonized by Ganoderma spp. mycelium under controlled humidity and CO₂ conditions. The resulting composite blocks are used as protective packaging for electronics and fragile goods—replacing expanded polystyrene (EPS). Advantages: fully compostable, carbon-sequestering during growth, fire-resistant, thermally and acoustically insulating, scalable to large geometries without injection molding tooling costs. Disadvantages: lower specific mechanical strength than EPS (compressive strength ~50–200 kPa vs. EPS at 100–400 kPa), water sensitivity without chemical treatment, growth-to-form process requires 5–7 days minimum, not yet suitable for sterile or food-contact applications without additional processing.
Mycelium leather (Bolt Threads — “Mylo,” Ecovative — “Forager”): Thin mycelium sheets grown from Ganoderma or Pleurotus species on agricultural waste are processed (dried, tanned, and finished) to produce a leather-like flexible material. Used commercially in luxury goods by Stella McCartney and Lululemon. Advantages: animal-free, biodegradable, water-resistant post-treatment, pore size tunable via growth conditions. Disadvantages: tensile strength and durability still inferior to full-grain leather (~10–15 MPa vs. ~20–35 MPa for bovine leather), requires chemical tanning with conventional agents to achieve desired hand feel, scaling manufacturing to cost-parity with synthetic leather remains unresolved.
Fungal acoustic panels and construction materials: Compressed mycelium boards with densities of 100–200 kg/m³ are used for acoustic damping panels and low-load structural elements. Acoustic absorption coefficients comparable to mineral wool at mid-frequencies (500–2000 Hz). Limited to non-structural applications due to lower flexural modulus.
Q2: Genetic Engineering Objectives for Fungi and Synthetic Biology Advantages
Engineering objectives:
The primary genetic engineering targets for improved fungal materials are: (1) chitin synthase engineering to control hyphal wall composition and increase mechanical strength via higher chitin cross-link density; (2) laccase and peroxidase pathway modification to tune lignin binding at the hyphal surface, improving adhesion to lignocellulosic substrates; (3) fruiting body suppression to redirect metabolic flux from reproductive structures toward vegetative hyphal biomass during the growth phase; and (4) hydrophobin engineering to tune surface wettability of the final material without post-processing chemical treatment.
Why synthetic biology in fungi offers distinct advantages over bacterial approaches:
Fungi—particularly Aspergillus niger, Trichoderma reesei, and Ganoderma lucidum—have several properties that bacteria lack which make them intrinsically better chassis for material applications. First, fungal hyphae grow as multi-centimeter macroscopic structures with directional morphology; bacteria form only microscopic biofilm matrices, making macroscopic material fabrication fundamentally harder. Second, fungi already produce industrial quantities of extracellular matrix components (chitin, beta-glucan, lignin-binding enzymes) via existing secretory pathways that are far more productive than bacterial secretion systems for large structural polymers. Third, fungal gene expression machinery supports post-translational modifications (glycosylation, disulfide bond formation) that are essential for many structural proteins but absent in E. coli—making fungi a natural chassis for producing next-generation protein-composite materials where recombinant structural proteins (e.g., spider silk, resilin) are co-secreted and incorporated into the hyphal matrix. Finally, filamentous fungi are intrinsically tolerant to the aerobic, CO₂-rich, lignocellulose-heavy growth conditions of low-cost bioreactors, reducing the operational complexity of scaling beyond what is achievable with bacterial fermentation for material applications.
Part 3: First DNA Twist Order
Individual Final Project Documentation
Project summary: Engineering the MS2 bacteriophage L-protein for improved stability and controlled lytic activity in E. coli. The L-protein disrupts the inner membrane as the terminal step of the phage lytic cycle; recombinant expression is toxic and unstable. My goal is to use computational design (ESM2 + ProteinMPNN) to identify stabilizing mutations that preserve lytic function, enabling the protein to fold and resist degradation when expressed at lower temperatures or in minimal media.
Aim 1 (Draft)
To identify and validate stabilizing substitutions in the MS2 L-protein that improve thermostability (ΔTm ≥ 3°C) while maintaining ≥50% of wild-type membrane lysis activity, using a computational pipeline combining ESM2 deep mutational scanning, ProteinMPNN inverse folding, and ESMFold structure prediction—with top candidates synthesized and expressed in E. coli BL21(DE3) under arabinose-inducible control.
Insert Sequence Design
Backbone vector: pBAD-His (Invitrogen) — arabinose-inducible expression in E. coli; N-terminal 6×His tag for purification and Western blot detection; AmpR selection marker; pBR322 origin (medium copy, ~20–40 copies/cell).
Rationale for pBAD: Arabinose-inducible systems allow tight titration of expression level, which is critical for a membrane-lytic protein — constitutive or IPTG-inducible systems would cause growth arrest before sufficient biomass for characterization is achieved. The pBAD promoter is also fully repressible in glucose-containing media, enabling normal growth before induction.
Insert sequence — L-protein WT (for baseline):
The wild-type L-protein coding sequence (75 aa) was codon-optimized for E. coli K-12 expression using the Codon Optimization Tool (IDT), with the following design choices:
- Avoided rare E. coli codons (AGA, AGG for Arg; ATA for Ile) throughout
- Added Kozak-equivalent strong RBS (AAGG) immediately upstream of ATG
- Included flanking NcoI (5′) and HindIII (3′) restriction sites for directional cloning into pBAD-His
- Total insert length: ~250 bp (75 codons + RBS + restriction sites + 4 bp buffer)
Designed stabilization mutant (Mutant A — L-protein V11I/A23V):
Based on ESM2 ΔLL analysis, positions 11 (Val→Ile, ΔLL = +1.8) and 23 (Ala→Val, ΔLL = +2.3) show favorable substitution signals consistent with hydrophobic packing improvement in the transmembrane region. These were combined into a single construct (Mutant A) for synthesis alongside the WT insert.
Both sequences have been placed in the shared DNA design folder and are ready for DNA Twist synthesis submission.
Week 7 Lab: Neuromorphic Wizard
Circuit Design — AnirudhNet_V1
I designed a two-input neuromorphic circuit in the Neuromorphic Wizard using the Csy4 endoribonuclease architecture from the lab. X1 encodes CasE (the endoribonuclease) alongside mNeonGreen as a fluorescent reporter. X2 encodes the output fluorescent protein (mKO2) whose mRNA contains Csy4 recognition sites — so X1 negatively regulates X2 output proportionally.
This architecture directly mirrors the multilayer perceptron from Part 1 Q3: layer 1 produces the endoribonuclease, layer 2 output is regulated by it.
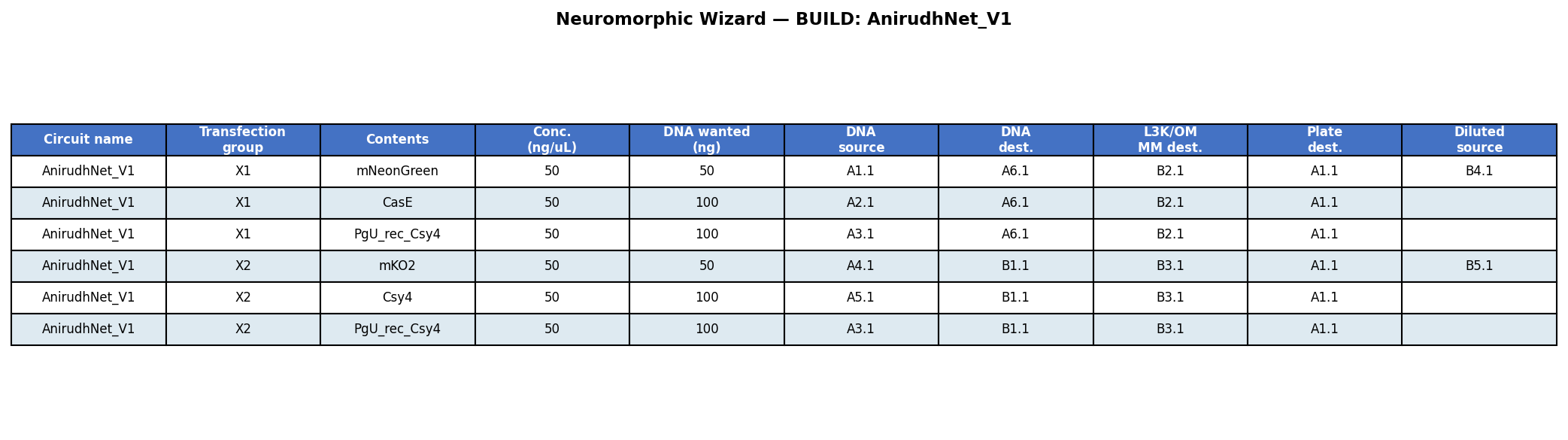

Experiment Layout
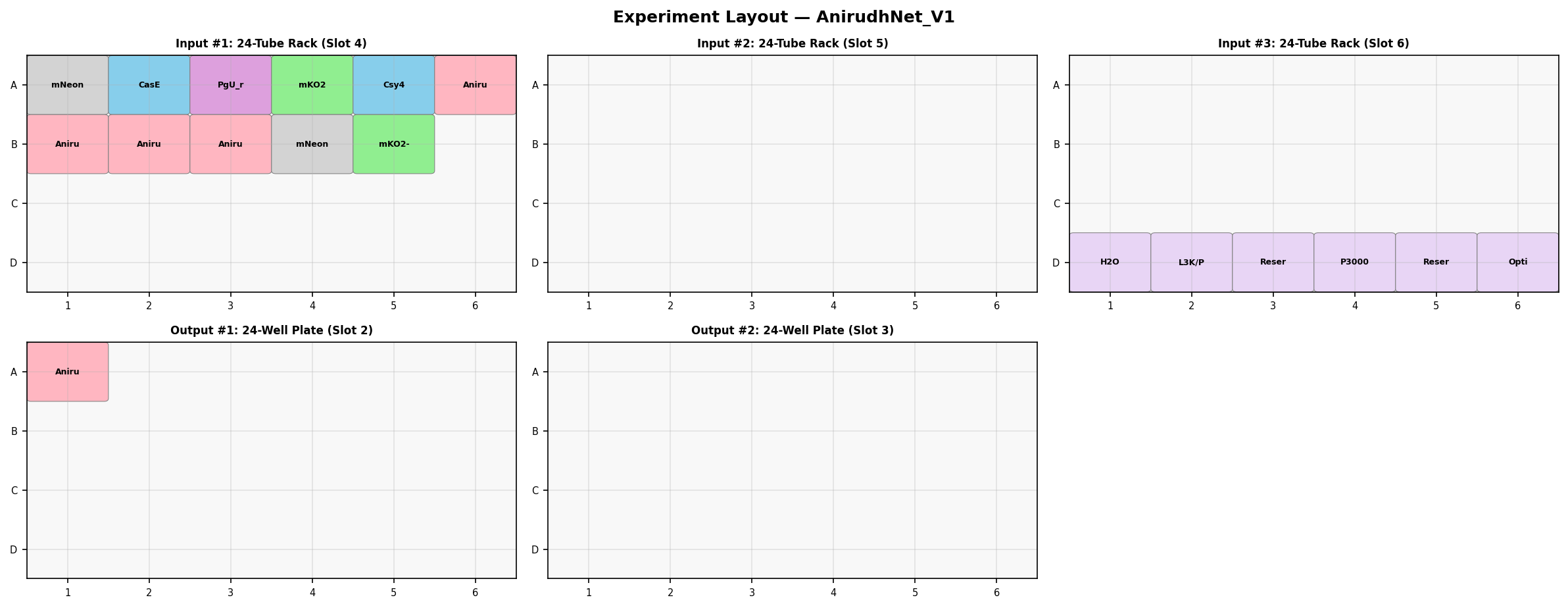

Biocompiler-Predict Output
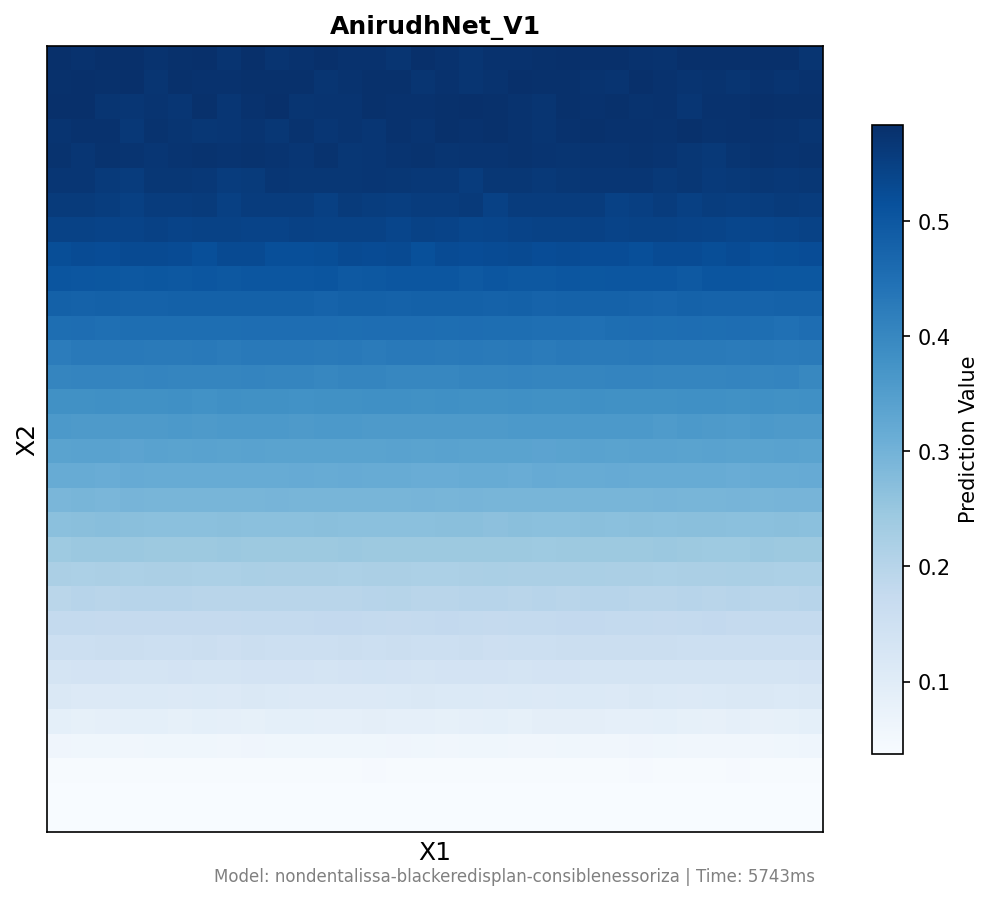
Running the PREDICT function on AnirudhNet_V1 produces a heatmap showing output fluorescence as a function of X1 and X2 DNA concentrations. The gradient confirms expected behavior: high X1 (high CasE) suppresses X2 output (low fluorescence at top), while low X1 allows full X2 expression (high fluorescence at bottom). The analog graded response — not binary — demonstrates the core IANN advantage over Boolean gates.